
diff --git a/deepdoc/README_tr.md b/deepdoc/README_tr.md
new file mode 100644
index 00000000000..434a4cce3ff
--- /dev/null
+++ b/deepdoc/README_tr.md
@@ -0,0 +1,136 @@
+[English](./README.md) | [简体中文](./README_zh.md) | Türkçe
+
+# *Deep*Doc
+
+- [*Deep*Doc](#deepdoc)
+ - [1. Giriş](#1-giriş)
+ - [2. Görsel İşleme](#2-görsel-i̇şleme)
+ - [3. Ayrıştırıcı](#3-ayrıştırıcı)
+ - [Özgeçmiş](#özgeçmiş)
+
+
+## 1. Giriş
+
+Farklı alanlardan, farklı formatlarda ve farklı erişim gereksinimleriyle gelen çok sayıda doküman için doğru bir analiz son derece zorlu bir görev haline gelmektedir. *Deep*Doc tam bu amaç için doğmuştur. Şu ana kadar *Deep*Doc'ta iki bileşen bulunmaktadır: görsel işleme ve ayrıştırıcı. OCR, yerleşim tanıma ve TSR sonuçlarımızla ilgileniyorsanız aşağıdaki test programlarını çalıştırabilirsiniz.
+
+```bash
+python deepdoc/vision/t_ocr.py -h
+usage: t_ocr.py [-h] --inputs INPUTS [--output_dir OUTPUT_DIR]
+
+options:
+ -h, --help show this help message and exit
+ --inputs INPUTS Directory where to store images or PDFs, or a file path to a single image or PDF
+ --output_dir OUTPUT_DIR
+ Directory where to store the output images. Default: './ocr_outputs'
+```
+
+```bash
+python deepdoc/vision/t_recognizer.py -h
+usage: t_recognizer.py [-h] --inputs INPUTS [--output_dir OUTPUT_DIR] [--threshold THRESHOLD] [--mode {layout,tsr}]
+
+options:
+ -h, --help show this help message and exit
+ --inputs INPUTS Directory where to store images or PDFs, or a file path to a single image or PDF
+ --output_dir OUTPUT_DIR
+ Directory where to store the output images. Default: './layouts_outputs'
+ --threshold THRESHOLD
+ A threshold to filter out detections. Default: 0.5
+ --mode {layout,tsr} Task mode: layout recognition or table structure recognition
+```
+
+Modellerimiz HuggingFace üzerinden sunulmaktadır. HuggingFace modellerini indirmekte sorun yaşıyorsanız, bu yardımcı olabilir!
+
+```bash
+export HF_ENDPOINT=https://hf-mirror.com
+```
+
+
+## 2. Görsel İşleme
+
+İnsanlar olarak sorunları çözmek için görsel bilgiyi kullanırız.
+
+ - **OCR (Optik Karakter Tanıma)**. Birçok doküman görsel olarak sunulduğundan veya en azından görsele dönüştürülebildiğinden, OCR metin çıkarımı için çok temel, önemli ve hatta evrensel bir çözümdür.
+ ```bash
+ python deepdoc/vision/t_ocr.py --inputs=gorsel_veya_pdf_yolu --output_dir=sonuc_klasoru
+ ```
+ Girdi, görseller veya PDF'ler içeren bir dizin ya da tek bir görsel veya PDF dosyası olabilir.
+ Sonuçların konumlarını gösteren görsellerin ve OCR metnini içeren txt dosyalarının bulunduğu `sonuc_klasoru` klasörüne bakabilirsiniz.
+
+

+
+
+ - **Yerleşim Tanıma (Layout Recognition)**. Farklı alanlardan gelen dokümanlar farklı yerleşimlere sahip olabilir; gazete, dergi, kitap ve özgeçmiş gibi dokümanlar yerleşim açısından birbirinden farklıdır. Yalnızca makine doğru bir yerleşim analizi yapabildiğinde, metin parçalarının ardışık olup olmadığına, bu parçanın Tablo Yapısı Tanıma (TSR) ile mi işlenmesi gerektiğine veya bu parçanın bir şekil olup bu başlıkla mı açıklandığına karar verebilir.
+ Çoğu durumu kapsayan 10 temel yerleşim bileşenimiz vardır:
+ - Metin
+ - Başlık
+ - Şekil
+ - Şekil açıklaması
+ - Tablo
+ - Tablo açıklaması
+ - Üst bilgi
+ - Alt bilgi
+ - Referans
+ - Denklem
+
+ Yerleşim algılama sonuçlarını görmek için aşağıdaki komutu deneyin.
+ ```bash
+ python deepdoc/vision/t_recognizer.py --inputs=gorsel_veya_pdf_yolu --threshold=0.2 --mode=layout --output_dir=sonuc_klasoru
+ ```
+ Girdi, görseller veya PDF'ler içeren bir dizin ya da tek bir görsel veya PDF dosyası olabilir.
+ Aşağıdaki gibi algılama sonuçlarını gösteren görsellerin bulunduğu `sonuc_klasoru` klasörüne bakabilirsiniz:
+
+

+
+
+ - **TSR (Tablo Yapısı Tanıma)**. Veri tablosu, sayılar veya metin dahil verileri sunmak için sıklıkla kullanılan bir yapıdır. Bir tablonun yapısı; hiyerarşik başlıklar, birleştirilmiş hücreler ve yansıtılmış satır başlıkları gibi çok karmaşık olabilir. TSR'nin yanı sıra, içeriği LLM tarafından iyi anlaşılabilecek cümlelere dönüştürüyoruz.
+ TSR görevi için beş etiketimiz vardır:
+ - Sütun
+ - Satır
+ - Sütun başlığı
+ - Yansıtılmış satır başlığı
+ - Birleştirilmiş hücre
+
+ Algılama sonuçlarını görmek için aşağıdaki komutu deneyin.
+ ```bash
+ python deepdoc/vision/t_recognizer.py --inputs=gorsel_veya_pdf_yolu --threshold=0.2 --mode=tsr --output_dir=sonuc_klasoru
+ ```
+ Girdi, görseller veya PDF'ler içeren bir dizin ya da tek bir görsel veya PDF dosyası olabilir.
+ Algılama sonuçlarını gösteren görsellerin ve HTML sayfalarının bulunduğu `sonuc_klasoru` klasörüne bakabilirsiniz:
+
+

+
+
+ - **Tablo Otomatik Döndürme**. Tabloların yanlış yönde olabileceği (90°, 180° veya 270° döndürülmüş) taranmış PDF'ler için, PDF ayrıştırıcısı tablo yapısı tanıma işleminden önce en iyi döndürme açısını OCR güven puanlarını kullanarak otomatik olarak algılar. Bu, döndürülmüş tablolar için OCR doğruluğunu ve tablo yapısı algılamasını önemli ölçüde artırır.
+
+ Özellik 4 döndürme açısını (0°, 90°, 180°, 270°) değerlendirir ve en yüksek OCR güvenine sahip olanı seçer. En iyi yönlendirmeyi belirledikten sonra, doğru döndürülmüş tablo görseli üzerinde OCR'yi yeniden gerçekleştirir.
+
+ Bu özellik **varsayılan olarak etkindir**. Ortam değişkeni ile kontrol edebilirsiniz:
+ ```bash
+ # Tablo otomatik döndürmeyi devre dışı bırak
+ export TABLE_AUTO_ROTATE=false
+
+ # Tablo otomatik döndürmeyi etkinleştir (varsayılan)
+ export TABLE_AUTO_ROTATE=true
+ ```
+
+ Veya API parametresi ile:
+ ```python
+ from deepdoc.parser import PdfParser
+
+ parser = PdfParser()
+ # Bu çağrı için otomatik döndürmeyi devre dışı bırak
+ boxes, tables = parser(pdf_path, auto_rotate_tables=False)
+ ```
+
+
+## 3. Ayrıştırıcı
+
+PDF, DOCX, EXCEL ve PPT olmak üzere dört doküman formatının kendine özgü ayrıştırıcısı vardır. En karmaşık olanı, PDF'nin esnekliği nedeniyle PDF ayrıştırıcısıdır. PDF ayrıştırıcısının çıktısı şunları içerir:
+ - PDF'deki konumlarıyla birlikte metin parçaları (sayfa numarası ve dikdörtgen konumları).
+ - PDF'den kırpılmış görsel ve doğal dil cümlelerine çevrilmiş içerikleriyle tablolar.
+ - Açıklama ve şekil içindeki metinlerle birlikte şekiller.
+
+### Özgeçmiş
+
+Özgeçmiş çok karmaşık bir doküman türüdür. Çeşitli yerleşimlere sahip yapılandırılmamış metinden oluşan bir özgeçmiş, yaklaşık yüz alanı kapsayan yapılandırılmış veriye dönüştürülebilir.
+Ayrıştırıcıyı henüz açık kaynak olarak yayınlamadık; ayrıştırma prosedüründen sonraki işleme yöntemini açık kaynak olarak sunmaktayız.
diff --git a/deepdoc/parser/__init__.py b/deepdoc/parser/__init__.py
index 809a56edf70..a34b1de0f39 100644
--- a/deepdoc/parser/__init__.py
+++ b/deepdoc/parser/__init__.py
@@ -15,6 +15,7 @@
#
from .docx_parser import RAGFlowDocxParser as DocxParser
+from .epub_parser import RAGFlowEpubParser as EpubParser
from .excel_parser import RAGFlowExcelParser as ExcelParser
from .html_parser import RAGFlowHtmlParser as HtmlParser
from .json_parser import RAGFlowJsonParser as JsonParser
@@ -29,6 +30,7 @@
"PdfParser",
"PlainParser",
"DocxParser",
+ "EpubParser",
"ExcelParser",
"PptParser",
"HtmlParser",
@@ -37,4 +39,3 @@
"TxtParser",
"MarkdownElementExtractor",
]
-
diff --git a/deepdoc/parser/docling_parser.py b/deepdoc/parser/docling_parser.py
index e8df1cfd4ee..a2ebc400255 100644
--- a/deepdoc/parser/docling_parser.py
+++ b/deepdoc/parser/docling_parser.py
@@ -17,6 +17,8 @@
import logging
import re
+import base64
+import os
from dataclasses import dataclass
from enum import Enum
from io import BytesIO
@@ -25,6 +27,7 @@
from typing import Any, Callable, Iterable, Optional
import pdfplumber
+import requests
from PIL import Image
try:
@@ -38,6 +41,8 @@
class RAGFlowPdfParser:
pass
+from deepdoc.parser.utils import extract_pdf_outlines
+
class DoclingContentType(str, Enum):
IMAGE = "image"
@@ -55,16 +60,60 @@ class _BBox:
y1: float
+def _extract_bbox_from_prov(item, prov_attr: str = "prov") -> Optional[_BBox]:
+ prov = getattr(item, prov_attr, None)
+ if not prov:
+ return None
+
+ prov_item = prov[0] if isinstance(prov, list) else prov
+ pn = getattr(prov_item, "page_no", None)
+ bb = getattr(prov_item, "bbox", None)
+ if pn is None or bb is None:
+ return None
+
+ coords = [getattr(bb, attr) for attr in ("l", "t", "r", "b")]
+ if None in coords:
+ return None
+
+ return _BBox(page_no=int(pn), x0=coords[0], y0=coords[1], x1=coords[2], y1=coords[3])
+
+
class DoclingParser(RAGFlowPdfParser):
- def __init__(self):
+ def __init__(self, docling_server_url: str = "", request_timeout: int = 600):
self.logger = logging.getLogger(self.__class__.__name__)
self.page_images: list[Image.Image] = []
self.page_from = 0
self.page_to = 10_000
self.outlines = []
-
-
- def check_installation(self) -> bool:
+ self.docling_server_url = (docling_server_url or "").rstrip("/")
+ self.request_timeout = request_timeout
+
+ def _effective_server_url(self, docling_server_url: Optional[str] = None) -> str:
+ return (docling_server_url or self.docling_server_url or "").rstrip("/") or (
+ os.environ.get("DOCLING_SERVER_URL", "").rstrip("/")
+ )
+
+ @staticmethod
+ def _is_http_endpoint_valid(url: str, timeout: int = 5) -> bool:
+ try:
+ response = requests.head(url, timeout=timeout, allow_redirects=True)
+ return response.status_code in [200, 301, 302, 307, 308]
+ except Exception:
+ try:
+ response = requests.get(url, timeout=timeout, allow_redirects=True)
+ return response.status_code in [200, 301, 302, 307, 308]
+ except Exception:
+ return False
+
+ def check_installation(self, docling_server_url: Optional[str] = None) -> bool:
+ server_url = self._effective_server_url(docling_server_url)
+ if server_url:
+ for path in ("/openapi.json", "/docs", "/v1/convert/source"):
+ if self._is_http_endpoint_valid(f"{server_url}{path}", timeout=5):
+ return True
+ self.logger.warning(f"[Docling] external server not reachable: {server_url}")
+ return False
+
if DocumentConverter is None:
self.logger.warning("[Docling] 'docling' is not importable, please: pip install docling")
return False
@@ -168,34 +217,22 @@ def crop(self, text: str, ZM: int = 1, need_position: bool = False):
def _iter_doc_items(self, doc) -> Iterable[tuple[str, Any, Optional[_BBox]]]:
for t in getattr(doc, "texts", []):
- parent=getattr(t, "parent", "")
- ref=getattr(parent,"cref","")
- label=getattr(t, "label", "")
- if (label in ("section_header","text",) and ref in ("#/body",)) or label in ("list_item",):
+ parent = getattr(t, "parent", "")
+ ref = getattr(parent, "cref", "")
+ label = getattr(t, "label", "")
+ if (label in ("section_header", "text") and ref in ("#/body",)) or label in ("list_item",):
text = getattr(t, "text", "") or ""
- bbox = None
- if getattr(t, "prov", None):
- pn = getattr(t.prov[0], "page_no", None)
- bb = getattr(t.prov[0], "bbox", None)
- bb = [getattr(bb, "l", None),getattr(bb, "t", None),getattr(bb, "r", None),getattr(bb, "b", None)]
- if pn and bb and len(bb) == 4:
- bbox = _BBox(page_no=int(pn), x0=bb[0], y0=bb[1], x1=bb[2], y1=bb[3])
+ bbox = _extract_bbox_from_prov(t)
yield (DoclingContentType.TEXT.value, text, bbox)
for item in getattr(doc, "texts", []):
if getattr(item, "label", "") in ("FORMULA",):
text = getattr(item, "text", "") or ""
- bbox = None
- if getattr(item, "prov", None):
- pn = getattr(item.prov, "page_no", None)
- bb = getattr(item.prov, "bbox", None)
- bb = [getattr(bb, "l", None),getattr(bb, "t", None),getattr(bb, "r", None),getattr(bb, "b", None)]
- if pn and bb and len(bb) == 4:
- bbox = _BBox(int(pn), bb[0], bb[1], bb[2], bb[3])
+ bbox = _extract_bbox_from_prov(item)
yield (DoclingContentType.EQUATION.value, text, bbox)
- def _transfer_to_sections(self, doc, parse_method: str) -> list[tuple[str, str]]:
- sections: list[tuple[str, str]] = []
+ def _transfer_to_sections(self, doc, parse_method: str) -> list[tuple[str, ...]]:
+ sections: list[tuple[str, ...]] = []
for typ, payload, bbox in self._iter_doc_items(doc):
if typ == DoclingContentType.TEXT.value:
section = payload.strip()
@@ -207,7 +244,7 @@ def _transfer_to_sections(self, doc, parse_method: str) -> list[tuple[str, str]]
continue
tag = self._make_line_tag(bbox) if isinstance(bbox,_BBox) else ""
- if parse_method == "manual":
+ if parse_method in {"manual", "pipeline"}:
sections.append((section, typ, tag))
elif parse_method == "paper":
sections.append((section + tag, typ))
@@ -248,16 +285,9 @@ def _transfer_to_tables(self, doc):
for tab in getattr(doc, "tables", []):
img = None
positions = ""
- if getattr(tab, "prov", None):
- pn = getattr(tab.prov[0], "page_no", None)
- bb = getattr(tab.prov[0], "bbox", None)
- if pn is not None and bb is not None:
- left = getattr(bb, "l", None)
- top = getattr(bb, "t", None)
- right = getattr(bb, "r", None)
- bott = getattr(bb, "b", None)
- if None not in (left, top, right, bott):
- img, positions = self.cropout_docling_table(int(pn), (float(left), float(top), float(right), float(bott)))
+ bbox = _extract_bbox_from_prov(tab)
+ if bbox:
+ img, positions = self.cropout_docling_table(bbox.page_no, (bbox.x0, bbox.y0, bbox.x1, bbox.y1))
html = ""
try:
html = tab.export_to_html(doc=doc)
@@ -267,16 +297,9 @@ def _transfer_to_tables(self, doc):
for pic in getattr(doc, "pictures", []):
img = None
positions = ""
- if getattr(pic, "prov", None):
- pn = getattr(pic.prov[0], "page_no", None)
- bb = getattr(pic.prov[0], "bbox", None)
- if pn is not None and bb is not None:
- left = getattr(bb, "l", None)
- top = getattr(bb, "t", None)
- right = getattr(bb, "r", None)
- bott = getattr(bb, "b", None)
- if None not in (left, top, right, bott):
- img, positions = self.cropout_docling_table(int(pn), (float(left), float(top), float(right), float(bott)))
+ bbox = _extract_bbox_from_prov(pic)
+ if bbox:
+ img, positions = self.cropout_docling_table(bbox.page_no, (bbox.x0, bbox.y0, bbox.x1, bbox.y1))
captions = ""
try:
captions = pic.caption_text(doc=doc)
@@ -285,6 +308,141 @@ def _transfer_to_tables(self, doc):
tables.append(((img, [captions]), positions if positions else ""))
return tables
+ @staticmethod
+ def _sections_from_remote_text(text: str, parse_method: str) -> list[tuple[str, ...]]:
+ txt = (text or "").strip()
+ if not txt:
+ return []
+ if parse_method in {"manual", "pipeline"}:
+ return [(txt, DoclingContentType.TEXT.value, "")]
+ if parse_method == "paper":
+ return [(txt, DoclingContentType.TEXT.value)]
+ return [(txt, "")]
+
+ @staticmethod
+ def _extract_remote_document_entries(payload: Any) -> list[dict[str, Any]]:
+ if not isinstance(payload, dict):
+ return []
+ if isinstance(payload.get("document"), dict):
+ return [payload["document"]]
+ if isinstance(payload.get("documents"), list):
+ return [d for d in payload["documents"] if isinstance(d, dict)]
+ if isinstance(payload.get("results"), list):
+ docs = []
+ for it in payload["results"]:
+ if isinstance(it, dict):
+ if isinstance(it.get("document"), dict):
+ docs.append(it["document"])
+ elif isinstance(it.get("result"), dict):
+ docs.append(it["result"])
+ else:
+ docs.append(it)
+ return docs
+ return []
+
+ def _parse_pdf_remote(
+ self,
+ filepath: str | PathLike[str],
+ binary: BytesIO | bytes | None = None,
+ callback: Optional[Callable] = None,
+ *,
+ parse_method: str = "raw",
+ docling_server_url: Optional[str] = None,
+ request_timeout: Optional[int] = None,
+ ):
+ server_url = self._effective_server_url(docling_server_url)
+ if not server_url:
+ raise RuntimeError("[Docling] DOCLING_SERVER_URL is not configured.")
+
+ timeout = request_timeout or self.request_timeout
+ if binary is not None:
+ if isinstance(binary, (bytes, bytearray)):
+ pdf_bytes = bytes(binary)
+ else:
+ pdf_bytes = bytes(binary.getbuffer())
+ else:
+ src_path = Path(filepath)
+ if not src_path.exists():
+ raise FileNotFoundError(f"PDF not found: {src_path}")
+ with open(src_path, "rb") as f:
+ pdf_bytes = f.read()
+
+ if callback:
+ callback(0.2, f"[Docling] Requesting external server: {server_url}")
+
+ filename = Path(filepath).name or "input.pdf"
+ b64 = base64.b64encode(pdf_bytes).decode("ascii")
+ v1_payload = {
+ "options": {
+ "from_formats": ["pdf"],
+ "to_formats": ["json", "md", "text"],
+ },
+ "sources": [
+ {
+ "kind": "file",
+ "filename": filename,
+ "base64_string": b64,
+ }
+ ],
+ }
+ v1alpha_payload = {
+ "options": {
+ "from_formats": ["pdf"],
+ "to_formats": ["json", "md", "text"],
+ },
+ "file_sources": [
+ {
+ "filename": filename,
+ "base64_string": b64,
+ }
+ ],
+ }
+ errors = []
+ response_json = None
+ for endpoint, payload in (
+ ("/v1/convert/source", v1_payload),
+ ("/v1alpha/convert/source", v1alpha_payload),
+ ):
+ try:
+ resp = requests.post(
+ f"{server_url}{endpoint}",
+ json=payload,
+ timeout=timeout,
+ )
+ if resp.status_code < 300:
+ response_json = resp.json()
+ break
+ errors.append(f"{endpoint}: HTTP {resp.status_code} {resp.text[:300]}")
+ except Exception as exc:
+ errors.append(f"{endpoint}: {exc}")
+
+ if response_json is None:
+ raise RuntimeError("[Docling] remote convert failed: " + " | ".join(errors))
+
+ docs = self._extract_remote_document_entries(response_json)
+ if not docs:
+ raise RuntimeError("[Docling] remote response does not contain parsed documents.")
+
+ sections: list[tuple[str, ...]] = []
+ tables = []
+ for doc in docs:
+ md = doc.get("md_content")
+ txt = doc.get("text_content")
+ if isinstance(md, str) and md.strip():
+ sections.extend(self._sections_from_remote_text(md, parse_method=parse_method))
+ elif isinstance(txt, str) and txt.strip():
+ sections.extend(self._sections_from_remote_text(txt, parse_method=parse_method))
+
+ json_content = doc.get("json_content")
+ if isinstance(json_content, dict):
+ md_fallback = json_content.get("md_content")
+ if isinstance(md_fallback, str) and md_fallback.strip() and not sections:
+ sections.extend(self._sections_from_remote_text(md_fallback, parse_method=parse_method))
+
+ if callback:
+ callback(0.95, f"[Docling] Remote sections: {len(sections)}")
+ return sections, tables
+
def parse_pdf(
self,
filepath: str | PathLike[str],
@@ -295,12 +453,26 @@ def parse_pdf(
lang: Optional[str] = None,
method: str = "auto",
delete_output: bool = True,
- parse_method: str = "raw"
+ parse_method: str = "raw",
+ docling_server_url: Optional[str] = None,
+ request_timeout: Optional[int] = None,
):
+ self.outlines = extract_pdf_outlines(binary if binary is not None else filepath)
- if not self.check_installation():
+ if not self.check_installation(docling_server_url=docling_server_url):
raise RuntimeError("Docling not available, please install `docling`")
+ server_url = self._effective_server_url(docling_server_url)
+ if server_url:
+ return self._parse_pdf_remote(
+ filepath=filepath,
+ binary=binary,
+ callback=callback,
+ parse_method=parse_method,
+ docling_server_url=server_url,
+ request_timeout=request_timeout,
+ )
+
if binary is not None:
tmpdir = Path(output_dir) if output_dir else Path.cwd() / ".docling_tmp"
tmpdir.mkdir(parents=True, exist_ok=True)
diff --git a/deepdoc/parser/docx_parser.py b/deepdoc/parser/docx_parser.py
index 2a65841e246..0257a320f7f 100644
--- a/deepdoc/parser/docx_parser.py
+++ b/deepdoc/parser/docx_parser.py
@@ -20,9 +20,54 @@
from collections import Counter
from rag.nlp import rag_tokenizer
from io import BytesIO
-
+import logging
+from docx.image.exceptions import (
+ InvalidImageStreamError,
+ UnexpectedEndOfFileError,
+ UnrecognizedImageError,
+)
+from rag.utils.lazy_image import LazyImage
class RAGFlowDocxParser:
+ def get_picture(self, document, paragraph):
+ imgs = paragraph._element.xpath(".//pic:pic")
+ if not imgs:
+ return None
+ image_blobs = []
+ for img in imgs:
+ embed = img.xpath(".//a:blip/@r:embed")
+ if not embed:
+ continue
+ embed = embed[0]
+ image_blob = None
+ try:
+ related_part = document.part.related_parts[embed]
+ except Exception as e:
+ logging.warning(f"Skipping image due to unexpected error getting related_part: {e}")
+ continue
+
+ try:
+ image = related_part.image
+ if image is not None:
+ image_blob = image.blob
+ except (
+ UnrecognizedImageError,
+ UnexpectedEndOfFileError,
+ InvalidImageStreamError,
+ UnicodeDecodeError,
+ ) as e:
+ logging.info(f"Damaged image encountered, attempting blob fallback: {e}")
+ except Exception as e:
+ logging.warning(f"Unexpected error getting image, attempting blob fallback: {e}")
+
+ if image_blob is None:
+ image_blob = getattr(related_part, "blob", None)
+ if image_blob:
+ image_blobs.append(image_blob)
+ if not image_blobs:
+ return None
+ return LazyImage(image_blobs)
+
def __extract_table_content(self, tb):
df = []
diff --git a/deepdoc/parser/epub_parser.py b/deepdoc/parser/epub_parser.py
new file mode 100644
index 00000000000..5badd7c33b6
--- /dev/null
+++ b/deepdoc/parser/epub_parser.py
@@ -0,0 +1,145 @@
+#
+# Copyright 2025 The InfiniFlow Authors. All Rights Reserved.
+#
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in compliance with the License.
+# You may obtain a copy of the License at
+#
+# http://www.apache.org/licenses/LICENSE-2.0
+#
+# Unless required by applicable law or agreed to in writing, software
+# distributed under the License is distributed on an "AS IS" BASIS,
+# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
+# See the License for the specific language governing permissions and
+# limitations under the License.
+#
+
+import logging
+import warnings
+import zipfile
+from io import BytesIO
+from xml.etree import ElementTree
+
+from .html_parser import RAGFlowHtmlParser
+
+# OPF XML namespaces
+_OPF_NS = "http://www.idpf.org/2007/opf"
+_CONTAINER_NS = "urn:oasis:names:tc:opendocument:xmlns:container"
+
+# Media types that contain readable XHTML content
+_XHTML_MEDIA_TYPES = {"application/xhtml+xml", "text/html", "text/xml"}
+
+logger = logging.getLogger(__name__)
+
+
+class RAGFlowEpubParser:
+ """Parse EPUB files by extracting XHTML content in spine (reading) order
+ and delegating to RAGFlowHtmlParser for chunking."""
+
+ def __call__(self, fnm, binary=None, chunk_token_num=512):
+ if binary is not None:
+ if not binary:
+ logger.warning(
+ "RAGFlowEpubParser received an empty EPUB binary payload for %r",
+ fnm,
+ )
+ raise ValueError("Empty EPUB binary payload")
+ zf = zipfile.ZipFile(BytesIO(binary))
+ else:
+ zf = zipfile.ZipFile(fnm)
+
+ try:
+ content_items = self._get_spine_items(zf)
+ all_sections = []
+ html_parser = RAGFlowHtmlParser()
+
+ for item_path in content_items:
+ try:
+ html_bytes = zf.read(item_path)
+ except KeyError:
+ continue
+ if not html_bytes:
+ logger.debug("Skipping empty EPUB content item: %s", item_path)
+ continue
+ with warnings.catch_warnings():
+ warnings.filterwarnings("ignore", category=UserWarning)
+ sections = html_parser(
+ item_path, binary=html_bytes, chunk_token_num=chunk_token_num
+ )
+ all_sections.extend(sections)
+
+ return all_sections
+ finally:
+ zf.close()
+
+ @staticmethod
+ def _get_spine_items(zf):
+ """Return content file paths in spine (reading) order."""
+ # 1. Find the OPF file path from META-INF/container.xml
+ try:
+ container_xml = zf.read("META-INF/container.xml")
+ except KeyError:
+ return RAGFlowEpubParser._fallback_xhtml_order(zf)
+
+ try:
+ container_root = ElementTree.fromstring(container_xml)
+ except ElementTree.ParseError:
+ logger.warning("Failed to parse META-INF/container.xml; falling back to XHTML order.")
+ return RAGFlowEpubParser._fallback_xhtml_order(zf)
+
+ rootfile_el = container_root.find(f".//{{{_CONTAINER_NS}}}rootfile")
+ if rootfile_el is None:
+ return RAGFlowEpubParser._fallback_xhtml_order(zf)
+
+ opf_path = rootfile_el.get("full-path", "")
+ if not opf_path:
+ return RAGFlowEpubParser._fallback_xhtml_order(zf)
+
+ # Base directory of the OPF file (content paths are relative to it)
+ opf_dir = opf_path.rsplit("/", 1)[0] + "/" if "/" in opf_path else ""
+
+ # 2. Parse the OPF file
+ try:
+ opf_xml = zf.read(opf_path)
+ except KeyError:
+ return RAGFlowEpubParser._fallback_xhtml_order(zf)
+
+ try:
+ opf_root = ElementTree.fromstring(opf_xml)
+ except ElementTree.ParseError:
+ logger.warning("Failed to parse OPF file '%s'; falling back to XHTML order.", opf_path)
+ return RAGFlowEpubParser._fallback_xhtml_order(zf)
+
+ # 3. Build id->href+mediatype map from
+ manifest = {}
+ for item in opf_root.findall(f".//{{{_OPF_NS}}}item"):
+ item_id = item.get("id", "")
+ href = item.get("href", "")
+ media_type = item.get("media-type", "")
+ if item_id and href:
+ manifest[item_id] = (href, media_type)
+
+ # 4. Walk to get reading order
+ spine_items = []
+ for itemref in opf_root.findall(f".//{{{_OPF_NS}}}itemref"):
+ idref = itemref.get("idref", "")
+ if idref not in manifest:
+ continue
+ href, media_type = manifest[idref]
+ if media_type not in _XHTML_MEDIA_TYPES:
+ continue
+ spine_items.append(opf_dir + href)
+
+ return (
+ spine_items if spine_items else RAGFlowEpubParser._fallback_xhtml_order(zf)
+ )
+
+ @staticmethod
+ def _fallback_xhtml_order(zf):
+ """Fallback: return all .xhtml/.html files sorted alphabetically."""
+ return sorted(
+ n
+ for n in zf.namelist()
+ if n.lower().endswith((".xhtml", ".html", ".htm"))
+ and not n.startswith("META-INF/")
+ )
diff --git a/deepdoc/parser/excel_parser.py b/deepdoc/parser/excel_parser.py
index 2fe3420192c..acbd98f228a 100644
--- a/deepdoc/parser/excel_parser.py
+++ b/deepdoc/parser/excel_parser.py
@@ -18,9 +18,9 @@
import pandas as pd
from openpyxl import Workbook, load_workbook
-from PIL import Image
from rag.nlp import find_codec
+from rag.utils.lazy_image import LazyImage
# copied from `/openpyxl/cell/cell.py`
ILLEGAL_CHARACTERS_RE = re.compile(r"[\000-\010]|[\013-\014]|[\016-\037]")
@@ -74,9 +74,16 @@ def clean_string(s):
return df.apply(lambda col: col.map(clean_string))
+ @staticmethod
+ def _fill_worksheet_from_dataframe(ws, df: pd.DataFrame):
+ for col_num, column_name in enumerate(df.columns, 1):
+ ws.cell(row=1, column=col_num, value=column_name)
+ for row_num, row in enumerate(df.values, 2):
+ for col_num, value in enumerate(row, 1):
+ ws.cell(row=row_num, column=col_num, value=value)
+
@staticmethod
def _dataframe_to_workbook(df):
- # if contains multiple sheets use _dataframes_to_workbook
if isinstance(df, dict) and len(df) > 1:
return RAGFlowExcelParser._dataframes_to_workbook(df)
@@ -84,30 +91,19 @@ def _dataframe_to_workbook(df):
wb = Workbook()
ws = wb.active
ws.title = "Data"
-
- for col_num, column_name in enumerate(df.columns, 1):
- ws.cell(row=1, column=col_num, value=column_name)
-
- for row_num, row in enumerate(df.values, 2):
- for col_num, value in enumerate(row, 1):
- ws.cell(row=row_num, column=col_num, value=value)
-
+ RAGFlowExcelParser._fill_worksheet_from_dataframe(ws, df)
return wb
-
+
@staticmethod
def _dataframes_to_workbook(dfs: dict):
wb = Workbook()
default_sheet = wb.active
wb.remove(default_sheet)
-
+
for sheet_name, df in dfs.items():
df = RAGFlowExcelParser._clean_dataframe(df)
ws = wb.create_sheet(title=sheet_name)
- for col_num, column_name in enumerate(df.columns, 1):
- ws.cell(row=1, column=col_num, value=column_name)
- for row_num, row in enumerate(df.values, 2):
- for col_num, value in enumerate(row, 1):
- ws.cell(row=row_num, column=col_num, value=value)
+ RAGFlowExcelParser._fill_worksheet_from_dataframe(ws, df)
return wb
@staticmethod
@@ -126,7 +122,7 @@ def _extract_images_from_worksheet(ws, sheetname=None):
for img in images:
try:
img_bytes = img._data()
- pil_img = Image.open(BytesIO(img_bytes)).convert("RGB")
+ lazy_img = LazyImage([img_bytes])
anchor = img.anchor
if hasattr(anchor, "_from") and hasattr(anchor, "_to"):
@@ -143,7 +139,7 @@ def _extract_images_from_worksheet(ws, sheetname=None):
item = {
"sheet": sheetname or ws.title,
- "image": pil_img,
+ "image": lazy_img,
"image_description": "",
"row_from": r1,
"col_from": c1,
diff --git a/deepdoc/parser/figure_parser.py b/deepdoc/parser/figure_parser.py
index ec5e333de28..e062f462538 100644
--- a/deepdoc/parser/figure_parser.py
+++ b/deepdoc/parser/figure_parser.py
@@ -20,29 +20,36 @@
from common.constants import LLMType
from api.db.services.llm_service import LLMBundle
+from api.db.joint_services.tenant_model_service import get_tenant_default_model_by_type
from common.connection_utils import timeout
from rag.app.picture import vision_llm_chunk as picture_vision_llm_chunk
from rag.prompts.generator import vision_llm_figure_describe_prompt, vision_llm_figure_describe_prompt_with_context
from rag.nlp import append_context2table_image4pdf
+from rag.utils.lazy_image import ensure_pil_image, open_image_for_processing, is_image_like
# need to delete before pr
def vision_figure_parser_figure_data_wrapper(figures_data_without_positions):
if not figures_data_without_positions:
return []
- return [
- (
- (figure_data[1], [figure_data[0]]),
- [(0, 0, 0, 0, 0)],
+ res = []
+ for figure_data in figures_data_without_positions:
+ img = ensure_pil_image(figure_data[1])
+ if not isinstance(img, Image.Image):
+ continue
+ res.append(
+ (
+ (img, [figure_data[0]]),
+ [(0, 0, 0, 0, 0)],
+ )
)
- for figure_data in figures_data_without_positions
- if isinstance(figure_data[1], Image.Image)
- ]
+ return res
def vision_figure_parser_docx_wrapper(sections, tbls, callback=None,**kwargs):
if not sections:
return tbls
try:
- vision_model = LLMBundle(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model_config = get_tenant_default_model_by_type(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model = LLMBundle(kwargs["tenant_id"], vision_model_config)
callback(0.7, "Visual model detected. Attempting to enhance figure extraction...")
except Exception:
vision_model = None
@@ -61,13 +68,14 @@ def vision_figure_parser_figure_xlsx_wrapper(images,callback=None, **kwargs):
if not images:
return []
try:
- vision_model = LLMBundle(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model_config = get_tenant_default_model_by_type(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model = LLMBundle(kwargs["tenant_id"], vision_model_config)
callback(0.2, "Visual model detected. Attempting to enhance Excel image extraction...")
except Exception:
vision_model = None
if vision_model:
figures_data = [((
- img["image"], # Image.Image
+ img["image"], # Image.Image or LazyImage (converted by ensure_pil_image)
[img["image_description"]] # description list (must be list)
),
[
@@ -89,14 +97,15 @@ def vision_figure_parser_pdf_wrapper(tbls, callback=None, **kwargs):
parser_config = kwargs.get("parser_config", {})
context_size = max(0, int(parser_config.get("image_context_size", 0) or 0))
try:
- vision_model = LLMBundle(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model_config = get_tenant_default_model_by_type(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model = LLMBundle(kwargs["tenant_id"], vision_model_config)
callback(0.7, "Visual model detected. Attempting to enhance figure extraction...")
except Exception:
vision_model = None
if vision_model:
def is_figure_item(item):
- return isinstance(item[0][0], Image.Image) and isinstance(item[0][1], list)
+ return is_image_like(item[0][0]) and isinstance(item[0][1], list)
figures_data = [item for item in tbls if is_figure_item(item)]
figure_contexts = []
@@ -127,13 +136,17 @@ def vision_figure_parser_docx_wrapper_naive(chunks, idx_lst, callback=None, **kw
if not chunks:
return []
try:
- vision_model = LLMBundle(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model_config = get_tenant_default_model_by_type(kwargs["tenant_id"], LLMType.IMAGE2TEXT)
+ vision_model = LLMBundle(kwargs["tenant_id"], vision_model_config)
callback(0.7, "Visual model detected. Attempting to enhance figure extraction...")
except Exception:
vision_model = None
if vision_model:
@timeout(30, 3)
def worker(idx, ck):
+ img, close_after = open_image_for_processing(ck.get("image"), allow_bytes=True)
+ if not isinstance(img, Image.Image):
+ return idx, ""
context_above = ck.get("context_above", "")
context_below = ck.get("context_below", "")
if context_above or context_below:
@@ -149,13 +162,20 @@ def worker(idx, ck):
prompt = vision_llm_figure_describe_prompt()
logging.info(f"[VisionFigureParser] figure={idx} context_len=0 prompt=default")
- description_text = picture_vision_llm_chunk(
- binary=ck.get("image"),
- vision_model=vision_model,
- prompt=prompt,
- callback=callback,
- )
- return idx, description_text
+ try:
+ description_text = picture_vision_llm_chunk(
+ binary=img,
+ vision_model=vision_model,
+ prompt=prompt,
+ callback=callback,
+ )
+ return idx, description_text
+ finally:
+ if close_after and isinstance(img, Image.Image):
+ try:
+ img.close()
+ except Exception:
+ pass
with ThreadPoolExecutor(max_workers=10) as executor:
futures = [
@@ -187,13 +207,19 @@ def _extract_figures_info(self, figures_data):
# position
if len(item) == 2 and isinstance(item[0], tuple) and len(item[0]) == 2 and isinstance(item[1], list) and isinstance(item[1][0], tuple) and len(item[1][0]) == 5:
img_desc = item[0]
- assert len(img_desc) == 2 and isinstance(img_desc[0], Image.Image) and isinstance(img_desc[1], list), "Should be (figure, [description])"
- self.figures.append(img_desc[0])
+ img = ensure_pil_image(img_desc[0])
+ if img is None:
+ continue
+ assert len(img_desc) == 2 and isinstance(img_desc[1], list), "Should be (figure, [description])"
+ self.figures.append(img)
self.descriptions.append(img_desc[1])
self.positions.append(item[1])
else:
- assert len(item) == 2 and isinstance(item[0], Image.Image) and isinstance(item[1], list), f"Unexpected form of figure data: get {len(item)=}, {item=}"
- self.figures.append(item[0])
+ img = ensure_pil_image(item[0])
+ if img is None:
+ continue
+ assert len(item) == 2 and isinstance(item[1], list), f"Unexpected form of figure data: get {len(item)=}, {item=}"
+ self.figures.append(img)
self.descriptions.append(item[1])
def _assemble(self):
diff --git a/deepdoc/parser/html_parser.py b/deepdoc/parser/html_parser.py
index dcf33a8bbd1..f4d360c6413 100644
--- a/deepdoc/parser/html_parser.py
+++ b/deepdoc/parser/html_parser.py
@@ -33,7 +33,7 @@ def get_encoding(file):
"table", "pre", "code", "blockquote",
"figure", "figcaption"
]
-TITLE_TAGS = {"h1": "#", "h2": "##", "h3": "###", "h4": "#####", "h5": "#####", "h6": "######"}
+TITLE_TAGS = {"h1": "#", "h2": "##", "h3": "###", "h4": "####", "h5": "#####", "h6": "######"}
class RAGFlowHtmlParser:
diff --git a/deepdoc/parser/markdown_parser.py b/deepdoc/parser/markdown_parser.py
index 900ef525ccf..e911a22ac8e 100644
--- a/deepdoc/parser/markdown_parser.py
+++ b/deepdoc/parser/markdown_parser.py
@@ -56,7 +56,7 @@ def replace_tables_with_rendered_html(pattern, table_list, render=True):
""",
re.VERBOSE,
)
- working_text = replace_tables_with_rendered_html(border_table_pattern, tables)
+ working_text = replace_tables_with_rendered_html(border_table_pattern, tables, render=separate_tables)
# Borderless Markdown table
no_border_table_pattern = re.compile(
@@ -68,7 +68,7 @@ def replace_tables_with_rendered_html(pattern, table_list, render=True):
""",
re.VERBOSE,
)
- working_text = replace_tables_with_rendered_html(no_border_table_pattern, tables)
+ working_text = replace_tables_with_rendered_html(no_border_table_pattern, tables, render=separate_tables)
# Replace any TAGS e.g. to
TAGS = ["table", "td", "tr", "th", "tbody", "thead", "div"]
diff --git a/deepdoc/parser/mineru_parser.py b/deepdoc/parser/mineru_parser.py
index cc4c99c76b8..25a0627ff41 100644
--- a/deepdoc/parser/mineru_parser.py
+++ b/deepdoc/parser/mineru_parser.py
@@ -35,6 +35,7 @@
from strenum import StrEnum
from deepdoc.parser.pdf_parser import RAGFlowPdfParser
+from deepdoc.parser.utils import extract_pdf_outlines
LOCK_KEY_pdfplumber = "global_shared_lock_pdfplumber"
if LOCK_KEY_pdfplumber not in sys.modules:
@@ -73,6 +74,8 @@ class MinerUContentType(StrEnum):
'Thai': 'th',
'Greek': 'el',
'Hindi': 'devanagari',
+ 'Bulgarian': 'cyrillic',
+ 'Turkish': 'latin',
}
@@ -339,6 +342,11 @@ def _line_tag(self, bx):
pn = [bx["page_idx"] + 1]
positions = bx.get("bbox", (0, 0, 0, 0))
x0, top, x1, bott = positions
+ # Normalize flipped coordinates (MinerU may report inverted bbox for flipped images)
+ if x0 > x1:
+ x0, x1 = x1, x0
+ if top > bott:
+ top, bott = bott, top
if hasattr(self, "page_images") and self.page_images and len(self.page_images) > bx["page_idx"]:
page_width, page_height = self.page_images[bx["page_idx"]].size
@@ -428,6 +436,12 @@ def crop(self, text, ZM=1, need_position=False):
img0 = self.page_images[pns[0]]
x0, y0, x1, y1 = int(left), int(top), int(right), int(min(bottom, img0.size[1]))
+ if x0 > x1:
+ x0, x1 = x1, x0
+ if y0 > y1:
+ y0, y1 = y1, y0
+ if x1 <= x0 or y1 <= y0:
+ continue
crop0 = img0.crop((x0, y0, x1, y1))
imgs.append(crop0)
if 0 < ii < len(poss) - 1:
@@ -441,6 +455,13 @@ def crop(self, text, ZM=1, need_position=False):
continue
page = self.page_images[pn]
x0, y0, x1, y1 = int(left), 0, int(right), int(min(bottom, page.size[1]))
+ if x0 > x1:
+ x0, x1 = x1, x0
+ if y0 > y1:
+ y0, y1 = y1, y0
+ if x1 <= x0 or y1 <= y0:
+ bottom -= page.size[1]
+ continue
cimgp = page.crop((x0, y0, x1, y1))
imgs.append(cimgp)
if 0 < ii < len(poss) - 1:
@@ -556,7 +577,7 @@ def _transfer_to_sections(self, outputs: list[dict[str, Any]], parse_method: str
case MinerUContentType.DISCARDED:
continue # Skip discarded blocks entirely
- if section and parse_method == "manual":
+ if section and parse_method in {"manual", "pipeline"}:
sections.append((section, output["type"], self._line_tag(output)))
elif section and parse_method == "paper":
sections.append((section + self._line_tag(output), output["type"]))
@@ -582,6 +603,7 @@ def parse_pdf(
) -> tuple:
import shutil
+ self.outlines = extract_pdf_outlines(binary if binary is not None else filepath)
temp_pdf = None
created_tmp_dir = False
diff --git a/deepdoc/parser/paddleocr_parser.py b/deepdoc/parser/paddleocr_parser.py
index 85db63b862d..a23852e89c0 100644
--- a/deepdoc/parser/paddleocr_parser.py
+++ b/deepdoc/parser/paddleocr_parser.py
@@ -36,6 +36,8 @@
class RAGFlowPdfParser:
pass
+from deepdoc.parser.utils import extract_pdf_outlines
+
AlgorithmType = Literal["PaddleOCR-VL"]
SectionTuple = tuple[str, ...]
@@ -59,11 +61,22 @@ def _remove_images_from_markdown(markdown: str) -> str:
return _MARKDOWN_IMAGE_PATTERN.sub("", markdown)
+def _normalize_bbox(bbox: list[Any] | tuple[Any, ...]) -> tuple[float, float, float, float]:
+ if len(bbox) < 4:
+ return 0.0, 0.0, 0.0, 0.0
+
+ left, top, right, bottom = (float(bbox[0]), float(bbox[1]), float(bbox[2]), float(bbox[3]))
+ if left > right:
+ left, right = right, left
+ if top > bottom:
+ top, bottom = bottom, top
+ return left, top, right, bottom
+
+
@dataclass
class PaddleOCRVLConfig:
"""Configuration for PaddleOCR-VL algorithm."""
- use_doc_orientation_classify: Optional[bool] = False
use_doc_orientation_classify: Optional[bool] = False
use_doc_unwarping: Optional[bool] = False
use_layout_detection: Optional[bool] = None
@@ -199,6 +212,7 @@ def __init__(
"""Initialize PaddleOCR parser."""
super().__init__()
+ self.outlines = []
self.api_url = api_url.rstrip("/") if api_url else os.getenv("PADDLEOCR_API_URL", "")
self.access_token = access_token or os.getenv("PADDLEOCR_ACCESS_TOKEN")
self.algorithm = algorithm
@@ -241,6 +255,7 @@ def parse_pdf(
**kwargs: Any,
) -> ParseResult:
"""Parse PDF document using PaddleOCR API."""
+ self.outlines = extract_pdf_outlines(binary if binary is not None else filepath)
# Create configuration - pass all kwargs to capture VL config parameters
config_dict = {
"api_url": api_url if api_url is not None else self.api_url,
@@ -393,10 +408,11 @@ def _transfer_to_sections(self, result: dict[str, Any], algorithm: AlgorithmType
label = block.get("block_label", "")
block_bbox = block.get("block_bbox", [0, 0, 0, 0])
+ left, top, right, bottom = _normalize_bbox(block_bbox)
- tag = f"@@{page_idx + 1}\t{block_bbox[0] // self._ZOOMIN}\t{block_bbox[2] // self._ZOOMIN}\t{block_bbox[1] // self._ZOOMIN}\t{block_bbox[3] // self._ZOOMIN}##"
+ tag = f"@@{page_idx + 1}\t{left // self._ZOOMIN}\t{right // self._ZOOMIN}\t{top // self._ZOOMIN}\t{bottom // self._ZOOMIN}##"
- if parse_method == "manual":
+ if parse_method in {"manual", "pipeline"}:
sections.append((block_content, label, tag))
elif parse_method == "paper":
sections.append((block_content + tag, label))
@@ -409,7 +425,7 @@ def _transfer_to_tables(self, result: dict[str, Any]) -> list[TableTuple]:
"""Convert API response to table tuples."""
return []
- def __images__(self, fnm, page_from=0, page_to=100, callback=None):
+ def __images__(self, fnm, page_from=0, page_to=10**9, callback=None):
"""Generate page images from PDF for cropping."""
self.page_from = page_from
self.page_to = page_to
@@ -509,6 +525,16 @@ def crop(self, text: str, need_position: bool = False):
img0 = self.page_images[pns[0]]
x0, y0, x1, y1 = int(left), int(top), int(right), int(min(bottom, img0.size[1]))
+ if x0 > x1:
+ x0, x1 = x1, x0
+ if y0 > y1:
+ y0, y1 = y1, y0
+ x0 = max(0, min(x0, img0.size[0]))
+ x1 = max(0, min(x1, img0.size[0]))
+ y0 = max(0, min(y0, img0.size[1]))
+ y1 = max(0, min(y1, img0.size[1]))
+ if x1 <= x0 or y1 <= y0:
+ continue
crop0 = img0.crop((x0, y0, x1, y1))
imgs.append(crop0)
if 0 < ii < len(poss) - 1:
@@ -521,6 +547,17 @@ def crop(self, text: str, need_position: bool = False):
continue
page = self.page_images[pn]
x0, y0, x1, y1 = int(left), 0, int(right), int(min(bottom, page.size[1]))
+ if x0 > x1:
+ x0, x1 = x1, x0
+ if y0 > y1:
+ y0, y1 = y1, y0
+ x0 = max(0, min(x0, page.size[0]))
+ x1 = max(0, min(x1, page.size[0]))
+ y0 = max(0, min(y0, page.size[1]))
+ y1 = max(0, min(y1, page.size[1]))
+ if x1 <= x0 or y1 <= y0:
+ bottom -= page.size[1]
+ continue
cimgp = page.crop((x0, y0, x1, y1))
imgs.append(cimgp)
if 0 < ii < len(poss) - 1:
@@ -532,21 +569,25 @@ def crop(self, text: str, need_position: bool = False):
return None, None
return
- height = 0
+ total_height = 0
+ max_width = 0
+ img_sizes = []
for img in imgs:
- height += img.size[1] + GAP
- height = int(height)
- width = int(np.max([i.size[0] for i in imgs]))
- pic = Image.new("RGB", (width, height), (245, 245, 245))
- height = 0
- for ii, img in enumerate(imgs):
- if ii == 0 or ii + 1 == len(imgs):
+ w, h = img.size
+ img_sizes.append((w, h))
+ max_width = max(max_width, w)
+ total_height += h + GAP
+
+ pic = Image.new("RGB", (max_width, int(total_height)), (245, 245, 245))
+ current_height = 0
+ imgs_count = len(imgs)
+ for ii, (img, (w, h)) in enumerate(zip(imgs, img_sizes)):
+ if ii == 0 or ii + 1 == imgs_count:
img = img.convert("RGBA")
- overlay = Image.new("RGBA", img.size, (0, 0, 0, 0))
- overlay.putalpha(128)
+ overlay = Image.new("RGBA", img.size, (0, 0, 0, 128))
img = Image.alpha_composite(img, overlay).convert("RGB")
- pic.paste(img, (0, int(height)))
- height += img.size[1] + GAP
+ pic.paste(img, (0, int(current_height)))
+ current_height += h + GAP
if need_position:
return pic, positions
diff --git a/deepdoc/parser/pdf_parser.py b/deepdoc/parser/pdf_parser.py
index 6681e4a893a..b3a6adec8b5 100644
--- a/deepdoc/parser/pdf_parser.py
+++ b/deepdoc/parser/pdf_parser.py
@@ -22,6 +22,7 @@
import re
import sys
import threading
+import unicodedata
from collections import Counter, defaultdict
from copy import deepcopy
from io import BytesIO
@@ -37,10 +38,10 @@
from sklearn.metrics import silhouette_score
from common.file_utils import get_project_base_directory
-from common.misc_utils import pip_install_torch
from deepdoc.vision import OCR, AscendLayoutRecognizer, LayoutRecognizer, Recognizer, TableStructureRecognizer
from rag.nlp import rag_tokenizer
from rag.prompts.generator import vision_llm_describe_prompt
+from deepdoc.parser.utils import extract_pdf_outlines
from common import settings
@@ -89,14 +90,9 @@ def __init__(self, **kwargs):
self.tbl_det = TableStructureRecognizer()
self.updown_cnt_mdl = xgb.Booster()
- try:
- pip_install_torch()
- import torch.cuda
-
- if torch.cuda.is_available():
- self.updown_cnt_mdl.set_param({"device": "cuda"})
- except Exception:
- logging.info("No torch found.")
+ # xgboost model is very small; using CPU explicitly
+ self.updown_cnt_mdl.set_param({"device": "cpu"})
+ logging.info("updown_cnt_mdl initialized on CPU")
try:
model_dir = os.path.join(get_project_base_directory(), "rag/res/deepdoc")
self.updown_cnt_mdl.load_model(os.path.join(model_dir, "updown_concat_xgb.model"))
@@ -197,6 +193,127 @@ def _has_color(self, o):
return False
return True
+ # CID pattern regex for unmapped font characters from pdfminer
+ _CID_PATTERN = re.compile(r"\(cid\s*:\s*\d+\s*\)")
+
+ @staticmethod
+ def _is_garbled_char(ch):
+ """Check if a single character is garbled (unmappable from PDF font encoding).
+
+ A character is considered garbled if it falls into Unicode Private Use Areas
+ or certain replacement/control character ranges that typically indicate
+ pdfminer failed to map a CID to a valid Unicode codepoint.
+ """
+ if not ch:
+ return False
+ cp = ord(ch)
+ if 0xE000 <= cp <= 0xF8FF:
+ return True
+ if 0xF0000 <= cp <= 0xFFFFF:
+ return True
+ if 0x100000 <= cp <= 0x10FFFF:
+ return True
+ if cp == 0xFFFD:
+ return True
+ if cp < 0x20 and ch not in ('\t', '\n', '\r'):
+ return True
+ if 0x80 <= cp <= 0x9F:
+ return True
+ cat = unicodedata.category(ch)
+ if cat in ("Cn", "Cs"):
+ return True
+ return False
+
+ @staticmethod
+ def _is_garbled_text(text, threshold=0.5):
+ """Check if a text string contains too many garbled characters.
+
+ Examines each character and determines if the overall proportion
+ of garbled characters exceeds the given threshold. Also detects
+ pdfminer's CID placeholder patterns like '(cid:123)'.
+ """
+ if not text or not text.strip():
+ return False
+ if RAGFlowPdfParser._CID_PATTERN.search(text):
+ return True
+ garbled_count = 0
+ total = 0
+ for ch in text:
+ if ch.isspace():
+ continue
+ total += 1
+ if RAGFlowPdfParser._is_garbled_char(ch):
+ garbled_count += 1
+ if total == 0:
+ return False
+ return garbled_count / total >= threshold
+
+ @staticmethod
+ def _has_subset_font_prefix(fontname):
+ """Check if a font name has a subset prefix (e.g. 'DY1+ZLQDm1-1').
+
+ PDF subset fonts use a 6-letter uppercase tag followed by '+' before
+ the actual font name. Some tools use shorter tags (e.g. 'DY1+').
+ """
+ if not fontname:
+ return False
+ return bool(re.match(r"^[A-Z0-9]{2,6}\+", fontname))
+
+ @staticmethod
+ def _is_garbled_by_font_encoding(page_chars, min_chars=20):
+ """Detect garbled text caused by broken font encoding mappings.
+
+ Some PDFs (especially older Chinese standards) embed custom fonts that
+ map CJK glyphs to ASCII codepoints. The extracted text appears as
+ random ASCII punctuation/symbols instead of actual CJK characters.
+
+ Detection strategy: if a significant proportion of characters come from
+ subset-embedded fonts and the page produces overwhelmingly ASCII
+ (punctuation, digits, symbols) with virtually no CJK/Hangul/Kana
+ characters, the page is likely garbled due to broken font encoding.
+ """
+ if not page_chars or len(page_chars) < min_chars:
+ return False
+
+ subset_font_count = 0
+ total_non_space = 0
+ ascii_punct_sym = 0
+ cjk_like = 0
+
+ for c in page_chars:
+ text = c.get("text", "")
+ fontname = c.get("fontname", "")
+ if not text or text.isspace():
+ continue
+ total_non_space += 1
+
+ if RAGFlowPdfParser._has_subset_font_prefix(fontname):
+ subset_font_count += 1
+
+ cp = ord(text[0])
+ if (0x2E80 <= cp <= 0x9FFF or 0xF900 <= cp <= 0xFAFF
+ or 0x20000 <= cp <= 0x2FA1F
+ or 0xAC00 <= cp <= 0xD7AF
+ or 0x3040 <= cp <= 0x30FF):
+ cjk_like += 1
+ elif (0x21 <= cp <= 0x2F or 0x3A <= cp <= 0x40
+ or 0x5B <= cp <= 0x60 or 0x7B <= cp <= 0x7E):
+ ascii_punct_sym += 1
+
+ if total_non_space < min_chars:
+ return False
+
+ subset_ratio = subset_font_count / total_non_space
+ if subset_ratio < 0.3:
+ return False
+
+ cjk_ratio = cjk_like / total_non_space
+ punct_ratio = ascii_punct_sym / total_non_space
+ if cjk_ratio < 0.05 and punct_ratio > 0.4:
+ return True
+
+ return False
+
def _evaluate_table_orientation(self, table_img, sample_ratio=0.3):
"""
Evaluate the best rotation orientation for a table image.
@@ -585,7 +702,7 @@ def _insert_ocr_boxes(ocr_results, page_index, table_x0, table_top, insert_at, t
def __ocr(self, pagenum, img, chars, ZM=3, device_id: int | None = None):
start = timer()
bxs = self.ocr.detect(np.array(img), device_id)
- logging.info(f"__ocr detecting boxes of a image cost ({timer() - start}s)")
+ logging.info(f"__ocr detecting boxes of an image cost ({timer() - start}s)")
start = timer()
if not bxs:
@@ -618,14 +735,40 @@ def __ocr(self, pagenum, img, chars, ZM=3, device_id: int | None = None):
if not b["chars"]:
del b["chars"]
continue
- m_ht = np.mean([c["height"] for c in b["chars"]])
- for c in Recognizer.sort_Y_firstly(b["chars"], m_ht):
+ box_chars = b["chars"]
+ m_ht = np.mean([c["height"] for c in box_chars])
+ garbled_count = 0
+ total_count = 0
+ for c in Recognizer.sort_Y_firstly(box_chars, m_ht):
if c["text"] == " " and b["text"]:
if re.match(r"[0-9a-zA-Zа-яА-Я,.?;:!%%]", b["text"][-1]):
b["text"] += " "
else:
b["text"] += c["text"]
+ for ch in c["text"]:
+ if not ch.isspace():
+ total_count += 1
+ if self._is_garbled_char(ch):
+ garbled_count += 1
del b["chars"]
+ # If the majority of characters from pdfplumber are garbled,
+ # clear the text so OCR recognition will be used as fallback.
+ # Strategy 1: PUA / unmapped CID characters
+ if total_count > 0 and garbled_count / total_count >= 0.5:
+ logging.info(
+ "Page %d: detected garbled pdfplumber text (garbled=%d/%d), falling back to OCR for box at (%.1f, %.1f)",
+ pagenum, garbled_count, total_count, b["x0"], b["top"],
+ )
+ b["text"] = ""
+ continue
+ # Strategy 2: font-encoding garbling — all chars are ASCII
+ # punctuation from subset fonts (no CJK output)
+ if total_count > 0 and self._is_garbled_by_font_encoding(box_chars, min_chars=5):
+ logging.info(
+ "Page %d: detected font-encoding garbled text (%d chars), falling back to OCR for box at (%.1f, %.1f)",
+ pagenum, total_count, b["x0"], b["top"],
+ )
+ b["text"] = ""
logging.info(f"__ocr sorting {len(chars)} chars cost {timer() - start}s")
start = timer()
@@ -1400,34 +1543,40 @@ def __images__(self, fnm, zoomin=3, page_from=0, page_to=299, callback=None):
logging.warning(f"Failed to extract characters for pages {page_from}-{page_to}: {str(e)}")
self.page_chars = [[] for _ in range(page_to - page_from)] # If failed to extract, using empty list instead.
+ # Detect garbled pages and clear their chars so the OCR
+ # path will be used instead. Two detection strategies:
+ # 1) PUA / unmapped CID characters (threshold=0.3)
+ # 2) Font-encoding garbling: subset fonts mapping CJK to ASCII
+ for pi, page_ch in enumerate(self.page_chars):
+ if not page_ch:
+ continue
+ # Strategy 1: PUA / CID garbling
+ sample = page_ch if len(page_ch) <= 200 else page_ch[:200]
+ sample_text = "".join(c.get("text", "") for c in sample)
+ if self._is_garbled_text(sample_text, threshold=0.3):
+ logging.warning(
+ "Page %d: pdfplumber extracted mostly garbled characters (%d chars), "
+ "clearing to use OCR fallback.",
+ page_from + pi + 1, len(page_ch),
+ )
+ self.page_chars[pi] = []
+ continue
+ # Strategy 2: font-encoding garbling (CJK mapped to ASCII)
+ if self._is_garbled_by_font_encoding(page_ch):
+ logging.warning(
+ "Page %d: detected font-encoding garbled text "
+ "(subset fonts with no CJK output, %d chars), "
+ "clearing to use OCR fallback.",
+ page_from + pi + 1, len(page_ch),
+ )
+ self.page_chars[pi] = []
+
self.total_page = len(self.pdf.pages)
except Exception as e:
logging.exception(f"RAGFlowPdfParser __images__, exception: {e}")
logging.info(f"__images__ dedupe_chars cost {timer() - start}s")
- self.outlines = []
- try:
- with pdf2_read(fnm if isinstance(fnm, str) else BytesIO(fnm)) as pdf:
- self.pdf = pdf
-
- outlines = self.pdf.outline
-
- def dfs(arr, depth):
- for a in arr:
- if isinstance(a, dict):
- self.outlines.append((a["/Title"], depth))
- continue
- dfs(a, depth + 1)
-
- dfs(outlines, 0)
-
- except Exception as e:
- logging.warning(f"Outlines exception: {e}")
-
- if not self.outlines:
- logging.warning("Miss outlines")
-
logging.debug("Images converted.")
self.is_english = [
re.search(r"[ a-zA-Z0-9,/¸;:'\[\]\(\)!@#$%^&*\"?<>._-]{30,}", "".join(random.choices([c["text"] for c in self.page_chars[i]], k=min(100, len(self.page_chars[i])))))
@@ -1535,6 +1684,7 @@ def __call__(self, fnm, need_image=True, zoomin=3, return_html=False, auto_rotat
if auto_rotate_tables is None:
auto_rotate_tables = os.getenv("TABLE_AUTO_ROTATE", "true").lower() in ("true", "1", "yes")
+ self.outlines = extract_pdf_outlines(fnm)
self.__images__(fnm, zoomin)
self._layouts_rec(zoomin)
self._table_transformer_job(zoomin, auto_rotate=auto_rotate_tables)
@@ -1546,6 +1696,7 @@ def __call__(self, fnm, need_image=True, zoomin=3, return_html=False, auto_rotat
def parse_into_bboxes(self, fnm, callback=None, zoomin=3):
start = timer()
+ self.outlines = extract_pdf_outlines(fnm)
self.__images__(fnm, zoomin, callback=callback)
if callback:
callback(0.40, "OCR finished ({:.2f}s)".format(timer() - start))
@@ -1594,19 +1745,41 @@ def min_rectangle_distance(rect1, rect2):
return math.sqrt(dx * dx + dy * dy) # + (pn2-pn1)*10000
for (img, txt), poss in tbls_or_figs:
- bboxes = [(i, (b["page_number"], b["x0"], b["x1"], b["top"], b["bottom"])) for i, b in enumerate(self.boxes)]
- dists = [
- (min_rectangle_distance((pn, left, right, top + self.page_cum_height[pn], bott + self.page_cum_height[pn]), rect), i) for i, rect in bboxes for pn, left, right, top, bott in poss
- ]
- min_i = np.argmin(dists, axis=0)[0]
- min_i, rect = bboxes[dists[min_i][-1]]
+ # Positions coming from _extract_table_figure carry absolute 0-based page
+ # indices (page_from offset). Convert back to chunk-local indices so we
+ # stay consistent with self.boxes/page_cum_height, which are all relative
+ # to the current parsing window.
+ local_poss = []
+ for pn, left, right, top, bott in poss:
+ local_pn = pn - self.page_from
+ if 0 <= local_pn < len(self.page_cum_height) - 1:
+ local_poss.append((local_pn, left, right, top, bott))
+ else:
+ logging.debug(f"Skip out-of-range table/figure position pn={pn}, page_from={self.page_from}")
+ if not local_poss:
+ logging.debug("No valid local positions for table/figure; skip insertion.")
+ continue
+
if isinstance(txt, list):
txt = "\n".join(txt)
- pn, left, right, top, bott = poss[0]
- if self.boxes[min_i]["bottom"] < top + self.page_cum_height[pn]:
- min_i += 1
+ pn, left, right, top, bott = local_poss[0]
+ insert_at = len(self.boxes)
+ bboxes = [(i, (b["page_number"], b["x0"], b["x1"], b["top"], b["bottom"])) for i, b in enumerate(self.boxes)]
+ if bboxes:
+ dists = [
+ (min_rectangle_distance((cand_pn, cand_left, cand_right, cand_top + self.page_cum_height[cand_pn], cand_bott + self.page_cum_height[cand_pn]), rect), i)
+ for i, rect in bboxes
+ for cand_pn, cand_left, cand_right, cand_top, cand_bott in local_poss
+ ]
+ if dists:
+ nearest_bbox_idx = int(np.argmin([dist for dist, _ in dists]))
+ insert_at, _ = bboxes[dists[nearest_bbox_idx][-1]]
+ if self.boxes[insert_at]["bottom"] < top + self.page_cum_height[pn]:
+ insert_at += 1
+ else:
+ logging.debug("No text boxes available; append %s block directly.", layout_type)
self.boxes.insert(
- min_i,
+ insert_at,
{
"page_number": pn + 1,
"x0": left,
@@ -1771,27 +1944,14 @@ def get_position(self, bx, ZM):
class PlainParser:
def __call__(self, filename, from_page=0, to_page=100000, **kwargs):
- self.outlines = []
lines = []
try:
self.pdf = pdf2_read(filename if isinstance(filename, str) else BytesIO(filename))
for page in self.pdf.pages[from_page:to_page]:
lines.extend([t for t in page.extract_text().split("\n")])
-
- outlines = self.pdf.outline
-
- def dfs(arr, depth):
- for a in arr:
- if isinstance(a, dict):
- self.outlines.append((a["/Title"], depth))
- continue
- dfs(a, depth + 1)
-
- dfs(outlines, 0)
except Exception:
logging.exception("Outlines exception")
- if not self.outlines:
- logging.warning("Miss outlines")
+ self.outlines = extract_pdf_outlines(filename)
return [(line, "") for line in lines], []
diff --git a/deepdoc/parser/resume/entities/corporations.py b/deepdoc/parser/resume/entities/corporations.py
index 0396281deed..50359673032 100644
--- a/deepdoc/parser/resume/entities/corporations.py
+++ b/deepdoc/parser/resume/entities/corporations.py
@@ -29,11 +29,12 @@
).fillna(0)
GOODS["cid"] = GOODS["cid"].astype(str)
GOODS = GOODS.set_index(["cid"])
-CORP_TKS = json.load(
- open(os.path.join(current_file_path, "res/corp.tks.freq.json"), "r",encoding="utf-8")
-)
-GOOD_CORP = json.load(open(os.path.join(current_file_path, "res/good_corp.json"), "r",encoding="utf-8"))
-CORP_TAG = json.load(open(os.path.join(current_file_path, "res/corp_tag.json"), "r",encoding="utf-8"))
+with open(os.path.join(current_file_path, "res/corp.tks.freq.json"), "r", encoding="utf-8") as f:
+ CORP_TKS = json.load(f)
+with open(os.path.join(current_file_path, "res/good_corp.json"), "r", encoding="utf-8") as f:
+ GOOD_CORP = json.load(f)
+with open(os.path.join(current_file_path, "res/corp_tag.json"), "r", encoding="utf-8") as f:
+ CORP_TAG = json.load(f)
def baike(cid, default_v=0):
diff --git a/deepdoc/parser/resume/entities/schools.py b/deepdoc/parser/resume/entities/schools.py
index 4425236beb1..5763ca48be5 100644
--- a/deepdoc/parser/resume/entities/schools.py
+++ b/deepdoc/parser/resume/entities/schools.py
@@ -25,7 +25,8 @@
os.path.join(current_file_path, "res/schools.csv"), sep="\t", header=0
).fillna("")
TBL["name_en"] = TBL["name_en"].map(lambda x: x.lower().strip())
-GOOD_SCH = json.load(open(os.path.join(current_file_path, "res/good_sch.json"), "r",encoding="utf-8"))
+with open(os.path.join(current_file_path, "res/good_sch.json"), "r", encoding="utf-8") as f:
+ GOOD_SCH = json.load(f)
GOOD_SCH = set([re.sub(r"[,. &()()]+", "", c) for c in GOOD_SCH])
diff --git a/deepdoc/parser/tcadp_parser.py b/deepdoc/parser/tcadp_parser.py
index af1c9034895..6a37f0befd0 100644
--- a/deepdoc/parser/tcadp_parser.py
+++ b/deepdoc/parser/tcadp_parser.py
@@ -39,6 +39,7 @@
from common.config_utils import get_base_config
from deepdoc.parser.pdf_parser import RAGFlowPdfParser
+from deepdoc.parser.utils import extract_pdf_outlines
class TencentCloudAPIClient:
@@ -392,6 +393,7 @@ def parse_pdf(
) -> tuple:
"""Parse PDF document"""
+ self.outlines = extract_pdf_outlines(binary if binary else filepath)
temp_file = None
created_tmp_dir = False
diff --git a/deepdoc/parser/txt_parser.py b/deepdoc/parser/txt_parser.py
index 64e200cbc66..6abf8591da8 100644
--- a/deepdoc/parser/txt_parser.py
+++ b/deepdoc/parser/txt_parser.py
@@ -40,7 +40,10 @@ def add_chunk(t):
cks.append(t)
tk_nums.append(tnum)
else:
- cks[-1] += t
+ if cks[-1]:
+ cks[-1] += "\n" + t
+ else:
+ cks[-1] += t
tk_nums[-1] += tnum
dels = []
diff --git a/deepdoc/parser/utils.py b/deepdoc/parser/utils.py
index 85a3554955b..b36af08fa59 100644
--- a/deepdoc/parser/utils.py
+++ b/deepdoc/parser/utils.py
@@ -14,12 +14,16 @@
# limitations under the License.
#
+from io import BytesIO
+
+from pypdf import PdfReader as pdf2_read
+
from rag.nlp import find_codec
def get_text(fnm: str, binary=None) -> str:
txt = ""
- if binary:
+ if binary is not None:
encoding = find_codec(binary)
txt = binary.decode(encoding, errors="ignore")
else:
@@ -30,3 +34,21 @@ def get_text(fnm: str, binary=None) -> str:
break
txt += line
return txt
+
+
+def extract_pdf_outlines(source):
+ try:
+ with pdf2_read(source if isinstance(source, str) else BytesIO(source)) as pdf:
+ outlines = []
+
+ def dfs(nodes, depth):
+ for node in nodes:
+ if isinstance(node, list):
+ dfs(node, depth + 1)
+ else:
+ outlines.append((node["/Title"], depth, pdf.get_destination_page_number(node) + 1))
+
+ dfs(pdf.outline, 0)
+ return outlines
+ except Exception:
+ return []
diff --git a/deepdoc/vision/__init__.py b/deepdoc/vision/__init__.py
index 6b88b792d6b..8d6c6c398a2 100644
--- a/deepdoc/vision/__init__.py
+++ b/deepdoc/vision/__init__.py
@@ -60,9 +60,8 @@ def images_and_outputs(fnm):
pdf_pages(fnm)
return
try:
- fp = open(fnm, "rb")
- binary = fp.read()
- fp.close()
+ with open(fnm, "rb") as fp:
+ binary = fp.read()
images.append(Image.open(io.BytesIO(binary)).convert("RGB"))
outputs.append(os.path.split(fnm)[-1])
except Exception:
diff --git a/deepdoc/vision/layout_recognizer.py b/deepdoc/vision/layout_recognizer.py
index 5b79e2bf5c6..be1f8667cec 100644
--- a/deepdoc/vision/layout_recognizer.py
+++ b/deepdoc/vision/layout_recognizer.py
@@ -17,7 +17,7 @@
import logging
import math
import os
-# import re
+import re
from collections import Counter
from copy import deepcopy
@@ -62,9 +62,8 @@ def __init__(self, domain):
def __call__(self, image_list, ocr_res, scale_factor=3, thr=0.2, batch_size=16, drop=True):
def __is_garbage(b):
- return False
- # patt = [r"^•+$", "^[0-9]{1,2} / ?[0-9]{1,2}$", r"^[0-9]{1,2} of [0-9]{1,2}$", "^http://[^ ]{12,}", "\\(cid *: *[0-9]+ *\\)"]
- # return any([re.search(p, b["text"]) for p in patt])
+ patt = [r"\(cid\s*:\s*\d+\s*\)"]
+ return any([re.search(p, b.get("text", "")) for p in patt])
if self.client:
layouts = self.client.predict(image_list)
diff --git a/deepdoc/vision/ocr.py b/deepdoc/vision/ocr.py
index 1f573bda595..d5e546a3c59 100644
--- a/deepdoc/vision/ocr.py
+++ b/deepdoc/vision/ocr.py
@@ -670,19 +670,13 @@ def detect(self, img, device_id: int | None = None):
if device_id is None:
device_id = 0
- time_dict = {'det': 0, 'rec': 0, 'cls': 0, 'all': 0}
-
if img is None:
- return None, None, time_dict
+ return None
- start = time.time()
- dt_boxes, elapse = self.text_detector[device_id](img)
- time_dict['det'] = elapse
+ dt_boxes, _ = self.text_detector[device_id](img)
if dt_boxes is None:
- end = time.time()
- time_dict['all'] = end - start
- return None, None, time_dict
+ return None
return zip(self.sorted_boxes(dt_boxes), [
("", 0) for _ in range(len(dt_boxes))])
diff --git a/deepdoc/vision/operators.py b/deepdoc/vision/operators.py
index 65d2efa4cb0..43b55ccd3a9 100644
--- a/deepdoc/vision/operators.py
+++ b/deepdoc/vision/operators.py
@@ -22,6 +22,7 @@
import numpy as np
import math
from PIL import Image
+from rag.utils.lazy_image import ensure_pil_image
class DecodeImage:
@@ -128,8 +129,9 @@ def __init__(self, scale=None, mean=None, std=None, order='chw', **kwargs):
def __call__(self, data):
img = data['image']
from PIL import Image
- if isinstance(img, Image.Image):
- img = np.array(img)
+ pil = ensure_pil_image(img)
+ if isinstance(pil, Image.Image):
+ img = np.array(pil)
assert isinstance(img,
np.ndarray), "invalid input 'img' in NormalizeImage"
data['image'] = (
@@ -147,8 +149,9 @@ def __init__(self, **kwargs):
def __call__(self, data):
img = data['image']
from PIL import Image
- if isinstance(img, Image.Image):
- img = np.array(img)
+ pil = ensure_pil_image(img)
+ if isinstance(pil, Image.Image):
+ img = np.array(pil)
data['image'] = img.transpose((2, 0, 1))
return data
diff --git a/deepdoc/vision/table_structure_recognizer.py b/deepdoc/vision/table_structure_recognizer.py
index 0cd762576c1..e0892c2d720 100644
--- a/deepdoc/vision/table_structure_recognizer.py
+++ b/deepdoc/vision/table_structure_recognizer.py
@@ -394,7 +394,7 @@ def __html_table(cap, hdset, tbl):
@staticmethod
def __desc_table(cap, hdr_rowno, tbl, is_english):
- # get text of every colomn in header row to become header text
+ # get text of every column in header row to become header text
clmno = len(tbl[0])
rowno = len(tbl)
headers = {}
diff --git a/docker/.env b/docker/.env
index 7e1bdf801bc..9fdf4e3ea1f 100644
--- a/docker/.env
+++ b/docker/.env
@@ -28,7 +28,7 @@ DEVICE=${DEVICE:-cpu}
COMPOSE_PROFILES=${DOC_ENGINE},${DEVICE}
# The version of Elasticsearch.
-STACK_VERSION=8.11.3
+STACK_VERSION=${STACK_VERSION:-8.11.3}
# The hostname where the Elasticsearch service is exposed
ES_HOST=es01
@@ -118,7 +118,7 @@ MYSQL_DBNAME=rag_flow
MYSQL_PORT=3306
# The port used to expose the MySQL service to the host machine,
# allowing EXTERNAL access to the MySQL database running inside the Docker container.
-EXPOSE_MYSQL_PORT=5455
+EXPOSE_MYSQL_PORT=3306
# The maximum size of communication packets sent to the MySQL server
MYSQL_MAX_PACKET=1073741824
@@ -152,13 +152,18 @@ SVR_WEB_HTTPS_PORT=443
SVR_HTTP_PORT=9380
ADMIN_SVR_HTTP_PORT=9381
SVR_MCP_PORT=9382
+GO_HTTP_PORT=9384
+GO_ADMIN_PORT=9383
+
+# API_PROXY_SCHEME=hybrid # go and python hybrid deploy mode
+API_PROXY_SCHEME=python # use pure python server deployment
# The RAGFlow Docker image to download. v0.22+ doesn't include embedding models.
-RAGFLOW_IMAGE=infiniflow/ragflow:v0.24.0
+RAGFLOW_IMAGE=infiniflow/ragflow:latest
# If you cannot download the RAGFlow Docker image:
-# RAGFLOW_IMAGE=swr.cn-north-4.myhuaweicloud.com/infiniflow/ragflow:v0.24.0
-# RAGFLOW_IMAGE=registry.cn-hangzhou.aliyuncs.com/infiniflow/ragflow:v0.24.0
+# RAGFLOW_IMAGE=swr.cn-north-4.myhuaweicloud.com/infiniflow/ragflow:v0.25.0
+# RAGFLOW_IMAGE=registry.cn-hangzhou.aliyuncs.com/infiniflow/ragflow:v0.25.0
#
# - For the `nightly` edition, uncomment either of the following:
# RAGFLOW_IMAGE=swr.cn-north-4.myhuaweicloud.com/infiniflow/ragflow:nightly
@@ -256,6 +261,10 @@ REGISTER_ENABLED=1
# SANDBOX_ENABLE_SECCOMP=false
# SANDBOX_MAX_MEMORY=256m # b, k, m, g
# SANDBOX_TIMEOUT=10s # s, m, 1m30s
+# The MinIO bucket name for storing sandbox-generated artifacts (charts, files, etc.).
+SANDBOX_ARTIFACT_BUCKET=sandbox-artifacts
+# Number of days before sandbox artifacts are automatically deleted from storage.
+SANDBOX_ARTIFACT_EXPIRE_DAYS=7
# Enable DocLing
USE_DOCLING=false
@@ -276,4 +285,7 @@ DOTNET_SYSTEM_GLOBALIZATION_INVARIANT=1
# Used for ThreadPoolExecutor
-THREAD_POOL_MAX_WORKERS=128
\ No newline at end of file
+THREAD_POOL_MAX_WORKERS=128
+
+#Option to disable login form for SSO
+DISABLE_PASSWORD_LOGIN=false
diff --git a/docker/README.md b/docker/README.md
index c6422bad8c7..b2a9b2fd70e 100644
--- a/docker/README.md
+++ b/docker/README.md
@@ -79,7 +79,7 @@ The [.env](./.env) file contains important environment variables for Docker.
- `SVR_HTTP_PORT`
The port used to expose RAGFlow's HTTP API service to the host machine, allowing **external** access to the service running inside the Docker container. Defaults to `9380`.
- `RAGFLOW-IMAGE`
- The Docker image edition. Defaults to `infiniflow/ragflow:v0.24.0`. The RAGFlow Docker image does not include embedding models.
+ The Docker image edition. Defaults to `infiniflow/ragflow:v0.25.0`. The RAGFlow Docker image does not include embedding models.
> [!TIP]
diff --git a/docker/docker-compose-base.yml b/docker/docker-compose-base.yml
index f82f8027333..1030136bb5e 100644
--- a/docker/docker-compose-base.yml
+++ b/docker/docker-compose-base.yml
@@ -36,7 +36,7 @@ services:
opensearch01:
profiles:
- opensearch
- image: hub.icert.top/opensearchproject/opensearch:2.19.1
+ image: opensearchproject/opensearch:2.19.1
volumes:
- osdata01:/usr/share/opensearch/data
ports:
@@ -72,7 +72,7 @@ services:
infinity:
profiles:
- infinity
- image: infiniflow/infinity:v0.7.0-dev2
+ image: infiniflow/infinity:v0.7.0-dev5
volumes:
- infinity_data:/var/infinity
- ./infinity_conf.toml:/infinity_conf.toml
@@ -202,7 +202,7 @@ services:
restart: unless-stopped
minio:
- image: quay.io/minio/minio:RELEASE.2025-06-13T11-33-47Z
+ image: pgsty/minio:RELEASE.2026-03-25T00-00-00Z
command: ["server", "--console-address", ":9001", "/data"]
ports:
- ${MINIO_PORT}:9000
diff --git a/docker/docker-compose.yml b/docker/docker-compose.yml
index a32c2b609ef..6eba5825d6c 100644
--- a/docker/docker-compose.yml
+++ b/docker/docker-compose.yml
@@ -34,11 +34,13 @@ services:
- ${SVR_HTTP_PORT}:9380
- ${ADMIN_SVR_HTTP_PORT}:9381
- ${SVR_MCP_PORT}:9382 # entry for MCP (host_port:docker_port). The docker_port must match the value you set for `mcp-port` above.
+ - ${GO_HTTP_PORT}:9384
+ - ${GO_ADMIN_PORT}:9383
volumes:
- ./ragflow-logs:/ragflow/logs
- - ./nginx/ragflow.conf:/etc/nginx/conf.d/ragflow.conf
- - ./nginx/proxy.conf:/etc/nginx/proxy.conf
- - ./nginx/nginx.conf:/etc/nginx/nginx.conf
+ # - ./nginx/ragflow.conf:/etc/nginx/conf.d/ragflow.conf
+ # - ./nginx/proxy.conf:/etc/nginx/proxy.conf
+ # - ./nginx/nginx.conf:/etc/nginx/nginx.conf
- ./service_conf.yaml.template:/ragflow/conf/service_conf.yaml.template
- ./entrypoint.sh:/ragflow/entrypoint.sh
env_file: .env
@@ -84,9 +86,9 @@ services:
- ${SVR_MCP_PORT}:9382 # entry for MCP (host_port:docker_port). The docker_port must match the value you set for `mcp-port` above.
volumes:
- ./ragflow-logs:/ragflow/logs
- - ./nginx/ragflow.conf:/etc/nginx/conf.d/ragflow.conf
- - ./nginx/proxy.conf:/etc/nginx/proxy.conf
- - ./nginx/nginx.conf:/etc/nginx/nginx.conf
+ # - ./nginx/ragflow.conf:/etc/nginx/conf.d/ragflow.conf
+ # - ./nginx/proxy.conf:/etc/nginx/proxy.conf
+ # - ./nginx/nginx.conf:/etc/nginx/nginx.conf
- ./service_conf.yaml.template:/ragflow/conf/service_conf.yaml.template
- ./entrypoint.sh:/ragflow/entrypoint.sh
env_file: .env
diff --git a/docker/entrypoint.sh b/docker/entrypoint.sh
index 4fb5cbde3dd..79f77fe43ab 100755
--- a/docker/entrypoint.sh
+++ b/docker/entrypoint.sh
@@ -2,6 +2,9 @@
set -e
+echo "Start RAGFlow cluster, version: "
+cat /ragflow/VERSION
+
# -----------------------------------------------------------------------------
# Usage and command-line argument parsing
# -----------------------------------------------------------------------------
@@ -175,6 +178,27 @@ done < "${TEMPLATE_FILE}"
export LD_LIBRARY_PATH="/usr/lib/x86_64-linux-gnu/"
PY=python3
+# -----------------------------------------------------------------------------
+# Select Nginx Configuration based on API_PROXY_SCHEME
+# -----------------------------------------------------------------------------
+NGINX_CONF_DIR="/etc/nginx/conf.d"
+if [ -n "$API_PROXY_SCHEME" ]; then
+ if [[ "${API_PROXY_SCHEME}" == "hybrid" ]]; then
+ cp -f "$NGINX_CONF_DIR/ragflow.conf.hybrid" "$NGINX_CONF_DIR/ragflow.conf"
+ echo "Applied nginx config: ragflow.conf.hybrid"
+ elif [[ "${API_PROXY_SCHEME}" == "go" ]]; then
+ cp -f "$NGINX_CONF_DIR/ragflow.conf.golang" "$NGINX_CONF_DIR/ragflow.conf"
+ echo "Applied nginx config: ragflow.conf.golang (default)"
+ else
+ cp -f "$NGINX_CONF_DIR/ragflow.conf.python" "$NGINX_CONF_DIR/ragflow.conf"
+ echo "Applied nginx config: ragflow.conf.python"
+ fi
+else
+ # Default to python backend
+ cp -f "$NGINX_CONF_DIR/ragflow.conf.python" "$NGINX_CONF_DIR/ragflow.conf"
+ echo "Default: applied nginx config: ragflow.conf.python"
+fi
+
# -----------------------------------------------------------------------------
# Function(s)
# -----------------------------------------------------------------------------
@@ -212,36 +236,82 @@ function ensure_docling() {
|| uv pip install -i https://pypi.tuna.tsinghua.edu.cn/simple --extra-index-url https://pypi.org/simple --no-cache-dir "docling${DOCLING_PIN}"
}
+function ensure_db_init() {
+ echo "Initializing database tables..."
+ "$PY" -c "from api.db.db_models import init_database_tables as init_web_db; init_web_db()"
+ echo "Database tables initialized."
+}
+
+function wait_for_server() {
+ local url="$1"
+ local server_name="$2"
+ local timeout=90
+ local interval=2
+ local start_time=$(date +%s)
+
+ echo "Waiting for $server_name to be ready at $url..."
+ while ! curl -f -s -o /dev/null "$url"; do
+ if [ $(($(date +%s) - start_time)) -gt $timeout ]; then
+ echo "Timeout waiting for $server_name after $timeout seconds"
+ return 1
+ fi
+ sleep $interval
+ done
+ echo "$server_name is ready."
+}
+
# -----------------------------------------------------------------------------
# Start components based on flags
# -----------------------------------------------------------------------------
ensure_docling
+ensure_db_init
if [[ "${ENABLE_WEBSERVER}" -eq 1 ]]; then
echo "Starting nginx..."
/usr/sbin/nginx
- echo "Starting ragflow_server..."
while true; do
- "$PY" api/ragflow_server.py ${INIT_SUPERUSER_ARGS} &
- wait;
+ echo "Attempt to start RAGFlow server..."
+ "$PY" api/ragflow_server.py ${INIT_SUPERUSER_ARGS}
+ echo "RAGFlow python server started."
sleep 1;
done &
+
+ if [[ "${API_PROXY_SCHEME}" == "hybrid" ]]; then
+ while true; do
+ echo "Attempt to start RAGFlow go server..."
+ wait_for_server "http://127.0.0.1:9380/healthz" "ragflow_server"
+ echo "Starting RAGFlow go server..."
+ bin/server_main
+ sleep 1;
+ done &
+ fi
fi
-if [[ "${ENABLE_DATASYNC}" -eq 1 ]]; then
- echo "Starting data sync..."
+
+if [[ "${ENABLE_ADMIN_SERVER}" -eq 1 ]]; then
while true; do
- "$PY" rag/svr/sync_data_source.py &
- wait;
+ echo "Attempt to start Admin python server..."
+ "$PY" admin/server/admin_server.py
+ echo "Admin python server started"
sleep 1;
done &
+
+ if [[ "${API_PROXY_SCHEME}" == "hybrid" ]]; then
+ while true; do
+ echo "Attempt to starting Admin go server..."
+ wait_for_server "http://127.0.0.1:9381/api/v1/admin/ping" "admin_server"
+ echo "Starting Admin go server..."
+ bin/admin_server
+ sleep 1;
+ done &
+ fi
fi
-if [[ "${ENABLE_ADMIN_SERVER}" -eq 1 ]]; then
- echo "Starting admin_server..."
+if [[ "${ENABLE_DATASYNC}" -eq 1 ]]; then
+ echo "Starting data sync..."
while true; do
- "$PY" admin/server/admin_server.py &
+ "$PY" rag/svr/sync_data_source.py &
wait;
sleep 1;
done &
diff --git a/docker/migration.sh b/docker/migration.sh
index 35adab505e7..b881dbc45e7 100644
--- a/docker/migration.sh
+++ b/docker/migration.sh
@@ -1,8 +1,8 @@
#!/bin/bash
# RAGFlow Data Migration Script
-# Usage: ./migration.sh [backup|restore] [backup_folder]
-#
+# Usage: ./migration.sh [-p project_name] [backup|restore] [backup_folder]
+#
# This script helps you backup and restore RAGFlow Docker volumes
# including MySQL, MinIO, Redis, and Elasticsearch data.
@@ -11,35 +11,55 @@ set -e # Exit on any error
# Default values
DEFAULT_BACKUP_FOLDER="backup"
-VOLUMES=("docker_mysql_data" "docker_minio_data" "docker_redis_data" "docker_esdata01")
+DEFAULT_PROJECT_NAME="docker"
+VOLUME_BASES=("mysql_data" "minio_data" "redis_data" "esdata01")
BACKUP_FILES=("mysql_backup.tar.gz" "minio_backup.tar.gz" "redis_backup.tar.gz" "es_backup.tar.gz")
+# Build volume names from project name and base names
+build_volume_names() {
+ VOLUMES=()
+ for base in "${VOLUME_BASES[@]}"; do
+ VOLUMES+=("${PROJECT_NAME}_${base}")
+ done
+}
+
# Function to display help information
show_help() {
echo "RAGFlow Data Migration Tool"
echo ""
echo "USAGE:"
- echo " $0 [backup_folder]"
+ echo " $0 [-p project_name] [backup_folder]"
echo ""
echo "OPERATIONS:"
echo " backup - Create backup of all RAGFlow data volumes"
echo " restore - Restore RAGFlow data volumes from backup"
echo " help - Show this help message"
echo ""
+ echo "OPTIONS:"
+ echo " -p project_name - Docker Compose project name (default: '$DEFAULT_PROJECT_NAME')"
+ echo " Use this when you started RAGFlow with 'docker compose -p '"
+ echo ""
echo "PARAMETERS:"
- echo " backup_folder - Name of backup folder (default: '$DEFAULT_BACKUP_FOLDER')"
+ echo " backup_folder - Name of backup folder (default: '$DEFAULT_BACKUP_FOLDER')"
echo ""
echo "EXAMPLES:"
- echo " $0 backup # Backup to './backup' folder"
- echo " $0 backup my_backup # Backup to './my_backup' folder"
- echo " $0 restore # Restore from './backup' folder"
- echo " $0 restore my_backup # Restore from './my_backup' folder"
+ echo " $0 backup # Backup with default project name 'docker'"
+ echo " $0 backup my_backup # Backup to './my_backup' folder"
+ echo " $0 restore # Restore from './backup' folder"
+ echo " $0 restore my_backup # Restore from './my_backup' folder"
+ echo " $0 -p ragflow backup # Backup volumes for project 'ragflow'"
+ echo " $0 -p ragflow restore my_backup # Restore volumes for project 'ragflow'"
+ echo ""
+ echo "DOCKER VOLUMES (with default project name '$DEFAULT_PROJECT_NAME'):"
+ echo " - ${DEFAULT_PROJECT_NAME}_mysql_data (MySQL database)"
+ echo " - ${DEFAULT_PROJECT_NAME}_minio_data (MinIO object storage)"
+ echo " - ${DEFAULT_PROJECT_NAME}_redis_data (Redis cache)"
+ echo " - ${DEFAULT_PROJECT_NAME}_esdata01 (Elasticsearch indices)"
echo ""
- echo "DOCKER VOLUMES:"
- echo " - docker_mysql_data (MySQL database)"
- echo " - docker_minio_data (MinIO object storage)"
- echo " - docker_redis_data (Redis cache)"
- echo " - docker_esdata01 (Elasticsearch indices)"
+ echo "NOTE:"
+ echo " If you started RAGFlow with 'docker compose -p myproject up', the volume"
+ echo " names will be prefixed with 'myproject' instead of 'docker'. In that case,"
+ echo " use '-p myproject' with this script to match the correct volumes."
}
# Function to check if Docker is running
@@ -60,23 +80,23 @@ volume_exists() {
# Function to check if any containers are using the target volumes
check_containers_using_volumes() {
echo "🔍 Checking for running containers that might be using target volumes..."
-
+
# Get all running containers
local running_containers=$(docker ps --format "{{.Names}}")
-
+
if [ -z "$running_containers" ]; then
echo "✅ No running containers found"
return 0
fi
-
+
# Check each running container for volume usage
local containers_using_volumes=()
local volume_usage_details=()
-
+
for container in $running_containers; do
# Get container's mount information
local mounts=$(docker inspect "$container" --format '{{range .Mounts}}{{.Source}}{{"|"}}{{end}}' 2>/dev/null || echo "")
-
+
# Check if any of our target volumes are used by this container
for volume in "${VOLUMES[@]}"; do
if echo "$mounts" | grep -q "$volume"; then
@@ -86,7 +106,7 @@ check_containers_using_volumes() {
fi
done
done
-
+
# If any containers are using our volumes, show error and exit
if [ ${#containers_using_volumes[@]} -gt 0 ]; then
echo ""
@@ -100,15 +120,19 @@ check_containers_using_volumes() {
echo " - $detail"
done
echo ""
- echo "🛑 SOLUTION: Stop the containers before performing backup/restore operations:"
- echo " docker-compose -f docker/.yml down"
+ if [ "$PROJECT_NAME" = "$DEFAULT_PROJECT_NAME" ]; then
+ echo "🛑 SOLUTION: Stop the containers before performing backup/restore operations:"
+ echo " docker compose -f docker/.yml down"
+ else
+ echo "🛑 SOLUTION: Stop the containers before performing backup/restore operations:"
+ echo " docker compose -p $PROJECT_NAME -f docker/.yml down"
+ fi
echo ""
- echo "💡 After backup/restore, you can restart with:"
- echo " docker-compose -f docker/.yml up -d"
+ echo "💡 After backup/restore, you can restart with the corresponding 'up -d' command."
echo ""
exit 1
fi
-
+
echo "✅ No containers are using target volumes, safe to proceed"
return 0
}
@@ -127,25 +151,28 @@ confirm_action() {
# Function to perform backup
perform_backup() {
local backup_folder=$1
-
+
echo "🚀 Starting RAGFlow data backup..."
echo "📁 Backup folder: $backup_folder"
+ echo "🏷️ Project name: $PROJECT_NAME"
echo ""
-
+
# Check if any containers are using the volumes
check_containers_using_volumes
-
+
# Create backup folder if it doesn't exist
mkdir -p "$backup_folder"
-
+
+ local total=${#VOLUMES[@]}
+
# Backup each volume
for i in "${!VOLUMES[@]}"; do
local volume="${VOLUMES[$i]}"
local backup_file="${BACKUP_FILES[$i]}"
local step=$((i + 1))
-
- echo "📦 Step $step/4: Backing up $volume..."
-
+
+ echo "📦 Step $step/$total: Backing up $volume..."
+
if volume_exists "$volume"; then
docker run --rm \
-v "$volume":/source \
@@ -157,10 +184,10 @@ perform_backup() {
fi
echo ""
done
-
+
echo "🎉 Backup completed successfully!"
echo "📍 Backup location: $(pwd)/$backup_folder"
-
+
# List backup files with sizes
echo ""
echo "📋 Backup files created:"
@@ -175,20 +202,21 @@ perform_backup() {
# Function to perform restore
perform_restore() {
local backup_folder=$1
-
+
echo "🔄 Starting RAGFlow data restore..."
echo "📁 Backup folder: $backup_folder"
+ echo "🏷️ Project name: $PROJECT_NAME"
echo ""
-
+
# Check if any containers are using the volumes
check_containers_using_volumes
-
+
# Check if backup folder exists
if [ ! -d "$backup_folder" ]; then
echo "❌ Error: Backup folder '$backup_folder' does not exist"
exit 1
fi
-
+
# Check if all backup files exist
local missing_files=()
for backup_file in "${BACKUP_FILES[@]}"; do
@@ -196,7 +224,7 @@ perform_restore() {
missing_files+=("$backup_file")
fi
done
-
+
if [ ${#missing_files[@]} -gt 0 ]; then
echo "❌ Error: Missing backup files:"
for file in "${missing_files[@]}"; do
@@ -205,7 +233,7 @@ perform_restore() {
echo "Please ensure all backup files are present in '$backup_folder'"
exit 1
fi
-
+
# Check for existing volumes and warn user
local existing_volumes=()
for volume in "${VOLUMES[@]}"; do
@@ -213,7 +241,7 @@ perform_restore() {
existing_volumes+=("$volume")
fi
done
-
+
if [ ${#existing_volumes[@]} -gt 0 ]; then
echo "⚠️ WARNING: The following Docker volumes already exist:"
for volume in "${existing_volumes[@]}"; do
@@ -222,23 +250,25 @@ perform_restore() {
echo ""
echo "🔴 IMPORTANT: Restoring will OVERWRITE existing data!"
echo "💡 Recommendation: Create a backup of your current data first:"
- echo " $0 backup current_backup_$(date +%Y%m%d_%H%M%S)"
+ echo " $0 -p $PROJECT_NAME backup current_backup_$(date +%Y%m%d_%H%M%S)"
echo ""
-
+
if ! confirm_action "Do you want to continue with the restore operation?"; then
echo "❌ Restore operation cancelled by user"
exit 0
fi
fi
-
+
+ local total=${#VOLUMES[@]}
+
# Create volumes and restore data
for i in "${!VOLUMES[@]}"; do
local volume="${VOLUMES[$i]}"
local backup_file="${BACKUP_FILES[$i]}"
local step=$((i + 1))
-
- echo "🔧 Step $step/4: Restoring $volume..."
-
+
+ echo "🔧 Step $step/$total: Restoring $volume..."
+
# Create volume if it doesn't exist
if ! volume_exists "$volume"; then
echo " 📋 Creating Docker volume: $volume"
@@ -246,18 +276,18 @@ perform_restore() {
else
echo " 📋 Using existing Docker volume: $volume"
fi
-
+
# Restore data
echo " 📥 Restoring data from $backup_file..."
docker run --rm \
-v "$volume":/target \
-v "$(pwd)/$backup_folder":/backup \
alpine tar xzf "/backup/$backup_file" -C /target
-
+
echo "✅ Successfully restored $volume"
echo ""
done
-
+
echo "🎉 Restore completed successfully!"
echo "💡 You can now start your RAGFlow services"
}
@@ -266,17 +296,38 @@ perform_restore() {
main() {
# Check if Docker is available
check_docker
-
- # Parse command line arguments
+
+ # Parse -p flag
+ PROJECT_NAME="$DEFAULT_PROJECT_NAME"
+ while [ $# -gt 0 ]; do
+ case "$1" in
+ -p)
+ if [ -z "${2:-}" ]; then
+ echo "❌ Error: -p requires a project name argument"
+ exit 1
+ fi
+ PROJECT_NAME="$2"
+ shift 2
+ ;;
+ *)
+ break
+ ;;
+ esac
+ done
+
+ # Build volume names based on project name
+ build_volume_names
+
+ # Parse remaining positional arguments
local operation=${1:-}
local backup_folder=${2:-$DEFAULT_BACKUP_FOLDER}
-
+
# Handle help or no arguments
if [ -z "$operation" ] || [ "$operation" = "help" ] || [ "$operation" = "-h" ] || [ "$operation" = "--help" ]; then
show_help
exit 0
fi
-
+
# Validate operation
case "$operation" in
backup)
@@ -295,4 +346,4 @@ main() {
}
# Run main function with all arguments
-main "$@"
\ No newline at end of file
+main "$@"
diff --git a/docker/nginx/ragflow.conf.golang b/docker/nginx/ragflow.conf.golang
new file mode 100644
index 00000000000..d5c9bb12924
--- /dev/null
+++ b/docker/nginx/ragflow.conf.golang
@@ -0,0 +1,33 @@
+server {
+ listen 80;
+ server_name _;
+ root /ragflow/web/dist;
+
+ gzip on;
+ gzip_min_length 1k;
+ gzip_comp_level 9;
+ gzip_types text/plain application/javascript application/x-javascript text/css application/xml text/javascript application/x-httpd-php image/jpeg image/gif image/png;
+ gzip_vary on;
+ gzip_disable "MSIE [1-6]\.";
+
+ location ~ ^/api/v1/admin {
+ proxy_pass http://127.0.0.1:9383;
+ include proxy.conf;
+ }
+
+ location ~ ^/(v1|api) {
+ proxy_pass http://127.0.0.1:9382;
+ include proxy.conf;
+ }
+
+ location / {
+ index index.html;
+ try_files $uri $uri/ /index.html;
+ }
+
+ # Cache-Control: max-age Expires
+ location ~ ^/static/(css|js|media)/ {
+ expires 10y;
+ access_log off;
+ }
+}
diff --git a/docker/nginx/ragflow.conf.hybrid b/docker/nginx/ragflow.conf.hybrid
new file mode 100644
index 00000000000..0fc5f508083
--- /dev/null
+++ b/docker/nginx/ragflow.conf.hybrid
@@ -0,0 +1,68 @@
+server {
+ listen 80;
+ server_name _;
+ root /ragflow/web/dist;
+
+ gzip on;
+ gzip_min_length 1k;
+ gzip_comp_level 9;
+ gzip_types text/plain application/javascript application/x-javascript text/css application/xml text/javascript application/x-httpd-php image/jpeg image/gif image/png;
+ gzip_vary on;
+ gzip_disable "MSIE [1-6]\.";
+
+ location ~ ^/api/v1/admin/roles_with_permission {
+ proxy_pass http://127.0.0.1:9381;
+ include proxy.conf;
+ }
+
+ location ~ ^/api/v1/admin/sandbox {
+ proxy_pass http://127.0.0.1:9381;
+ include proxy.conf;
+ }
+
+ location ~ ^/api/v1/admin/roles {
+ proxy_pass http://127.0.0.1:9381;
+ include proxy.conf;
+ }
+
+ location ~ ^/api/v1/admin/whitelist {
+ proxy_pass http://127.0.0.1:9381;
+ include proxy.conf;
+ }
+
+ location ~ ^/api/v1/admin/variables {
+ proxy_pass http://127.0.0.1:9381;
+ include proxy.conf;
+ }
+
+ location ~ ^/api/v1/admin {
+ proxy_pass http://127.0.0.1:9383;
+ include proxy.conf;
+ }
+
+ location ~ ^/v1/system/config {
+ proxy_pass http://127.0.0.1:9384;
+ include proxy.conf;
+ }
+
+ location ~ ^/v1/user/(login|logout) {
+ proxy_pass http://127.0.0.1:9384;
+ include proxy.conf;
+ }
+
+ location ~ ^/(v1|api) {
+ proxy_pass http://127.0.0.1:9380;
+ include proxy.conf;
+ }
+
+ location / {
+ index index.html;
+ try_files $uri $uri/ /index.html;
+ }
+
+ # Cache-Control: max-age Expires
+ location ~ ^/static/(css|js|media)/ {
+ expires 10y;
+ access_log off;
+ }
+}
\ No newline at end of file
diff --git a/docker/nginx/ragflow.conf b/docker/nginx/ragflow.conf.python
similarity index 94%
rename from docker/nginx/ragflow.conf
rename to docker/nginx/ragflow.conf.python
index a0609827624..d000e41519a 100644
--- a/docker/nginx/ragflow.conf
+++ b/docker/nginx/ragflow.conf.python
@@ -26,7 +26,7 @@ server {
try_files $uri $uri/ /index.html;
}
- # Cache-Control: max-age~@~AExpires
+ # Cache-Control: max-age Expires
location ~ ^/static/(css|js|media)/ {
expires 10y;
access_log off;
diff --git a/docker/service_conf.yaml.template b/docker/service_conf.yaml.template
index f283f08530e..a06e71f9e7f 100644
--- a/docker/service_conf.yaml.template
+++ b/docker/service_conf.yaml.template
@@ -19,6 +19,10 @@ minio:
host: '${MINIO_HOST:-minio}:9000'
bucket: '${MINIO_BUCKET:-}'
prefix_path: '${MINIO_PREFIX_PATH:-}'
+ # optional: set to true for HTTPS (SSL/TLS). Used by MinIO client and health check.
+ # secure: ${MINIO_SECURE:-false}
+ # optional: set to false to allow self-signed certificates (e.g. in development).
+ # verify: ${MINIO_VERIFY:-true}
es:
hosts: 'http://${ES_HOST:-es01}:9200'
username: '${ES_USER:-elastic}'
@@ -94,6 +98,7 @@ user_default_llm:
# secret: 'secret'
# tenant_id: 'tenant_id'
# container_name: 'container_name'
+# cloud: 'public' # Azure cloud: 'public', 'china', 'government', or 'germany'
# The OSS object storage uses the MySQL configuration above by default. If you need to switch to another object storage service, please uncomment and configure the following parameters.
# opendal:
# scheme: 'mysql' # Storage type, such as s3, oss, azure, etc.
@@ -139,12 +144,13 @@ user_default_llm:
# client_secret: "your_client_secret"
# redirect_uri: "https://your-app.com/v1/user/oauth/callback/github"
# authentication:
-# client:
-# switch: false
-# http_app_key:
-# http_secret_key:
-# site:
-# switch: false
+# client:
+# switch: false
+# http_app_key:
+# http_secret_key:
+# site:
+# switch: false
+# disable_password_login: false
# permission:
# switch: false
# component: false
diff --git a/docs/administrator/_category_.json b/docs/administrator/_category_.json
new file mode 100644
index 00000000000..e00d9a56ab2
--- /dev/null
+++ b/docs/administrator/_category_.json
@@ -0,0 +1,11 @@
+{
+ "label": "Administrator guides",
+ "position": 4,
+ "link": {
+ "type": "generated-index",
+ "description": "Guides for system administrtors"
+ },
+ "customProps": {
+ "sidebarIcon": "LucideComputer"
+ }
+}
diff --git a/docs/guides/admin/_category_.json b/docs/administrator/admin/_category_.json
similarity index 53%
rename from docs/guides/admin/_category_.json
rename to docs/administrator/admin/_category_.json
index fa6d832fc8d..c05f2e48006 100644
--- a/docs/guides/admin/_category_.json
+++ b/docs/administrator/admin/_category_.json
@@ -1,9 +1,9 @@
{
- "label": "Administration",
- "position": 6,
+ "label": "Admin service",
+ "position": 3,
"link": {
"type": "generated-index",
- "description": "RAGFlow administration"
+ "description": "RAGFlow administration service"
},
"customProps": {
"categoryIcon": "LucideUserCog"
diff --git a/docs/guides/admin/admin_service.md b/docs/administrator/admin/admin_service.md
similarity index 100%
rename from docs/guides/admin/admin_service.md
rename to docs/administrator/admin/admin_service.md
diff --git a/docs/guides/admin/admin_ui.md b/docs/administrator/admin/admin_ui.md
similarity index 99%
rename from docs/guides/admin/admin_ui.md
rename to docs/administrator/admin/admin_ui.md
index 9584bb8cfc7..ae90bb97381 100644
--- a/docs/guides/admin/admin_ui.md
+++ b/docs/administrator/admin/admin_ui.md
@@ -32,7 +32,6 @@ The service status page displays of all services within the RAGFlow system.
- **Extra Info**: Display additional configuration information of a service in a dialog.
- **Service Details**: Display detailed status information of a service in a dialog. According to service's type, a service's status information could be displayed as a plain text, a key-value data list, a data table or a bar chart.
-
### User management
The user management page provides comprehensive tools for managing all users in the RAGFlow system.
diff --git a/docs/guides/admin/ragflow_cli.md b/docs/administrator/admin/ragflow_cli.md
similarity index 99%
rename from docs/guides/admin/ragflow_cli.md
rename to docs/administrator/admin/ragflow_cli.md
index f682d6be64d..a6ed02f0061 100644
--- a/docs/guides/admin/ragflow_cli.md
+++ b/docs/administrator/admin/ragflow_cli.md
@@ -16,7 +16,7 @@ The RAGFlow CLI is a command-line-based system administration tool that offers a
2. Install ragflow-cli.
```bash
- pip install ragflow-cli==0.24.0
+ pip install ragflow-cli==0.25.0
```
3. Launch the CLI client:
@@ -439,7 +439,7 @@ show_version
+-----------------------+
| version |
+-----------------------+
-| v0.24.0-24-g6f60e9f9e |
+| v0.25.0-24-g6f60e9f9e |
+-----------------------+
```
diff --git a/docs/administrator/backup_and_migration.md b/docs/administrator/backup_and_migration.md
new file mode 100644
index 00000000000..8a55691b68e
--- /dev/null
+++ b/docs/administrator/backup_and_migration.md
@@ -0,0 +1,313 @@
+---
+sidebar_position: 2
+slug: /migration
+sidebar_custom_props: {
+ categoryIcon: LucideLocateFixed
+}
+---
+
+# Backup & migration
+
+- [Data migration](#data-migration)
+- [Migrate from multi-bucket to single-bucket mode](#migrate-from-multi-bucket-to-single-bucket-mode)
+
+## Data migration
+
+:::info KUDOS
+This document is contributed by our community contributor [TreeDy](https://github.com/Treedy2020). We may not actively maintain this document.
+:::
+
+A common scenario is processing large datasets on a powerful instance (e.g., with a GPU) and then migrating the entire RAGFlow service to a different production environment (e.g., a CPU-only server). This guide explains how to safely back up and restore your data using our provided migration script.
+
+### Identify your data
+
+By default, RAGFlow uses Docker volumes to store all persistent data, including your database, uploaded files, and search indexes. You can see these volumes by running:
+
+```bash
+docker volume ls
+```
+
+The output will look similar to this:
+
+```text
+DRIVER VOLUME NAME
+local docker_esdata01
+local docker_minio_data
+local docker_mysql_data
+local docker_redis_data
+```
+
+These volumes contain all the data you need to migrate.
+
+:::note
+The volume name prefix (e.g., `docker_`) comes from the Docker Compose project name. By default it is `docker` (derived from the directory name). If you started RAGFlow with `docker compose -p `, your volumes will be prefixed with `_` instead, for example `ragflow_mysql_data`.
+:::
+
+### Step 1: Stop RAGFlow services
+
+Before starting the migration, you must stop all running RAGFlow services on the **source machine**. Navigate to the project's root directory and run:
+
+```bash
+docker compose -f docker/docker-compose.yml down
+```
+
+If you started RAGFlow with a custom project name (e.g., `docker compose -p ragflow`), include it in the command:
+
+```bash
+docker compose -p ragflow -f docker/docker-compose.yml down
+```
+
+**Important:** Do **not** use the `-v` flag (e.g., `docker compose down -v`), as this will delete all your data volumes. The migration script includes a check and will prevent you from running it if services are active.
+
+### Step 2: Back up your data
+
+We provide a convenient script to package all your data volumes into a single backup folder.
+
+For a quick reference of the script's commands and options, you can run:
+```bash
+bash docker/migration.sh help
+```
+
+To create a backup, run the following command from the project's root directory:
+
+```bash
+bash docker/migration.sh backup
+```
+
+This will create a `backup/` folder in your project root containing compressed archives of your data volumes.
+
+You can also specify a custom name for your backup folder:
+
+```bash
+bash docker/migration.sh backup my_ragflow_backup
+```
+
+This will create a folder named `my_ragflow_backup/` instead.
+
+If you started RAGFlow with a custom project name (e.g., `docker compose -p ragflow`), use the `-p` flag so the script can find the correct volumes:
+
+```bash
+bash docker/migration.sh -p ragflow backup
+bash docker/migration.sh -p ragflow backup my_ragflow_backup
+```
+
+### Step 3: Transfer the backup folder
+
+Copy the entire backup folder (e.g., `backup/` or `my_ragflow_backup/`) from your source machine to the RAGFlow project directory on your **target machine**. You can use tools like `scp`, `rsync`, or a physical drive for the transfer.
+
+### Step 4: Restore your data
+
+On the **target machine**, ensure that RAGFlow services are not running. Then, use the migration script to restore your data from the backup folder.
+
+If your backup folder is named `backup/`, run:
+
+```bash
+bash docker/migration.sh restore
+```
+
+If you used a custom name, specify it in the command:
+
+```bash
+bash docker/migration.sh restore my_ragflow_backup
+```
+
+If the target machine uses a custom project name, use the `-p` flag to ensure the volumes are created with the correct prefix:
+
+```bash
+bash docker/migration.sh -p ragflow restore
+bash docker/migration.sh -p ragflow restore my_ragflow_backup
+```
+
+The script will automatically create the necessary Docker volumes and unpack the data.
+
+**Note:** If the script detects that Docker volumes with the same names already exist on the target machine, it will warn you that restoring will overwrite the existing data and ask for confirmation before proceeding.
+
+### Step 5: Start RAGFlow services
+
+Once the restore process is complete, you can start the RAGFlow services on your new machine:
+
+```bash
+docker compose -f docker/docker-compose.yml up -d
+```
+
+If you use a custom project name:
+
+```bash
+docker compose -p ragflow -f docker/docker-compose.yml up -d
+```
+
+**Note:** If you already have built a service by docker compose before, you may need to backup your data for target machine like this guide above and run like:
+
+```bash
+# Please backup by `bash docker/migration.sh backup backup_dir_name` before you do the following line.
+# !!! this line -v flag will delete the original docker volume
+docker compose -f docker/docker-compose.yml down -v
+docker compose -f docker/docker-compose.yml up -d
+```
+
+Your RAGFlow instance is now running with all the data from your original machine.
+
+## Migrate from multi-bucket to single-bucket mode
+
+:::info KUDOS
+This document is contributed by our community contributor [arogan178](https://github.com/arogan178). We may not actively maintain this document.
+:::
+
+By default, RAGFlow creates one bucket per Knowledge Base (dataset) and one bucket per user folder. This can be problematic when:
+
+- Your cloud provider charges per bucket
+- Your IAM policy restricts bucket creation
+- You want all data organized in a single bucket with directory structure
+
+The **Single Bucket Mode** allows you to configure RAGFlow to use a single bucket with a directory structure instead of multiple buckets.
+
+### How it works
+
+#### Default mode (Multiple buckets)
+
+```
+bucket: kb_12345/
+ └── document_1.pdf
+bucket: kb_67890/
+ └── document_2.pdf
+bucket: folder_abc/
+ └── file_3.txt
+```
+
+#### Single bucket mode (with prefix_path)
+
+```
+bucket: ragflow-bucket/
+ └── ragflow/
+ ├── kb_12345/
+ │ └── document_1.pdf
+ ├── kb_67890/
+ │ └── document_2.pdf
+ └── folder_abc/
+ └── file_3.txt
+```
+
+### Configuration
+
+#### MinIO configuration
+
+Edit your `service_conf.yaml` or set environment variables:
+
+```yaml
+minio:
+ user: "your-access-key"
+ password: "your-secret-key"
+ host: "minio.example.com:443"
+ bucket: "ragflow-bucket" # Default bucket name
+ prefix_path: "ragflow" # Optional prefix path
+```
+
+Or using environment variables:
+
+```bash
+export MINIO_USER=your-access-key
+export MINIO_PASSWORD=your-secret-key
+export MINIO_HOST=minio.example.com:443
+export MINIO_BUCKET=ragflow-bucket
+export MINIO_PREFIX_PATH=ragflow
+```
+
+#### S3 configuration (already supported)
+
+```yaml
+s3:
+ access_key: "your-access-key"
+ secret_key: "your-secret-key"
+ endpoint_url: "https://s3.amazonaws.com"
+ bucket: "my-ragflow-bucket"
+ prefix_path: "production"
+ region: "us-east-1"
+```
+
+### IAM policy example
+
+When using single bucket mode, you only need permissions for one bucket:
+
+```json
+{
+ "Version": "2012-10-17",
+ "Statement": [
+ {
+ "Effect": "Allow",
+ "Action": ["s3:*"],
+ "Resource": [
+ "arn:aws:s3:::ragflow-bucket",
+ "arn:aws:s3:::ragflow-bucket/*"
+ ]
+ }
+ ]
+}
+```
+
+### Migration from multi-bucket to single bucket
+
+If you're migrating from multi-bucket mode to single-bucket mode:
+
+1. **Set environment variables** for the new configuration
+2. **Restart RAGFlow** services
+3. **Migrate existing data** (optional):
+
+```bash
+# Example using mc (MinIO Client)
+mc alias set old-minio http://old-minio:9000 ACCESS_KEY SECRET_KEY
+mc alias set new-minio https://new-minio:443 ACCESS_KEY SECRET_KEY
+
+# List all knowledge base buckets
+mc ls old-minio/ | grep kb_ | while read -r line; do
+ bucket=$(echo $line | awk '{print $5}')
+ # Copy each bucket to the new structure
+ mc cp --recursive old-minio/$bucket/ new-minio/ragflow-bucket/ragflow/$bucket/
+done
+```
+
+### Toggle between modes
+
+#### Enable single bucket mode
+
+```yaml
+minio:
+ bucket: "my-single-bucket"
+ prefix_path: "ragflow"
+```
+
+#### Disable (Use multi-bucket mode)
+
+```yaml
+minio:
+ # Leave bucket and prefix_path empty or commented out
+ # bucket: ''
+ # prefix_path: ''
+```
+
+### Troubleshooting
+
+#### Issue: Access Denied errors
+
+**Solution**: Ensure your IAM policy grants access to the bucket specified in the configuration.
+
+#### Issue: Files not found after switching modes
+
+**Solution**: The path structure changes between modes. You'll need to migrate existing data.
+
+#### Issue: Connection fails with HTTPS
+
+**Solution**: Ensure `secure: True` is set in the MinIO connection (automatically handled for port 443).
+
+### Storage backends supported
+
+- ✅ **MinIO** - Full support with single bucket mode
+- ✅ **AWS S3** - Full support with single bucket mode
+- ✅ **Alibaba OSS** - Full support with single bucket mode
+- ✅ **Azure Blob** - Uses container-based structure (different paradigm)
+- ⚠️ **OpenDAL** - Depends on underlying storage backend
+
+### Performance considerations
+
+- **Single bucket mode** may have slightly better performance for bucket listing operations
+- **Multi-bucket mode** provides better isolation and organization for large deployments
+- Choose based on your specific requirements and infrastructure constraints
diff --git a/docs/configurations.md b/docs/administrator/configurations.md
similarity index 98%
rename from docs/configurations.md
rename to docs/administrator/configurations.md
index 2b274c8e9b2..ec13939e3dc 100644
--- a/docs/configurations.md
+++ b/docs/administrator/configurations.md
@@ -1,8 +1,8 @@
---
-sidebar_position: 1
+sidebar_position: 0
slug: /configurations
sidebar_custom_props: {
- sidebarIcon: LucideCog
+ categoryIcon: LucideCog
}
---
# Configuration
@@ -103,7 +103,7 @@ RAGFlow utilizes MinIO as its object storage solution, leveraging its scalabilit
- `SVR_HTTP_PORT`
The port used to expose RAGFlow's HTTP API service to the host machine, allowing **external** access to the service running inside the Docker container. Defaults to `9380`.
- `RAGFLOW-IMAGE`
- The Docker image edition. Defaults to `infiniflow/ragflow:v0.24.0` (the RAGFlow Docker image without embedding models).
+ The Docker image edition. Defaults to `infiniflow/ragflow:v0.25.0` (the RAGFlow Docker image without embedding models).
:::tip NOTE
If you cannot download the RAGFlow Docker image, try the following mirrors.
@@ -111,7 +111,7 @@ If you cannot download the RAGFlow Docker image, try the following mirrors.
- For the `nightly` edition:
- `RAGFLOW_IMAGE=swr.cn-north-4.myhuaweicloud.com/infiniflow/ragflow:nightly` or,
- `RAGFLOW_IMAGE=registry.cn-hangzhou.aliyuncs.com/infiniflow/ragflow:nightly`.
-:::
+ :::
### Embedding service
diff --git a/docs/guides/tracing.mdx b/docs/administrator/tracing.mdx
similarity index 99%
rename from docs/guides/tracing.mdx
rename to docs/administrator/tracing.mdx
index 13cf99874b8..c9b4221e145 100644
--- a/docs/guides/tracing.mdx
+++ b/docs/administrator/tracing.mdx
@@ -1,5 +1,5 @@
---
-sidebar_position: 9
+sidebar_position: 5
slug: /tracing
sidebar_custom_props: {
categoryIcon: LucideLocateFixed
diff --git a/docs/guides/upgrade_ragflow.mdx b/docs/administrator/upgrade_ragflow.mdx
similarity index 92%
rename from docs/guides/upgrade_ragflow.mdx
rename to docs/administrator/upgrade_ragflow.mdx
index ef43384ddce..1fe5245eeed 100644
--- a/docs/guides/upgrade_ragflow.mdx
+++ b/docs/administrator/upgrade_ragflow.mdx
@@ -1,5 +1,5 @@
---
-sidebar_position: 11
+sidebar_position: 1
slug: /upgrade_ragflow
sidebar_custom_props: {
categoryIcon: LucideArrowBigUpDash
@@ -62,16 +62,16 @@ To upgrade RAGFlow, you must upgrade **both** your code **and** your Docker imag
git pull
```
-3. Switch to the latest, officially published release, e.g., `v0.24.0`:
+3. Switch to the latest, officially published release, e.g., `v0.25.0`:
```bash
- git checkout -f v0.24.0
+ git checkout -f v0.25.0
```
4. Update **ragflow/docker/.env**:
```bash
- RAGFLOW_IMAGE=infiniflow/ragflow:v0.24.0
+ RAGFLOW_IMAGE=infiniflow/ragflow:v0.25.0
```
5. Update the RAGFlow image and restart RAGFlow:
@@ -92,10 +92,10 @@ No, you do not need to. Upgrading RAGFlow in itself will *not* remove your uploa
1. From an environment with Internet access, pull the required Docker image.
2. Save the Docker image to a **.tar** file.
```bash
- docker save -o ragflow.v0.24.0.tar infiniflow/ragflow:v0.24.0
+ docker save -o ragflow.v0.25.0.tar infiniflow/ragflow:v0.25.0
```
3. Copy the **.tar** file to the target server.
4. Load the **.tar** file into Docker:
```bash
- docker load -i ragflow.v0.24.0.tar
+ docker load -i ragflow.v0.25.0.tar
```
diff --git a/docs/contribution/_category_.json b/docs/contribution/_category_.json
deleted file mode 100644
index a9bd348a8cc..00000000000
--- a/docs/contribution/_category_.json
+++ /dev/null
@@ -1,11 +0,0 @@
-{
- "label": "Contribution",
- "position": 8,
- "link": {
- "type": "generated-index",
- "description": "Miscellaneous contribution guides."
- },
- "customProps": {
- "sidebarIcon": "LucideHandshake"
- }
-}
diff --git a/docs/develop/_category_.json b/docs/develop/_category_.json
index c80693175f7..406b847f968 100644
--- a/docs/develop/_category_.json
+++ b/docs/develop/_category_.json
@@ -1,6 +1,6 @@
{
- "label": "Developers",
- "position": 4,
+ "label": "Developer guides",
+ "position": 5,
"link": {
"type": "generated-index",
"description": "Guides for hardcore developers"
diff --git a/docs/develop/build_docker_image.mdx b/docs/develop/build_docker_image.mdx
index 6cb2dede439..7e8462813c7 100644
--- a/docs/develop/build_docker_image.mdx
+++ b/docs/develop/build_docker_image.mdx
@@ -1,5 +1,5 @@
---
-sidebar_position: 1
+sidebar_position: 4
slug: /build_docker_image
sidebar_custom_props: {
categoryIcon: LucidePackage
@@ -49,7 +49,7 @@ After building the infiniflow/ragflow:nightly image, you are ready to launch a f
1. Edit Docker Compose Configuration
-Open the `docker/.env` file. Find the `RAGFLOW_IMAGE` setting and change the image reference from `infiniflow/ragflow:v0.24.0` to `infiniflow/ragflow:nightly` to use the pre-built image.
+Open the `docker/.env` file. Find the `RAGFLOW_IMAGE` setting and change the image reference from `infiniflow/ragflow:v0.25.0` to `infiniflow/ragflow:nightly` to use the pre-built image.
2. Launch the Service
diff --git a/docs/contribution/contributing.md b/docs/develop/contributing.md
similarity index 99%
rename from docs/contribution/contributing.md
rename to docs/develop/contributing.md
index 39b5e1a5503..e3f910672c6 100644
--- a/docs/contribution/contributing.md
+++ b/docs/develop/contributing.md
@@ -1,5 +1,5 @@
---
-sidebar_position: 1
+sidebar_position: 20
slug: /contributing
sidebar_custom_props: {
categoryIcon: LucideBookA
diff --git a/docs/develop/launch_ragflow_from_source.md b/docs/develop/launch_ragflow_from_source.md
index c193e2be373..22f127f34c2 100644
--- a/docs/develop/launch_ragflow_from_source.md
+++ b/docs/develop/launch_ragflow_from_source.md
@@ -1,5 +1,5 @@
---
-sidebar_position: 2
+sidebar_position: 3
slug: /launch_ragflow_from_source
sidebar_custom_props: {
categoryIcon: LucideMonitorPlay
@@ -90,7 +90,7 @@ docker compose -f docker/docker-compose-base.yml up -d
```
3. **Optional:** If you cannot access HuggingFace, set the HF_ENDPOINT environment variable to use a mirror site:
-
+
```bash
export HF_ENDPOINT=https://hf-mirror.com
```
diff --git a/docs/develop/mcp/_category_.json b/docs/develop/mcp/_category_.json
index eb7b1444aa9..aa67c45a958 100644
--- a/docs/develop/mcp/_category_.json
+++ b/docs/develop/mcp/_category_.json
@@ -1,6 +1,6 @@
{
"label": "MCP",
- "position": 40,
+ "position": 2,
"link": {
"type": "generated-index",
"description": "Guides and references on accessing RAGFlow's datasets via MCP."
diff --git a/docs/develop/mcp/launch_mcp_server.md b/docs/develop/mcp/launch_mcp_server.md
index 72a23aca19e..99633fd3238 100644
--- a/docs/develop/mcp/launch_mcp_server.md
+++ b/docs/develop/mcp/launch_mcp_server.md
@@ -196,7 +196,7 @@ docker logs docker-ragflow-cpu-1
## Security considerations
-As MCP technology is still at early stage and no official best practices for authentication or authorization have been established, RAGFlow currently uses [API key](./acquire_ragflow_api_key.md) to validate identity for the operations described earlier. However, in public environments, this makeshift solution could expose your MCP server to potential network attacks. Therefore, when running a local SSE server, it is recommended to bind only to localhost (`127.0.0.1`) rather than to all interfaces (`0.0.0.0`).
+As MCP technology is still at early stage and no official best practices for authentication or authorization have been established, RAGFlow currently uses [API key](../acquire_ragflow_api_key.md) to validate identity for the operations described earlier. However, in public environments, this makeshift solution could expose your MCP server to potential network attacks. Therefore, when running a local SSE server, it is recommended to bind only to localhost (`127.0.0.1`) rather than to all interfaces (`0.0.0.0`).
For further guidance, see the [official MCP documentation](https://modelcontextprotocol.io/docs/concepts/transports#security-considerations).
diff --git a/docs/develop/migrate_to_single_bucket_mode.md b/docs/develop/migrate_to_single_bucket_mode.md
deleted file mode 100644
index de7c8fe873b..00000000000
--- a/docs/develop/migrate_to_single_bucket_mode.md
+++ /dev/null
@@ -1,169 +0,0 @@
----
-sidebar_position: 20
-slug: /migrate_to_single_bucket_mode
----
-
-# Migrate from multi-Bucket to single-bucket mode
-
-By default, RAGFlow creates one bucket per Knowledge Base (dataset) and one bucket per user folder. This can be problematic when:
-
-- Your cloud provider charges per bucket
-- Your IAM policy restricts bucket creation
-- You want all data organized in a single bucket with directory structure
-
-The **Single Bucket Mode** allows you to configure RAGFlow to use a single bucket with a directory structure instead of multiple buckets.
-
-:::info KUDOS
-This document is contributed by our community contributor [arogan178](https://github.com/arogan178). We may not actively maintain this document.
-:::
-
-## How It Works
-
-### Default Mode (Multiple Buckets)
-
-```
-bucket: kb_12345/
- └── document_1.pdf
-bucket: kb_67890/
- └── document_2.pdf
-bucket: folder_abc/
- └── file_3.txt
-```
-
-### Single Bucket Mode (with prefix_path)
-
-```
-bucket: ragflow-bucket/
- └── ragflow/
- ├── kb_12345/
- │ └── document_1.pdf
- ├── kb_67890/
- │ └── document_2.pdf
- └── folder_abc/
- └── file_3.txt
-```
-
-## Configuration
-
-### MinIO Configuration
-
-Edit your `service_conf.yaml` or set environment variables:
-
-```yaml
-minio:
- user: "your-access-key"
- password: "your-secret-key"
- host: "minio.example.com:443"
- bucket: "ragflow-bucket" # Default bucket name
- prefix_path: "ragflow" # Optional prefix path
-```
-
-Or using environment variables:
-
-```bash
-export MINIO_USER=your-access-key
-export MINIO_PASSWORD=your-secret-key
-export MINIO_HOST=minio.example.com:443
-export MINIO_BUCKET=ragflow-bucket
-export MINIO_PREFIX_PATH=ragflow
-```
-
-### S3 Configuration (already supported)
-
-```yaml
-s3:
- access_key: "your-access-key"
- secret_key: "your-secret-key"
- endpoint_url: "https://s3.amazonaws.com"
- bucket: "my-ragflow-bucket"
- prefix_path: "production"
- region: "us-east-1"
-```
-
-## IAM Policy Example
-
-When using single bucket mode, you only need permissions for one bucket:
-
-```json
-{
- "Version": "2012-10-17",
- "Statement": [
- {
- "Effect": "Allow",
- "Action": ["s3:*"],
- "Resource": [
- "arn:aws:s3:::ragflow-bucket",
- "arn:aws:s3:::ragflow-bucket/*"
- ]
- }
- ]
-}
-```
-
-## Migration from Multi-Bucket to Single Bucket
-
-If you're migrating from multi-bucket mode to single-bucket mode:
-
-1. **Set environment variables** for the new configuration
-2. **Restart RAGFlow** services
-3. **Migrate existing data** (optional):
-
-```bash
-# Example using mc (MinIO Client)
-mc alias set old-minio http://old-minio:9000 ACCESS_KEY SECRET_KEY
-mc alias set new-minio https://new-minio:443 ACCESS_KEY SECRET_KEY
-
-# List all knowledge base buckets
-mc ls old-minio/ | grep kb_ | while read -r line; do
- bucket=$(echo $line | awk '{print $5}')
- # Copy each bucket to the new structure
- mc cp --recursive old-minio/$bucket/ new-minio/ragflow-bucket/ragflow/$bucket/
-done
-```
-
-## Toggle Between Modes
-
-### Enable Single Bucket Mode
-
-```yaml
-minio:
- bucket: "my-single-bucket"
- prefix_path: "ragflow"
-```
-
-### Disable (Use Multi-Bucket Mode)
-
-```yaml
-minio:
- # Leave bucket and prefix_path empty or commented out
- # bucket: ''
- # prefix_path: ''
-```
-
-## Troubleshooting
-
-### Issue: Access Denied errors
-
-**Solution**: Ensure your IAM policy grants access to the bucket specified in the configuration.
-
-### Issue: Files not found after switching modes
-
-**Solution**: The path structure changes between modes. You'll need to migrate existing data.
-
-### Issue: Connection fails with HTTPS
-
-**Solution**: Ensure `secure: True` is set in the MinIO connection (automatically handled for port 443).
-
-## Storage Backends Supported
-
-- ✅ **MinIO** - Full support with single bucket mode
-- ✅ **AWS S3** - Full support with single bucket mode
-- ✅ **Alibaba OSS** - Full support with single bucket mode
-- ✅ **Azure Blob** - Uses container-based structure (different paradigm)
-- ⚠️ **OpenDAL** - Depends on underlying storage backend
-
-## Performance Considerations
-
-- **Single bucket mode** may have slightly better performance for bucket listing operations
-- **Multi-bucket mode** provides better isolation and organization for large deployments
-- Choose based on your specific requirements and infrastructure constraints
diff --git a/docs/faq.mdx b/docs/faq.mdx
index cc7ab374b57..e52ff1cda03 100644
--- a/docs/faq.mdx
+++ b/docs/faq.mdx
@@ -1,5 +1,5 @@
---
-sidebar_position: 10
+sidebar_position: 20
slug: /faq
sidebar_custom_props: {
sidebarIcon: LucideCircleQuestionMark
@@ -68,11 +68,11 @@ These limitations led us to develop [Infinity](https://github.com/infiniflow/inf
---
-### Differences between demo.ragflow.io and a locally deployed open-source RAGFlow service?
+### Differences between cloud.ragflow.io and a locally deployed open-source RAGFlow service?
-demo.ragflow.io demonstrates the capabilities of RAGFlow Enterprise. Its DeepDoc models are pre-trained using proprietary data and it offers much more sophisticated team permission controls. Essentially, demo.ragflow.io serves as a preview of RAGFlow's forthcoming SaaS (Software as a Service) offering.
+cloud.ragflow.io demonstrates the capabilities of RAGFlow Enterprise. Its DeepDoc models are pre-trained using proprietary data and it offers much more sophisticated team permission controls. Essentially, cloud.ragflow.io serves as a preview of RAGFlow's forthcoming SaaS (Software as a Service) offering.
-You can deploy an open-source RAGFlow service and call it from a Python client or through RESTful APIs. However, this is not supported on demo.ragflow.io.
+You can deploy an open-source RAGFlow service and call it from a Python client or through RESTful APIs. However, this is not supported on cloud.ragflow.io.
---
@@ -182,6 +182,12 @@ To fix this issue, use https://hf-mirror.com instead:
---
+### `Fail to access model(Ollama/xxxxx)`
+
+Ollama may time out or fail during its first model load due to memory constraints or out-of-memory (OOM). It is best to test your local model in isolation first. If sharing hardware with other services, memory exhaustion is likely. To resolve this, switch to a smaller model or increase RAM.
+
+---
+
### `MaxRetryError: HTTPSConnectionPool(host='hf-mirror.com', port=443)`
This error suggests that you do not have Internet access or are unable to connect to hf-mirror.com. Try the following:
@@ -195,6 +201,20 @@ This error suggests that you do not have Internet access or are unable to connec
---
+### `RuntimeError: Unable to start Tika server.`
+
+This error is almost always caused by Java not being installed or not accessible in the environment. See [here](https://github.com/infiniflow/ragflow/issues/13194) for detailed instructions.
+
+---
+
+### `Cannot stat '/etc/nginx/conf.d/ragflow.conf.python': No such file or directory`
+
+To resolve this, either download the missing file from the corresponding tag on [GitHub](https://github.com/infiniflow/ragflow) or update `~/ragflow/docker/docker-compose.yml` as follows:
+
+
+
+---
+
### `WARNING: can't find /ragflow/rag/res/borker.tm`
Ignore this warning and continue. All system warnings can be ignored.
@@ -349,7 +369,7 @@ Your IP address or port number may be incorrect. If you are using the default co
A correct Ollama IP address and port is crucial to adding models to Ollama:
-- If you are on demo.ragflow.io, ensure that the server hosting Ollama has a publicly accessible IP address. Note that 127.0.0.1 is not a publicly accessible IP address.
+- If you are on cloud.ragflow.io, ensure that the server hosting Ollama has a publicly accessible IP address. Note that 127.0.0.1 is not a publicly accessible IP address.
- If you deploy RAGFlow locally, ensure that Ollama and RAGFlow are in the same LAN and can communicate with each other.
See [Deploy a local LLM](./guides/models/deploy_local_llm.mdx) for more information.
@@ -445,7 +465,7 @@ See [Acquire a RAGFlow API key](./develop/acquire_ragflow_api_key.md).
### How to upgrade RAGFlow?
-See [Upgrade RAGFlow](./guides/upgrade_ragflow.mdx) for more information.
+See [Upgrade RAGFlow](./administrator/upgrade_ragflow.mdx) for more information.
---
@@ -567,6 +587,24 @@ RAGFlow supports MinerU's `vlm-http-client` backend, enabling you to delegate do
When using the `vlm-http-client` backend, the RAGFlow server requires no GPU, only network connectivity. This enables cost-effective distributed deployment with multiple RAGFlow instances sharing one remote vLLM server.
:::
+### How to use an external Docling Serve server for document parsing?
+
+RAGFlow supports Docling in two modes:
+
+1. **Local Docling** (existing mode): install Docling in the RAGFlow runtime (`USE_DOCLING=true`) and parse in-process.
+2. **External Docling Serve** (remote mode): point RAGFlow to a Docling Serve endpoint.
+
+To enable remote mode, set:
+
+```bash
+DOCLING_SERVER_URL=http://your-docling-serve-host:5001
+```
+
+Behavior:
+
+- When `DOCLING_SERVER_URL` is set, RAGFlow sends PDFs to Docling Serve using `/v1/convert/source` (and falls back to `/v1alpha/convert/source` for older servers).
+- When `DOCLING_SERVER_URL` is not set, RAGFlow uses local in-process Docling.
+
### How to use PaddleOCR for document parsing?
From v0.24.0 onwards, RAGFlow includes PaddleOCR as an optional PDF parser. Please note that RAGFlow acts only as a *remote client* for PaddleOCR, calling the PaddleOCR API to parse PDFs and reading the returned files.
diff --git a/docs/guides/_category_.json b/docs/guides/_category_.json
index 18f4890a985..a5551f8b8b9 100644
--- a/docs/guides/_category_.json
+++ b/docs/guides/_category_.json
@@ -1,9 +1,9 @@
{
- "label": "Guides",
+ "label": "User guides",
"position": 3,
"link": {
"type": "generated-index",
- "description": "Guides for RAGFlow users and developers."
+ "description": "Guides for RAGFlow users."
},
"customProps": {
"sidebarIcon": "LucideBookMarked"
diff --git a/docs/guides/agent/agent_component_reference/code.mdx b/docs/guides/agent/agent_component_reference/code.mdx
index a9472ca5e03..d0af92cc184 100644
--- a/docs/guides/agent/agent_component_reference/code.mdx
+++ b/docs/guides/agent/agent_component_reference/code.mdx
@@ -23,7 +23,7 @@ We use gVisor to isolate code execution from the host system. Please follow [the
### 2. Ensure Sandbox is properly installed
-RAGFlow Sandbox is a secure, pluggable code execution backend. It serves as the code executor for the **Code** component. Please follow the [instructions here](https://github.com/infiniflow/ragflow/tree/main/sandbox) to install RAGFlow Sandbox.
+RAGFlow Sandbox is a secure, pluggable code execution backend. It serves as the code executor for the **Code** component. Please follow the [instructions here](https://github.com/infiniflow/ragflow/tree/main/agent/sandbox) to install RAGFlow Sandbox.
:::note Docker client version
The executor manager image now bundles Docker CLI `29.1.0` (API 1.44+). Older images shipped Docker 24.x and will fail against newer Docker daemons with `client version 1.43 is too old`. Pull the latest `infiniflow/sandbox-executor-manager:latest` or rebuild it in `./sandbox/executor_manager` if you encounter this error.
diff --git a/docs/guides/agent/agent_component_reference/docs_generator.md b/docs/guides/agent/agent_component_reference/docs_generator.md
deleted file mode 100644
index 3ed8e342af7..00000000000
--- a/docs/guides/agent/agent_component_reference/docs_generator.md
+++ /dev/null
@@ -1,241 +0,0 @@
----
-sidebar_position: 35
-slug: /docs_generator
----
-
-# Docs Generator component
-
-A component that generates downloadable PDF, DOCX, or TXT documents from markdown-style content with full Unicode support.
-
----
-
-The **Docs Generator** component enables you to create professional documents directly within your agent workflow. It accepts markdown-formatted text and converts it into downloadable files, making it ideal for generating reports, summaries, or any structured document output.
-
-## Key features
-
-- **Multiple output formats**: PDF, DOCX, and TXT
-- **Full Unicode support**: Automatic font switching for CJK (Chinese, Japanese, Korean), Arabic, Hebrew, and other non-Latin scripts
-- **Rich formatting**: Headers, lists, tables, code blocks, and more
-- **Customizable styling**: Fonts, margins, page size, and orientation
-- **Document extras**: Logo, watermark, page numbers, and timestamps
-- **Direct download**: Generates a download button for the chat interface
-
-## Prerequisites
-
-- Content to be converted into a document (typically from an **Agent** or other text-generating component).
-
-## Examples
-
-You can pair an **Agent** component with the **Docs Generator** to create dynamic documents based on user queries. The **Agent** generates the content, and the **Docs Generator** converts it into a downloadable file. Connect the output to a **Message** component to display the download button in the chat.
-
-A typical workflow looks like:
-
-```
-Begin → Agent → Docs Generator → Message
-```
-
-In the **Message** component, reference the `download` output variable from the **Docs Generator** to display a download button in the chat interface.
-
-## Configurations
-
-### Content
-
-The main text content to include in the document. Supports Markdown formatting:
-
-- **Bold**: `**text**` or `__text__`
-- **Italic**: `*text*` or `_text_`
-- **Inline code**: `` `code` ``
-- **Headings**: `# Heading 1`, `## Heading 2`, `### Heading 3`
-- **Bullet lists**: `- item` or `* item`
-- **Numbered lists**: `1. item`
-- **Tables**: `| Column 1 | Column 2 |`
-- **Horizontal lines**: `---`
-- **Code blocks**: ` ``` code ``` `
-
-:::tip NOTE
-Click **(x)** or type `/` to insert variables from upstream components.
-:::
-
-### Title
-
-Optional. The document title displayed at the top of the generated file.
-
-### Subtitle
-
-Optional. A subtitle displayed below the title.
-
-### Output format
-
-The file format for the generated document:
-
-- **PDF** (default): Portable Document Format with full styling support.
-- **DOCX**: Microsoft Word format.
-- **TXT**: Plain text format.
-
-### Logo image
-
-Optional. A logo image to display at the top of the document. You can either:
-
-- Upload an image file using the file picker
-- Paste an image path, URL, or base64-encoded data
-
-### Logo position
-
-The horizontal position of the logo:
-
-- **left** (default)
-- **center**
-- **right**
-
-### Logo dimensions
-
-- **Logo width**: Width in inches (default: `2.0`)
-- **Logo height**: Height in inches (default: `1.0`)
-
-### Font family
-
-The font used throughout the document:
-
-- **Helvetica** (default)
-- **Times-Roman**
-- **Courier**
-- **Helvetica-Bold**
-- **Times-Bold**
-
-### Font size
-
-The base font size in points. Defaults to `12`.
-
-### Title font size
-
-The font size for the document title. Defaults to `24`.
-
-### Page size
-
-The paper size for the document:
-
-- **A4** (default)
-- **Letter**
-
-### Orientation
-
-The page orientation:
-
-- **Portrait** (default)
-- **Landscape**
-
-### Margins
-
-Page margins in inches:
-
-- **Margin top**: Defaults to `1.0`
-- **Margin bottom**: Defaults to `1.0`
-- **Margin left**: Defaults to `1.0`
-- **Margin right**: Defaults to `1.0`
-
-### Filename
-
-Optional. Custom filename for the generated document. If left empty, a filename is auto-generated with a timestamp.
-
-### Output directory
-
-The server directory where generated documents are saved. Defaults to `/tmp/pdf_outputs`.
-
-### Add page numbers
-
-When enabled, page numbers are added to the footer of each page. Defaults to `true`.
-
-### Add timestamp
-
-When enabled, a generation timestamp is added to the document footer. Defaults to `true`.
-
-### Watermark text
-
-Optional. Text to display as a diagonal watermark across each page. Useful for marking documents as "Draft", "Confidential", etc.
-
-## Output
-
-The **Docs Generator** component provides the following output variables:
-
-| Variable name | Type | Description |
-|---------------|-----------|--------------------------------------------------------------|
-| `file_path` | `string` | The server path where the generated document is saved. |
-| `pdf_base64` | `string` | The document content encoded in base64 format. |
-| `download` | `string` | JSON containing download information for the chat interface. |
-| `success` | `boolean` | Indicates whether the document was generated successfully. |
-
-### Displaying the download button
-
-To display a download button in the chat, add a **Message** component after the **Docs Generator** and reference the `download` variable:
-
-1. Connect the **Docs Generator** output to a **Message** component.
-2. In the **Message** component's content field, type `/` and select `{Docs Generator_0@download}`.
-3. When the agent runs, a download button will appear in the chat, allowing users to download the generated document.
-
-The download button automatically handles:
-- File type detection (PDF, DOCX, TXT)
-- Proper MIME type for browser downloads
-- Base64 decoding for direct file delivery
-
-## Unicode and multi-language support
-
-The **Docs Generator** includes intelligent font handling for international content:
-
-### How it works
-
-1. **Content analysis**: The component scans the text for non-Latin characters.
-2. **Automatic font switching**: When CJK or other complex scripts are detected, the system automatically switches to a compatible CID font (STSong-Light for Chinese, HeiseiMin-W3 for Japanese, HYSMyeongJo-Medium for Korean).
-3. **Latin content**: For documents containing only Latin characters (including extended Latin, Cyrillic, and Greek), the user-selected font family is used.
-
-### Supported scripts
-
-| Script | Unicode Range | Font Used |
-|------------------------------|---------------|--------------------|
-| Chinese (CJK) | U+4E00–U+9FFF | STSong-Light |
-| Japanese (Hiragana/Katakana) | U+3040–U+30FF | HeiseiMin-W3 |
-| Korean (Hangul) | U+AC00–U+D7AF | HYSMyeongJo-Medium |
-| Arabic | U+0600–U+06FF | CID font fallback |
-| Hebrew | U+0590–U+05FF | CID font fallback |
-| Devanagari (Hindi) | U+0900–U+097F | CID font fallback |
-| Thai | U+0E00–U+0E7F | CID font fallback |
-
-### Font installation
-
-For full multi-language support in self-hosted deployments, ensure Unicode fonts are installed:
-
-**Linux (Debian/Ubuntu):**
-```bash
-apt-get install fonts-freefont-ttf fonts-noto-cjk
-```
-
-**Docker:** The official RAGFlow Docker image includes these fonts. For custom images, add the font packages to your Dockerfile:
-```dockerfile
-RUN apt-get update && apt-get install -y fonts-freefont-ttf fonts-noto-cjk
-```
-
-:::tip NOTE
-CID fonts (STSong-Light, HeiseiMin-W3, etc.) are built into ReportLab and do not require additional installation. They are used automatically when CJK content is detected.
-:::
-
-## Troubleshooting
-
-### Characters appear as boxes or question marks
-
-This indicates missing font support. Ensure:
-1. The content contains supported Unicode characters.
-2. For self-hosted deployments, Unicode fonts are installed on the server.
-3. The document is being viewed in a PDF reader that supports embedded fonts.
-
-### Download button not appearing
-
-Ensure:
-1. The **Message** component is connected after the **Docs Generator**.
-2. The `download` variable is correctly referenced using `/` (which appears as `{Docs Generator_0@download}` when copied).
-3. The document generation completed successfully (check `success` output).
-
-### Large tables not rendering correctly
-
-For tables with many columns or large cell content:
-- The component automatically converts wide tables to a definition list format for better readability.
-- Consider splitting large tables into multiple smaller tables.
-- Use landscape orientation for wide tables.
diff --git a/docs/guides/agent/agent_component_reference/execute_sql.md b/docs/guides/agent/agent_component_reference/execute_sql.md
index 30c9c9912fa..c08b73ffbbe 100644
--- a/docs/guides/agent/agent_component_reference/execute_sql.md
+++ b/docs/guides/agent/agent_component_reference/execute_sql.md
@@ -24,7 +24,7 @@ The **Execute SQL** tool enables you to connect to a relational database and run
## Examples
-You can pair an **Agent** component with the **Execute SQL** tool, with the **Agent** generating SQL statements and the **Execute SQL** tool handling database connection and query execution. An example of this setup can be found in the **SQL Assistant** Agent template shown below:
+You can pair an **Agent** component with the **Execute SQL** tool, with the **Agent** generating SQL statements and the **Execute SQL** tool handling database connection and query execution. An example of this setup can be found in the **Text-to-SQL data expert** Agent template shown below:
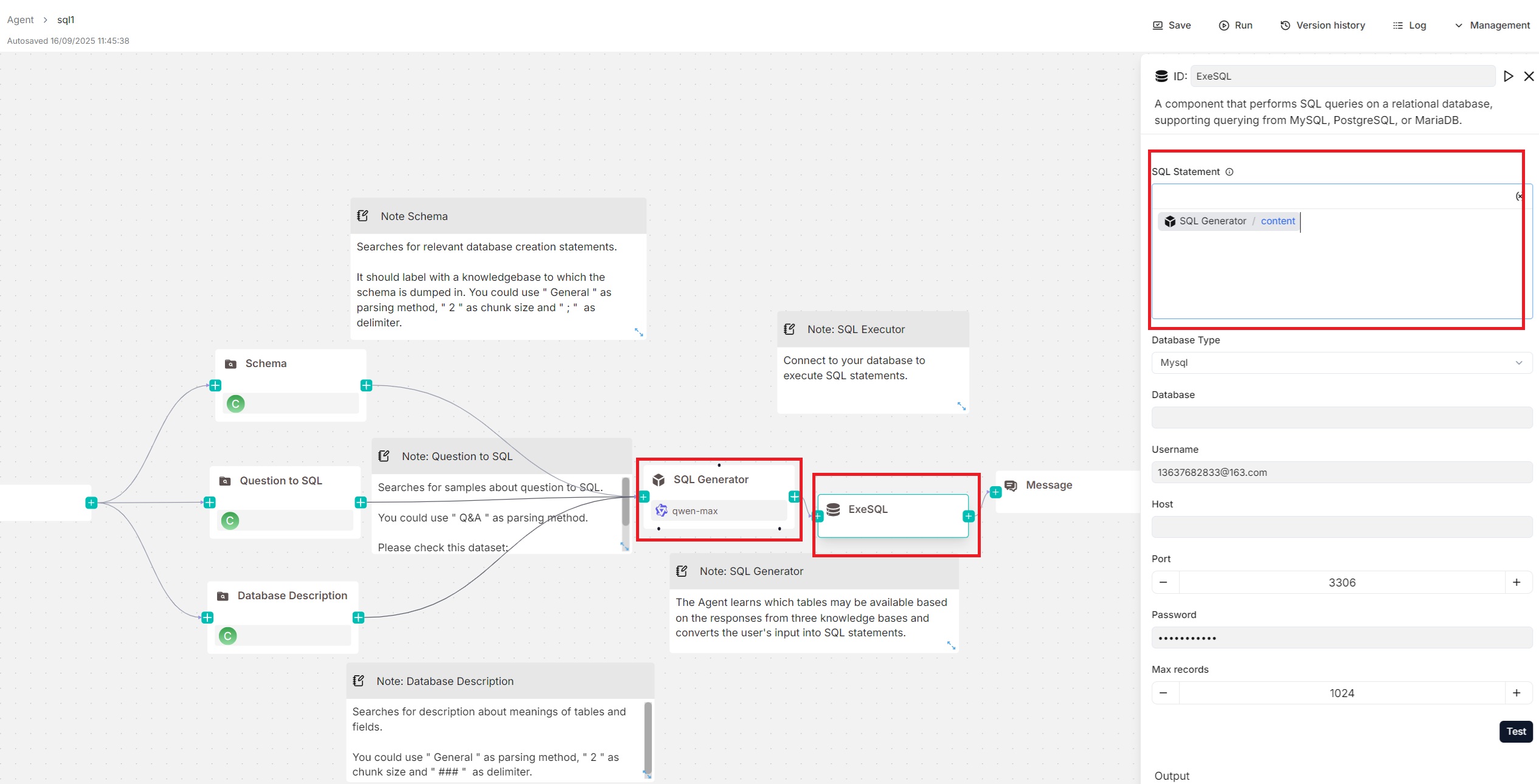
diff --git a/docs/guides/agent/agent_component_reference/parser.md b/docs/guides/agent/agent_component_reference/parser.md
index cdc0a9e1750..75b6341cb23 100644
--- a/docs/guides/agent/agent_component_reference/parser.md
+++ b/docs/guides/agent/agent_component_reference/parser.md
@@ -65,6 +65,12 @@ Starting from v0.22.0, RAGFlow includes MinerU (≥ 2.6.3) as an optional PDF p
- If you decide to use a chunking method from the **Built-in** dropdown, ensure it supports PDF parsing, then select **MinerU** from the **PDF parser** dropdown.
- If you use a custom ingestion pipeline instead, select **MinerU** in the **PDF parser** section of the **Parser** component.
+To use an external Docling Serve instance (instead of local in-process Docling), set:
+
+- `DOCLING_SERVER_URL`: The Docling Serve API endpoint (for example, `http://docling-host:5001`).
+
+When `DOCLING_SERVER_URL` is set, RAGFlow sends PDF content to Docling Serve (`/v1/convert/source`, with fallback to `/v1alpha/convert/source`) and ingests the returned markdown/text. If the variable is not set, RAGFlow keeps using local Docling (`USE_DOCLING=true` + installed package) behavior.
+
:::note
All MinerU environment variables are optional. When set, these values are used to auto-provision a MinerU OCR model for the tenant on first use. To avoid auto-provisioning, skip the environment variable settings and only configure MinerU from the **Model providers** page in the UI.
:::
diff --git a/docs/guides/agent/agent_component_reference/retrieval.mdx b/docs/guides/agent/agent_component_reference/retrieval.mdx
index 5295092ed1d..2cf791d4d8f 100644
--- a/docs/guides/agent/agent_component_reference/retrieval.mdx
+++ b/docs/guides/agent/agent_component_reference/retrieval.mdx
@@ -128,7 +128,7 @@ Select one or more languages for cross‑language search. If no language is sele
### Use knowledge graph
:::caution IMPORTANT
-Before enabling this feature, ensure you have properly [constructed a knowledge graph from each target dataset](../../dataset/construct_knowledge_graph.md).
+Before enabling this feature, ensure you have properly [constructed a knowledge graph from each target dataset](../../dataset/advanced/construct_knowledge_graph.md).
:::
Whether to use knowledge graph(s) in the specified dataset(s) during retrieval for multi-hop question answering. When enabled, this would involve iterative searches across entity, relationship, and community report chunks, greatly increasing retrieval time.
diff --git a/docs/guides/agent/agent_introduction.md b/docs/guides/agent/agent_introduction.md
index f310e503ddf..8e05ab7889f 100644
--- a/docs/guides/agent/agent_introduction.md
+++ b/docs/guides/agent/agent_introduction.md
@@ -43,7 +43,7 @@ We also provide templates catered to different business scenarios. You can eithe
-2. To create an agent from scratch, click **Create Agent**. Alternatively, to create an agent from one of our templates, click the desired card, such as **Deep Research**, name your agent in the pop-up dialogue, and click **OK** to confirm.
+2. To create an agent from scratch, click **Create Agent**. Alternatively, to create an agent from one of our templates, click the desired card, such as **Deep research**, name your agent in the pop-up dialogue, and click **OK** to confirm.
*You are now taken to the **no-code workflow editor** page.*
diff --git a/docs/guides/agent/embed_agent_into_webpage.md b/docs/guides/agent/embed_agent_into_webpage.md
index 97dae8b66c0..4676443e16e 100644
--- a/docs/guides/agent/embed_agent_into_webpage.md
+++ b/docs/guides/agent/embed_agent_into_webpage.md
@@ -11,7 +11,14 @@ You can use iframe to embed an agent into a third-party webpage.
1. Before proceeding, you must [acquire an API key](../models/llm_api_key_setup.md); otherwise, an error message would appear.
2. On the **Agent** page, click an intended agent to access its editing page.
-3. Click **Management > Embed into webpage** on the top right corner of the canvas to show the **iframe** window:
-4. Copy the iframe and embed it into your webpage.
+3. Click **Management > Embed into webpage** on the top right corner of the canvas to show the **Embed into webpage** dialog.
+4. Configure your embed options:
+ - **Embed Type**: Choose between Fullscreen Chat (traditional iframe) or Floating Widget (Intercom-style)
+ - **Theme**: Select Light or Dark theme (for fullscreen mode)
+ - **Hide avatar**: Toggle avatar visibility
+ - **Enable Streaming Responses**: Enable streaming for widget mode
+ - **Locale**: Select the language for the embedded agent
+5. Copy the generated iframe code and embed it into your webpage.
+6. **Chat in new tab**: Click the "Chat in new tab" button to preview the agent in a separate browser tab with your configured settings. This allows you to test the agent before embedding it.
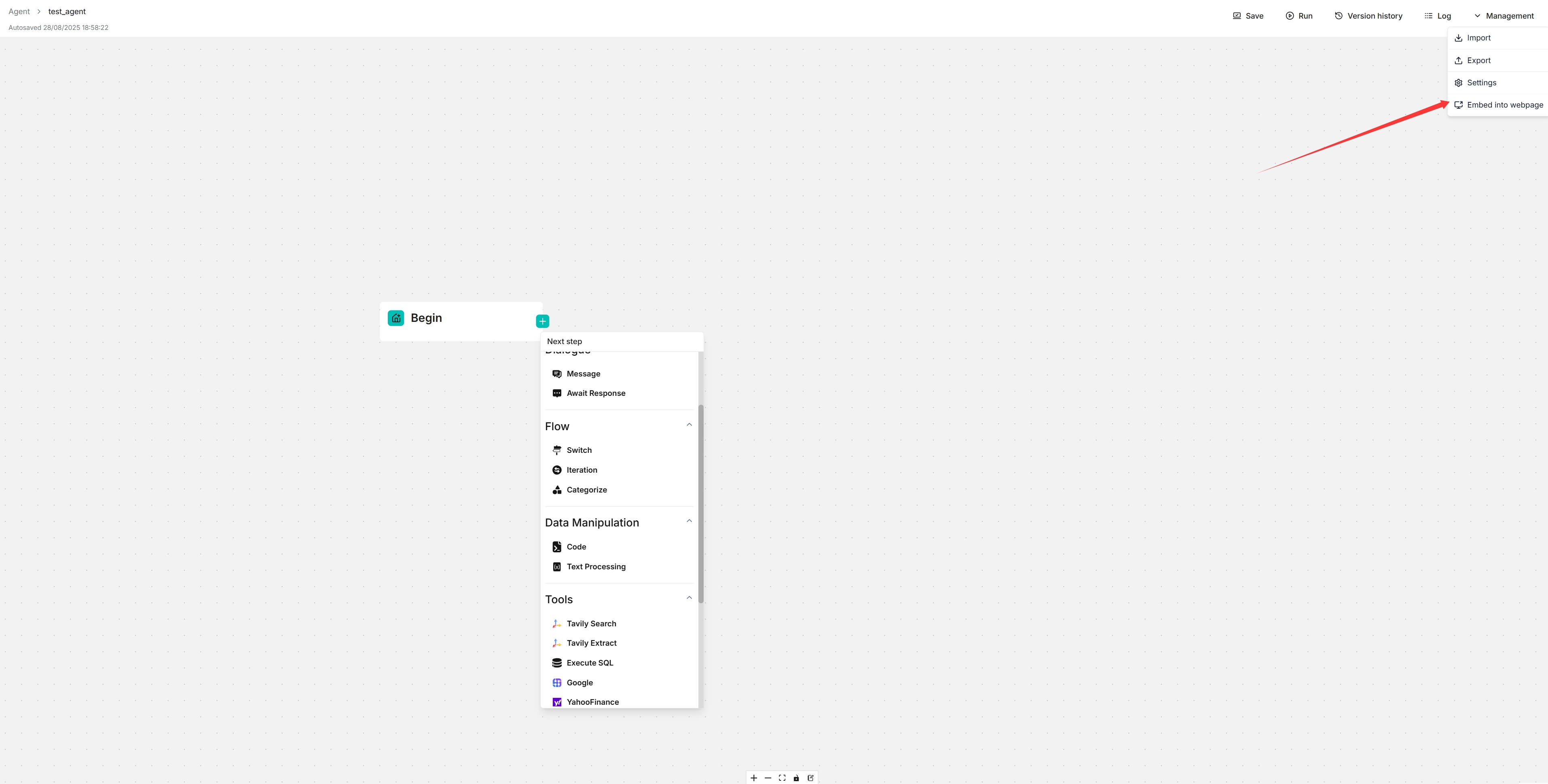
\ No newline at end of file
diff --git a/docs/guides/chat/implement_deep_research.md b/docs/guides/chat/implement_deep_research.md
index 2b07a4116e6..21f58f1e9fc 100644
--- a/docs/guides/chat/implement_deep_research.md
+++ b/docs/guides/chat/implement_deep_research.md
@@ -25,6 +25,6 @@ To activate this feature:
-*The following is a screenshot of a conversation that integrates Deep Research:*
+*The following is a screenshot of a conversation that integrates Deep research:*

\ No newline at end of file
diff --git a/docs/guides/chat/start_chat.md b/docs/guides/chat/start_chat.md
index e5066a8b297..501b9fabb73 100644
--- a/docs/guides/chat/start_chat.md
+++ b/docs/guides/chat/start_chat.md
@@ -40,7 +40,7 @@ You start an AI conversation by creating an assistant.
- **Top N** determines the *maximum* number of chunks to feed to the LLM. In other words, even if more chunks are retrieved, only the top N chunks are provided as input.
- **Multi-turn optimization** enhances user queries using existing context in a multi-round conversation. It is enabled by default. When enabled, it will consume additional LLM tokens and significantly increase the time to generate answers.
- **Use knowledge graph** indicates whether to use knowledge graph(s) in the specified dataset(s) during retrieval for multi-hop question answering. When enabled, this would involve iterative searches across entity, relationship, and community report chunks, greatly increasing retrieval time.
- - **Reasoning** indicates whether to generate answers through reasoning processes like Deepseek-R1/OpenAI o1. Once enabled, the chat model autonomously integrates Deep Research during question answering when encountering an unknown topic. This involves the chat model dynamically searching external knowledge and generating final answers through reasoning.
+ - **Reasoning** indicates whether to generate answers through reasoning processes like Deepseek-R1/OpenAI o1. Once enabled, the chat model autonomously integrates Deep research during question answering when encountering an unknown topic. This involves the chat model dynamically searching external knowledge and generating final answers through reasoning.
- **Rerank model** sets the reranker model to use. It is left empty by default.
- If **Rerank model** is left empty, the hybrid score system uses keyword similarity and vector similarity, and the default weight assigned to the vector similarity component is 1-0.7=0.3.
- If **Rerank model** is selected, the hybrid score system uses keyword similarity and reranker score, and the default weight assigned to the reranker score is 1-0.7=0.3.
diff --git a/docs/guides/dataset/add_data_source/_category_.json b/docs/guides/dataset/add_data_source/_category_.json
index 71b3d794d30..e4ba51baf42 100644
--- a/docs/guides/dataset/add_data_source/_category_.json
+++ b/docs/guides/dataset/add_data_source/_category_.json
@@ -1,5 +1,5 @@
{
- "label": "Add data source",
+ "label": "Add data sources",
"position": 18,
"link": {
"type": "generated-index",
diff --git a/docs/guides/dataset/add_data_source/add_confluence.md b/docs/guides/dataset/add_data_source/add_confluence.md
new file mode 100644
index 00000000000..273ceb107f1
--- /dev/null
+++ b/docs/guides/dataset/add_data_source/add_confluence.md
@@ -0,0 +1,58 @@
+---
+sidebar_position: 4
+slug: /add_confluence
+sidebar_custom_props: {
+ categoryIcon: SiGoogledrive
+}
+---
+
+# Add Confluence
+
+Integrate Confluence as a data source.
+
+---
+
+This guide outlines the integration of Confluence as a data source for RAGFlow.
+
+## Prerequisites
+
+Before configuring the connector, obtain the following credentials from your Atlassian account:
+
+- **Confluence user Email**: The email address of the account performing the indexing.
+- **Atlassian API Token**: Generated via [Atlassian Account Settings](https://id.atlassian.com/manage-profile/security/api-tokens).
+- **Confluence base URL**: The instance URL (e.g., `https://.atlassian.net/wiki`).
+
+## Configuration steps
+
+### Define Confluence as an external data source
+
+Navigate to the **Connectors** or **External Data Source** section in the RAGFlow Admin Panel and select **Confluence**. Enter the following in the popup window:
+
+- **Is Cloud** - A toggle indicating whether this is a Confluence Cloud instance.
+ - `Yes` (default): Confluence Cloud.
+ - `False`: Confluence Server/Data Center.
+- **Name**: *Required* A unique identifier for your Confluence connector (e.g., `Engineering-Wiki`).
+- **Confluence Username**: *Required*
+ - For Confluence Cloud: The full email address you use to log into Confluence.
+ - For Confluence Server/Data Center: Your login ID, often a shorthand name.
+- **Confluence Access Token**: *Required* The authentication key that allows RAGFlow to act on your behalf to read and index your wiki pages.
+ - For Confluence Cloud: An Atlassian API Token, a secure string generated from your global Atlassian account. Create one from id.atlassian.com/manage-profile/security/api-tokens.
+ - For Confluence Server/Data Center: Your Personal Access Token (PAT). You are required to log in to your company’s Confluence, click on your Profile Picture in the top right, select Settings, then, in the left-hand sidebar, look for Personal Access Tokens.
+- **Wiki Base URL**: The base URL of your confluence instance, e.g., https://your-domain.atlassian.net/wiki.
+- **Index Mode**
+ - `Everything`: (Default) Indexes all pages the provided credentials have access to.
+ - `Space`: RAGFlow restricts indexing only to the Space Keys you explicitly list in the configuration.
+ - **Space Keys:** Specify keys (e.g., `ENG, HR`) separated by commas to restrict indexing. Leave blank to index all accessible spaces.
+
+Once configuration is complete, click **Confirm** to save your changes.
+
+*RAGFlow validates the connection immediately.*
+
+### Link to a dataset
+
+Credentials alone do not trigger indexing. You must link the data source to a specific dataset:
+
+1. Navigate to the **Dataset** tab.
+2. Select or create the target Dataset.
+3. Navigate to the Dataset's **Configuration** page and select **Link data source**.
+4. Choose the previously created Confluence connector in the popup window.
diff --git a/docs/guides/dataset/add_data_source/add_github_repo.md b/docs/guides/dataset/add_data_source/add_github_repo.md
new file mode 100644
index 00000000000..9dc73831647
--- /dev/null
+++ b/docs/guides/dataset/add_data_source/add_github_repo.md
@@ -0,0 +1,67 @@
+---
+sidebar_position: 15
+slug: /add_github_repo
+sidebar_custom_props: {
+ categoryIcon: SiGoogledrive
+}
+---
+
+# Add GitHub repo
+
+Link your GitHub repo for pull request or issue synchronization.
+
+---
+
+This document explains how to link your GitHub repository to RAGFlow to synchronize pull requests and issues.
+
+## 1. GitHub configuration
+
+Before configuring RAGFlow, you must prepare your GitHub account and generate the necessary credentials.
+
+### Step a: Public email configuration
+
+To ensure smooth identity matching and permission synchronization between your organization and the RAG engine, it is a best practice to make your email visible.
+
+1. Go to your GitHub **Settings** > **Emails**.
+2. Uncheck "Keep my email addresses private".
+3. Go to **Public profile** and ensure your primary email is selected in the **Public email** dropdown.
+
+### Step b: Generate a personal access token (PAT)
+
+1. Navigate to **Settings** > **Developer settings** > **Personal access tokens** > **Tokens (classic)**.
+2. Click **Generate new token (classic)**.
+3. **Required scopes:** - **`repo` (Full control):** Essential for accessing private repositories, PRs, and issues.
+ - **`read:org` (Optional):** If you are syncing repositories across an entire organization.
+ - **`workflow` (Optional):** Recommended if you intend to index GitHub Action logs or CI/CD metadata.
+4. **Copy the token:** Save this immediately; it will not be displayed again.
+
+## 2. RAGFlow connector setup
+
+Once your GitHub token is ready, register the external data source within your RAGFlow instance.
+
+1. **Access data sources:** Click on your profile icon in RAGFlow and select **Data source**.
+2. **Add GitHub connector:** Click **+ Add** and select the **GitHub** icon.
+3. **Input configuration:**
+ - **Source name:** Name it based on the repository (e.g., `ragflow-repo`).
+ - **Repo owner:** The username or organization (e.g., `infiniflow`).
+ - **Repo name:** The repository identifier (e.g., `ragflow`).
+ - **Access token:** Paste the PAT generated in section 1.
+ - **Include Pull Request** Whether to include pull requests in the selected repo.
+ - **Include Issues** Whether to include issues in the selected repo.
+4. Click **Save** to confirm your changes.
+ *RAGFlow validates the connection immediately.*
+
+:::tip NOTE
+Currently deleted or modified files are not synchronized automatically. This feature is coming soon. Thanks to Gisselle-Gonzalez for requesting [this feature](https://github.com/infiniflow/ragflow/issues/13708).
+:::
+
+## 3. Dataset binding & ingestion
+
+Finally, link the connector to a specific knowledge base to begin the RAG process.
+
+1. **Create/select dataset:** Go to the **Dataset** tab and enter your target dataset.
+2. **Link external source:** Click **+ Add file** and select **External data source**.
+3. **Select GitHub source:** Pick the connector you just created.
+4. **Trigger initial sync:** - The files from the repo will appear in your file list.
+ - Select the files and click **Run/parsing**.
+ - **Parser selection:** For codebases, use the **"Naive"** parser for general text extraction or a specific code-aware template if available in your current version.
\ No newline at end of file
diff --git a/docs/guides/dataset/add_data_source/add_google_drive.md b/docs/guides/dataset/add_data_source/add_google_drive.md
index 57263094845..6e040a3b88b 100644
--- a/docs/guides/dataset/add_data_source/add_google_drive.md
+++ b/docs/guides/dataset/add_data_source/add_google_drive.md
@@ -5,76 +5,61 @@ sidebar_custom_props: {
categoryIcon: SiGoogledrive
}
---
-# Add Google Drive
-
-## 1. Create a Google Cloud Project
-You can either create a dedicated project for RAGFlow or use an existing
-Google Cloud external project.
-
-**Steps:**
-1. Open the project creation page\
-`https://console.cloud.google.com/projectcreate`
-
-2. Select **External** as the Audience
-
-3. Click **Create**
-
+# Add Google Drive
-------------------------------------------------------------------------
+Add Google Drive as one of the data sources in RAGFlow.
-## 2. Configure OAuth Consent Screen
+---
-1. Go to **APIs & Services → OAuth consent screen**
-2. Ensure **User Type = External**
-
-3. Add your test users under **Test Users** by entering email addresses
-
-
+This document provides step-by-step instructions for integrating Google Drive as a data source in RAGFlow.
-------------------------------------------------------------------------
+## 1. Create a Google Cloud project
-## 3. Create OAuth Client Credentials
+You can either create a dedicated project for RAGFlow or use an existing Google Cloud external project. In this case, we create a Google Cloud project from scratch:
-1. Navigate to:\
- `https://console.cloud.google.com/auth/clients`
-2. Create a **Web Application**
-
-3. Enter a name for the client
-4. Add the following **Authorized Redirect URIs**:
+1. Open the project creation page `https://console.cloud.google.com/projectcreate`:
+
+2. Under **App Information**, provide an App name and your Gmail account as user support email:
+
+3. Select **External**:
+ _Your app will start in testing mode and will only be available to a selected list of users._
+
+4: Click **Create** to confirm creation.
-```
-http://localhost:9380/v1/connector/google-drive/oauth/web/callback
-```
-
-- If using Docker deployment:
-
-**Authorized JavaScript origin:**
-```
-http://localhost:80
-```
+## 2. Configure OAuth Consent Screen
-
+You need to configure the OAuth Consent Screen because it is the step where you define how your app asks for permission and what specific data it wants to access on behalf of a user. It's a mandatory part of setting up OAuth 2.0 authentication with Google. Think of it as creating a standardized permission slip for your app. Without it, Google will not allow your app to request access to user data.
-- If running from source:
-**Authorized JavaScript origin:**
-```
-http://localhost:9222
-```
+1. Go to **APIs & Services** → **OAuth consent screen**.
+2. Ensure **User Type** is set to **External**:
+
+3. Under Under **Test Users**, click **+ Add users** to add test users:
+
+
-
+## 3. Create OAuth Client Credentials
-5. After saving, click **Download JSON**. This file will later be uploaded into RAGFlow.
+1. Navigate to `https://console.cloud.google.com/auth/clients`.
+2. Select **Web Application** as **Application type** for the created project:
+
+3. Enter a client name.
+4. Add `http://localhost:9380/v1/connector/google-drive/oauth/web/callback` as **Authorised redirect URIs**:
+5. Add **Authorised JavaScript origins**:
+ - If deploying RAGFlow from Docker, use `http://localhost:80`:
+ 
+ - If building RAGFlow from source, use `http://localhost:9222`
+ 
-
+6. After saving, click **Download JSON** in the popup window; this credential file will later be uploaded into RAGFlow.
-------------------------------------------------------------------------
+
## 4. Add Scopes
-1. Open **Data Access → Add or remove scopes**
+You need to add scopes to explicitly define the specific level of access your application requires from a user's Google Drive, such as read-only access to files. These scopes are presented to the user on the consent screen, ensuring transparency by showing exactly what permissions they are granted. To do so:
-2. Paste and add the following entries:
+1. Click **Data Access** → **Add or remove scopes**, and add the following entries and click **Update**:
```
https://www.googleapis.com/auth/drive.readonly
@@ -83,58 +68,46 @@ https://www.googleapis.com/auth/admin.directory.group.readonly
https://www.googleapis.com/auth/admin.directory.user.readonly
```
-
-3. Update and Save changes
-
-
-
-
-------------------------------------------------------------------------
-
-## 5. Enable Required APIs
-Navigate to the Google API Library:\
-`https://console.cloud.google.com/apis/library`
-
-
-Enable the following APIs:
-
-- Google Drive API
-- Admin SDK API
-- Google Sheets API
-- Google Docs API
-
+
+
-
+2. Click **Save** to save your data access changes:
-
+
-
+## 5. Enable required APIs
-
-
-
-
-
-
-------------------------------------------------------------------------
-
-## 6. Add Google Drive As a Data Source in RAGFlow
-
-1. Go to **Data Sources** inside RAGFlow
-2. Select **Google Drive**
-3. Upload the previously downloaded JSON credentials
-
-4. Enter the shared Google Drive folder link (https://drive.google.com/drive), such as:
-
-
-5. Click **Authorize with Google**
-A browser window will appear.
-
-Click: - **Continue** - **Select All → Continue** - Authorization should
-succeed - Select **OK** to add the data source
-
-
-
-
+You need to enable the required APIs (such as the Google Drive API) to formally grant your Google Cloud project permission to communicate with Google's services on behalf of your application. These APIs act as a gateway; even if you have valid OAuth credentials, Google will block requests to a disabled API. Enabling them ensures that when RAGFlow attempts to list or retrieve files, Google's servers recognize and authorize the request.
+1. Navigate to the Google API Library `https://console.cloud.google.com/apis/library`:
+
+2. Enable the following APIs:
+ - Google Drive API
+ - Admin SDK API
+ - Google Sheets API
+ - Google Docs API
+
+
+
+
+
+
+
+
+## 6. Add Google Drive as a data source in RAGFlow
+
+1. Go to **Data Sources** inside RAGFlow and select **Google Drive**.
+2. Under **OAuth Token JSON**, upload the previously downloaded JSON credentials you saved in [Section 2](#2-configure-oauth-consent-screen):
+
+3. Enter the url of the shared Google Drive folder link:
+
+4. Click **Authorize with Google**
+ _A browser window appears showing that Google hasn't verified this app._
+
+5. Click **Continue** → **Select All** → **Continue**.
+6. When the authorization succeeds, select **OK** to add the data source.
+
+
+
+
\ No newline at end of file
diff --git a/docs/guides/dataset/add_data_source/add_notion.md b/docs/guides/dataset/add_data_source/add_notion.md
new file mode 100644
index 00000000000..4535b6f3d99
--- /dev/null
+++ b/docs/guides/dataset/add_data_source/add_notion.md
@@ -0,0 +1,83 @@
+---
+sidebar_position: 5
+slug: /add_notion
+sidebar_custom_props: {
+ categoryIcon: SiGoogledrive
+}
+---
+
+# Add Notion
+
+Connecting your Notion workspace to RAGFlow allows you to ingest and sync your notes, databases, and documents directly into your dataset. Once configured, RAGFlow fetches data from the specified Notion pages to provide context for your RAG applications.
+
+## Prerequisites
+
+Before you begin, ensure you have:
+* A Notion account with **Workspace Owner** permissions (required to create integrations).
+* The specific pages or databases you intend to sync.
+
+---
+
+## Create an internal integration
+
+To allow RAGFlow to access your Notion data, you must first create an internal integration in the Notion developer portal to generate a secret token.
+
+1. Navigate to the [Notion My Integrations](https://www.notion.com/my-integrations) page.
+2. Click **+ New integration**.
+3. In the **Name** field, enter a name (e.g., "RAGFlow Connector").
+4. Select the **Associated workspace** where your data resides.
+5. Under **Capabilities**, ensure **Read content** is selected. RAGFlow does not require write or user-related permissions.
+6. Click **Submit**.
+7. Under the **Secrets** tab, click **Show** and then **Copy** to save your **Internal Integration Token**.
+
+---
+
+## Grant access to your pages
+
+By default, an integration has no access to any pages in your workspace. You must explicitly share the pages you want RAGFlow to index.
+
+1. Open the Notion page or database you wish to use as the root of your data source.
+2. Click the **...** (three dots) menu in the top-right corner.
+3. Scroll down to **Connect to** (or **Add connections**).
+4. Search for the integration you created (e.g., "RAGFlow Connector") and select it.
+5. Confirm the connection when prompted.
+
+:::tip NOTE
+If you share a parent page, all its nested child pages and databases will automatically be accessible to the integration.
+:::
+
+---
+
+## Identify the root page id
+
+The **Root Page Id** tells RAGFlow where to start indexing. You can find this in the URL of your Notion page.
+
+1. Open your target root page in a web browser.
+2. Look at the URL in the address bar. The page ID is the 32-character alphanumeric string at the end of the URL.
+ * **Format:** `https://www.notion.so/workspace-name/Page-Title-`**`11a047149aef80578303e705001bb90e`**
+3. Copy only the 32-character string (exclude any parameters following a `?`).
+
+---
+
+## Configure the notion connector in RAGFlow
+
+Once you have your token and ID, add the connector within the RAGFlow interface.
+
+| Field | Description | Required |
+| :--- | :--- | :--- |
+| **Name** | A unique label for this data source (e.g., `Engineering Wiki`). | Yes |
+| **Notion Integration Token** | The "Internal Integration Secret" copied from your Notion developer portal. | Yes |
+| **Root Page Id** | The 32-character ID of the top-level page you want to sync. | No |
+
+Once configuration is complete, click **Confirm** to save your changes.
+
+*RAGFlow validates the connection immediately.*
+
+### Link to a dataset
+
+Credentials alone do not trigger indexing. You must link the data source to a specific dataset:
+
+1. Navigate to the **Dataset** tab.
+2. Select or create the target Dataset.
+3. Navigate to the Dataset's **Configuration** page and select **Link data source**.
+4. Choose the previously created Notion connector in the popup window.
\ No newline at end of file
diff --git a/docs/guides/dataset/advanced/_category_.json b/docs/guides/dataset/advanced/_category_.json
new file mode 100644
index 00000000000..59b61230403
--- /dev/null
+++ b/docs/guides/dataset/advanced/_category_.json
@@ -0,0 +1,11 @@
+{
+ "label": "Advanced enrichment",
+ "position": 8,
+ "link": {
+ "type": "generated-index",
+ "description": "Advanced enrichment."
+ },
+ "customProps": {
+ "categoryIcon": "LucideFlower"
+ }
+}
diff --git a/docs/guides/dataset/auto_metadata.md b/docs/guides/dataset/advanced/auto_metadata.md
similarity index 99%
rename from docs/guides/dataset/auto_metadata.md
rename to docs/guides/dataset/advanced/auto_metadata.md
index 7a7b086361b..7814489d8e2 100644
--- a/docs/guides/dataset/auto_metadata.md
+++ b/docs/guides/dataset/advanced/auto_metadata.md
@@ -1,5 +1,5 @@
---
-sidebar_position: -6
+sidebar_position: 4
slug: /auto_metadata
sidebar_custom_props: {
categoryIcon: LucideFileCodeCorner
diff --git a/docs/guides/dataset/autokeyword_autoquestion.mdx b/docs/guides/dataset/advanced/autokeyword_autoquestion.mdx
similarity index 99%
rename from docs/guides/dataset/autokeyword_autoquestion.mdx
rename to docs/guides/dataset/advanced/autokeyword_autoquestion.mdx
index 3165a6a6b14..ae06006f118 100644
--- a/docs/guides/dataset/autokeyword_autoquestion.mdx
+++ b/docs/guides/dataset/advanced/autokeyword_autoquestion.mdx
@@ -1,5 +1,5 @@
---
-sidebar_position: 3
+sidebar_position: 0
slug: /autokeyword_autoquestion
sidebar_custom_props: {
categoryIcon: LucideSlidersHorizontal
diff --git a/docs/guides/dataset/construct_knowledge_graph.md b/docs/guides/dataset/advanced/construct_knowledge_graph.md
similarity index 99%
rename from docs/guides/dataset/construct_knowledge_graph.md
rename to docs/guides/dataset/advanced/construct_knowledge_graph.md
index b4eba1fd6b0..5b5f2198430 100644
--- a/docs/guides/dataset/construct_knowledge_graph.md
+++ b/docs/guides/dataset/advanced/construct_knowledge_graph.md
@@ -1,5 +1,5 @@
---
-sidebar_position: 8
+sidebar_position: 1
slug: /construct_knowledge_graph
sidebar_custom_props: {
categoryIcon: LucideWandSparkles
diff --git a/docs/guides/dataset/enable_raptor.md b/docs/guides/dataset/advanced/enable_raptor.md
similarity index 95%
rename from docs/guides/dataset/enable_raptor.md
rename to docs/guides/dataset/advanced/enable_raptor.md
index 54e36d2bf22..b312d7c94d2 100644
--- a/docs/guides/dataset/enable_raptor.md
+++ b/docs/guides/dataset/advanced/enable_raptor.md
@@ -1,5 +1,5 @@
---
-sidebar_position: 7
+sidebar_position: 2
slug: /enable_raptor
sidebar_custom_props: {
categoryIcon: LucideNetwork
@@ -34,7 +34,7 @@ The recursive clustering and summarization capture a broad understanding (by the
For multi-hop question-answering tasks involving complex, multistep reasoning, a semantic gap often exists between the question and its answer. As a result, searching with the question often fails to retrieve the relevant chunks that contribute to the correct answer. RAPTOR addresses this challenge by providing the chat model with richer and more context-aware and relevant chunks to summarize, enabling a holistic understanding without losing granular details.
:::tip NOTE
-Knowledge graphs can also be used for multi-hop question-answering tasks. See [Construct knowledge graph](./construct_knowledge_graph.md) for details. You may use either approach or both, but ensure you understand the memory, computational, and token costs involved.
+Knowledge graphs can also be used for multi-hop question-answering tasks. See [Construct knowledge graph](../advanced/construct_knowledge_graph.md) for details. You may use either approach or both, but ensure you understand the memory, computational, and token costs involved.
:::
## Prerequisites
diff --git a/docs/guides/dataset/extract_table_of_contents.md b/docs/guides/dataset/advanced/extract_table_of_contents.md
similarity index 98%
rename from docs/guides/dataset/extract_table_of_contents.md
rename to docs/guides/dataset/advanced/extract_table_of_contents.md
index fc86f78f466..8835d68dd3e 100644
--- a/docs/guides/dataset/extract_table_of_contents.md
+++ b/docs/guides/dataset/advanced/extract_table_of_contents.md
@@ -1,5 +1,5 @@
---
-sidebar_position: 4
+sidebar_position: 3
slug: /enable_table_of_contents
sidebar_custom_props: {
categoryIcon: LucideTableOfContents
diff --git a/docs/guides/dataset/configure_knowledge_base.md b/docs/guides/dataset/configure_knowledge_base.md
index 92fc1fec9ae..391dcee50bb 100644
--- a/docs/guides/dataset/configure_knowledge_base.md
+++ b/docs/guides/dataset/configure_knowledge_base.md
@@ -45,7 +45,7 @@ RAGFlow offers multiple built-in chunking template to facilitate chunking files
|--------------|-------------------------------------------------------------------------------|---------------------------------------------------------------------------------------------------------|
| General | Files are consecutively chunked based on a preset chunk token number. | MD, MDX, DOCX, XLSX, XLS (Excel 97-2003), PPT, PDF, TXT, JPEG, JPG, PNG, TIF, GIF, CSV, JSON, EML, HTML |
| Q&A | Retrieves relevant information and generates answers to respond to questions. | XLSX, XLS (Excel 97-2003), CSV/TXT |
-| Resume | Enterprise edition only. You can also try it out on demo.ragflow.io. | DOCX, PDF, TXT |
+| Resume | Enterprise edition only. You can also try it out on cloud.ragflow.io. | DOCX, PDF, TXT |
| Manual | | PDF |
| Table | The table mode uses TSI technology for efficient data parsing. | XLSX, XLS (Excel 97-2003), CSV/TXT |
| Paper | | PDF |
@@ -135,7 +135,7 @@ See [Run retrieval test](./run_retrieval_test.md) for details.
## Search for dataset
-As of RAGFlow v0.24.0, the search feature is still in a rudimentary form, supporting only dataset search by name.
+As of RAGFlow v0.25.0, the search feature is still in a rudimentary form, supporting only dataset search by name.
diff --git a/docs/guides/dataset/run_retrieval_test.md b/docs/guides/dataset/run_retrieval_test.md
index 973a2f2ed56..807d68278fc 100644
--- a/docs/guides/dataset/run_retrieval_test.md
+++ b/docs/guides/dataset/run_retrieval_test.md
@@ -18,7 +18,7 @@ During a retrieval test, chunks created from your specified chunking method are
- If no rerank model is selected, weighted keyword similarity will be combined with weighted vector cosine similarity.
- If a rerank model is selected, weighted keyword similarity will be combined with weighted vector reranking score.
-In contrast, chunks created from [knowledge graph construction](./construct_knowledge_graph.md) are retrieved solely using vector cosine similarity.
+In contrast, chunks created from [knowledge graph construction](./advanced/construct_knowledge_graph.md) are retrieved solely using vector cosine similarity.
## Prerequisites
@@ -94,4 +94,4 @@ If you have adjusted the default settings, such as keyword similarity weight or
### Is an LLM used when the Use Knowledge Graph switch is enabled?
-Yes, your LLM will be involved to analyze your query and extract the related entities and relationship from the knowledge graph. This also explains why additional tokens and time will be consumed.
\ No newline at end of file
+Yes, your LLM will be involved to analyze your query and extract the related entities and relationship from the knowledge graph. This also explains why additional tokens and time will be consumed.
diff --git a/docs/guides/dataset/select_pdf_parser.md b/docs/guides/dataset/select_pdf_parser.md
index fa2d068cb42..d96992f5af7 100644
--- a/docs/guides/dataset/select_pdf_parser.md
+++ b/docs/guides/dataset/select_pdf_parser.md
@@ -65,6 +65,12 @@ Starting from v0.22.0, RAGFlow includes MinerU (≥ 2.6.3) as an optional PDF p
- If you decide to use a chunking method from the **Built-in** dropdown, ensure it supports PDF parsing, then select **MinerU** from the **PDF parser** dropdown.
- If you use a custom ingestion pipeline instead, select **MinerU** in the **PDF parser** section of the **Parser** component.
+To use an external Docling Serve instance (instead of local in-process Docling), set:
+
+- `DOCLING_SERVER_URL`: The Docling Serve API endpoint (for example, `http://docling-host:5001`).
+
+When `DOCLING_SERVER_URL` is set, RAGFlow sends PDF content to Docling Serve (`/v1/convert/source`, with fallback to `/v1alpha/convert/source`) and ingests the returned markdown/text. If the variable is not set, RAGFlow keeps using local Docling (`USE_DOCLING=true` + installed package) behavior.
+
:::note
All MinerU environment variables are optional. When set, these values are used to auto-provision a MinerU OCR model for the tenant on first use. To avoid auto-provisioning, skip the environment variable settings and only configure MinerU from the **Model providers** page in the UI.
:::
diff --git a/docs/guides/dataset/set_metadata.md b/docs/guides/dataset/set_metadata.md
index 082fc70b540..6931281c2bc 100644
--- a/docs/guides/dataset/set_metadata.md
+++ b/docs/guides/dataset/set_metadata.md
@@ -31,4 +31,4 @@ Ensure that your metadata is in JSON format; otherwise, your updates will not be
### Can I set metadata for multiple documents at once?
-From v0.23.0 onwards, you can set metadata for each document individually or have the LLM auto-generate metadata for multiple files. See [Extract metadata](./auto_metadata.md) for details.
\ No newline at end of file
+From v0.23.0 onwards, you can set metadata for each document individually or have the LLM auto-generate metadata for multiple files. See [Extract metadata](./advanced/auto_metadata.md) for details.
\ No newline at end of file
diff --git a/docs/guides/manage_files.md b/docs/guides/manage_files.md
index bbb5b5ec143..4399bc71f97 100644
--- a/docs/guides/manage_files.md
+++ b/docs/guides/manage_files.md
@@ -89,4 +89,4 @@ RAGFlow's file management allows you to download an uploaded file:

-> As of RAGFlow v0.24.0, bulk download is not supported, nor can you download an entire folder.
+> As of RAGFlow v0.25.0, bulk download is not supported, nor can you download an entire folder.
diff --git a/docs/guides/migration/_category_.json b/docs/guides/migration/_category_.json
deleted file mode 100644
index 1099886f2ee..00000000000
--- a/docs/guides/migration/_category_.json
+++ /dev/null
@@ -1,11 +0,0 @@
-{
- "label": "Migration",
- "position": 5,
- "link": {
- "type": "generated-index",
- "description": "RAGFlow migration guide"
- },
- "customProps": {
- "categoryIcon": "LucideArrowRightLeft"
- }
-}
diff --git a/docs/guides/migration/migrate_from_docker_compose.md b/docs/guides/migration/migrate_from_docker_compose.md
deleted file mode 100644
index c2e8eeb5488..00000000000
--- a/docs/guides/migration/migrate_from_docker_compose.md
+++ /dev/null
@@ -1,108 +0,0 @@
-# Data Migration Guide
-
-A common scenario is processing large datasets on a powerful instance (e.g., with a GPU) and then migrating the entire RAGFlow service to a different production environment (e.g., a CPU-only server). This guide explains how to safely back up and restore your data using our provided migration script.
-
-## Identifying Your Data
-
-By default, RAGFlow uses Docker volumes to store all persistent data, including your database, uploaded files, and search indexes. You can see these volumes by running:
-
-```bash
-docker volume ls
-```
-
-The output will look similar to this:
-
-```text
-DRIVER VOLUME NAME
-local docker_esdata01
-local docker_minio_data
-local docker_mysql_data
-local docker_redis_data
-```
-
-These volumes contain all the data you need to migrate.
-
-## Step 1: Stop RAGFlow Services
-
-Before starting the migration, you must stop all running RAGFlow services on the **source machine**. Navigate to the project's root directory and run:
-
-```bash
-docker-compose -f docker/docker-compose.yml down
-```
-
-**Important:** Do **not** use the `-v` flag (e.g., `docker-compose down -v`), as this will delete all your data volumes. The migration script includes a check and will prevent you from running it if services are active.
-
-## Step 2: Back Up Your Data
-
-We provide a convenient script to package all your data volumes into a single backup folder.
-
-For a quick reference of the script's commands and options, you can run:
-```bash
-bash docker/migration.sh help
-```
-
-To create a backup, run the following command from the project's root directory:
-
-```bash
-bash docker/migration.sh backup
-```
-
-This will create a `backup/` folder in your project root containing compressed archives of your data volumes.
-
-You can also specify a custom name for your backup folder:
-
-```bash
-bash docker/migration.sh backup my_ragflow_backup
-```
-
-This will create a folder named `my_ragflow_backup/` instead.
-
-## Step 3: Transfer the Backup Folder
-
-Copy the entire backup folder (e.g., `backup/` or `my_ragflow_backup/`) from your source machine to the RAGFlow project directory on your **target machine**. You can use tools like `scp`, `rsync`, or a physical drive for the transfer.
-
-## Step 4: Restore Your Data
-
-On the **target machine**, ensure that RAGFlow services are not running. Then, use the migration script to restore your data from the backup folder.
-
-If your backup folder is named `backup/`, run:
-
-```bash
-bash docker/migration.sh restore
-```
-
-If you used a custom name, specify it in the command:
-
-```bash
-bash docker/migration.sh restore my_ragflow_backup
-```
-
-The script will automatically create the necessary Docker volumes and unpack the data.
-
-**Note:** If the script detects that Docker volumes with the same names already exist on the target machine, it will warn you that restoring will overwrite the existing data and ask for confirmation before proceeding.
-
-## Step 5: Start RAGFlow Services
-
-Once the restore process is complete, you can start the RAGFlow services on your new machine:
-
-```bash
-docker-compose -f docker/docker-compose.yml up -d
-```
-
-**Note:** If you already have built a service by docker-compose before, you may need to backup your data for target machine like this guide above and run like:
-
-```bash
-# Please backup by `sh docker/migration.sh backup backup_dir_name` before you do the following line.
-# !!! this line -v flag will delete the original docker volume
-docker-compose -f docker/docker-compose.yml down -v
-docker-compose -f docker/docker-compose.yml up -d
-```
-
-Your RAGFlow instance is now running with all the data from your original machine.
-
-
-
-
-
-
-
diff --git a/docs/guides/models/_category_.json b/docs/guides/models/_category_.json
index b4a996b4fa5..08f6f4ddf3a 100644
--- a/docs/guides/models/_category_.json
+++ b/docs/guides/models/_category_.json
@@ -1,6 +1,6 @@
{
"label": "Models",
- "position": -1,
+ "position": 8,
"link": {
"type": "generated-index",
"description": "Guides on model settings."
diff --git a/docs/guides/models/deploy_local_llm.mdx b/docs/guides/models/deploy_local_llm.mdx
index e7e3fbeaee3..2109ab5588f 100644
--- a/docs/guides/models/deploy_local_llm.mdx
+++ b/docs/guides/models/deploy_local_llm.mdx
@@ -9,11 +9,11 @@ sidebar_custom_props: {
import Tabs from '@theme/Tabs';
import TabItem from '@theme/TabItem';
-Deploy and run local models using Ollama, Xinference, VLLM ,SGLANG or other frameworks.
+Deploy and run local models using Ollama, Xinference, vLLM ,SGLang , GPUStack or other frameworks.
---
-RAGFlow supports deploying models locally using Ollama, Xinference, IPEX-LLM, or jina. If you have locally deployed models to leverage or wish to enable GPU or CUDA for inference acceleration, you can bind Ollama or Xinference into RAGFlow and use either of them as a local "server" for interacting with your local models.
+RAGFlow supports deploying models locally using Ollama, Xinference, IPEX-LLM, vLLM ,SGLang , GPUStack or jina. If you have locally deployed models to leverage or wish to enable GPU or CUDA for inference acceleration, you can bind Ollama or Xinference into RAGFlow and use either of them as a local "server" for interacting with your local models.
RAGFlow seamlessly integrates with Ollama and Xinference, without the need for further environment configurations. You can use them to deploy two types of local models in RAGFlow: chat models and embedding models.
@@ -316,28 +316,28 @@ To enable IPEX-LLM accelerated Ollama in RAGFlow, you must also complete the con
3. [Update System Model Settings](#6-update-system-model-settings)
4. [Update Chat Configuration](#7-update-chat-configuration)
-### 5. Deploy VLLM
+### 5. Deploy vLLM
ubuntu 22.04/24.04
```bash
- pip install vllm
- ```
+pip install vllm
+```
### 5.1 RUN VLLM WITH BEST PRACTISE
```bash
nohup vllm serve /data/Qwen3-8B --served-model-name Qwen3-8B-FP8 --dtype auto --port 1025 --gpu-memory-utilization 0.90 --tool-call-parser hermes --enable-auto-tool-choice > /var/log/vllm_startup1.log 2>&1 &
- ```
+```
you can get log info
```bash
- tail -f -n 100 /var/log/vllm_startup1.log
- ```
+tail -f -n 100 /var/log/vllm_startup1.log
+```
when see the follow ,it means vllm engine is ready for access
```bash
Starting vLLM API server 0 on http://0.0.0.0:1025
Started server process [19177]
Application startup complete.
- ```
+```
### 5.2 INTERGRATEING RAGFLOW WITH VLLM CHAT/EM/RERANK LLM WITH WEBUI
setting->model providers->search->vllm->add ,configure as follow:
@@ -350,6 +350,38 @@ select vllm chat model as default llm model as follow:
create chat->create conversations-chat as follow:
+### 6. Deploy GPUStack
+
+ubuntu 22.04/24.04
+
+### 6.1 RUN GPUStack WITH BEST PRACTISE
+
+```bash
+sudo docker run -d --name gpustack \
+ --restart unless-stopped \
+ -p 80:80 \
+ -p 10161:10161 \
+ --volume gpustack-data:/var/lib/gpustack \
+ gpustack/gpustack
+```
+you can get docker info
+```bash
+docker ps
+```
+when see the follow ,it means vllm engine is ready for access
+```bash
+root@gpustack-prod:~# docker ps
+CONTAINER ID IMAGE COMMAND CREATED STATUS PORTS NAMES
+abf59be84b1a gpustack/gpustack "/usr/bin/entrypoint…" 6 hours ago Up 6 hours 0.0.0.0:80->80/tcp, [::]:80->80/tcp, 0.0.0.0:10161->10161/tcp, [::]:10161->10161/tcp gpustack
+```
+### 6.2 INTERGRATEING RAGFLOW WITH GPUSTACK CHAT/EM/RERANK LLM WITH WEBUI
+
+setting->model providers->search->gpustack->add ,configure as follow:
+
+
+
+select gpustack chat model as default llm model as follow:
+
diff --git a/docs/guides/models/llm_api_key_setup.md b/docs/guides/models/llm_api_key_setup.md
index d2cf67597cc..704d31e395a 100644
--- a/docs/guides/models/llm_api_key_setup.md
+++ b/docs/guides/models/llm_api_key_setup.md
@@ -11,7 +11,7 @@ An API key is required for RAGFlow to interact with an online AI model. This gui
## Get model API key
-RAGFlow supports most mainstream LLMs. Please refer to [Supported Models](../../references/supported_models.mdx) for a complete list of supported models. You will need to apply for your model API key online. Note that most LLM providers grant newly-created accounts trial credit, which will expire in a couple of months, or a promotional amount of free quota.
+RAGFlow supports most mainstream LLMs. Please refer to [Supported Models](../../guides/models/supported_models.mdx) for a complete list of supported models. You will need to apply for your model API key online. Note that most LLM providers grant newly-created accounts trial credit, which will expire in a couple of months, or a promotional amount of free quota.
:::note
If you find your online LLM is not on the list, don't feel disheartened. The list is expanding, and you can [file a feature request](https://github.com/infiniflow/ragflow/issues/new?assignees=&labels=feature+request&projects=&template=feature_request.yml&title=%5BFeature+Request%5D%3A+) with us! Alternatively, if you have customized or locally-deployed models, you can [bind them to RAGFlow using Ollama, Xinference, or LocalAI](./deploy_local_llm.mdx).
diff --git a/docs/references/supported_models.mdx b/docs/guides/models/supported_models.mdx
similarity index 95%
rename from docs/references/supported_models.mdx
rename to docs/guides/models/supported_models.mdx
index d35f203a537..cc20e4120c2 100644
--- a/docs/references/supported_models.mdx
+++ b/docs/guides/models/supported_models.mdx
@@ -1,5 +1,5 @@
---
-sidebar_position: 1
+sidebar_position: 3
slug: /supported_models
sidebar_custom_props: {
categoryIcon: LucideBox
@@ -18,6 +18,7 @@ A complete list of models supported by RAGFlow, which will continue to expand.
| Provider | LLM | Image2Text | Speech2text | TTS | Embedding | Rerank | OCR |
| --------------------- | ------------------ | ------------------ | ------------------ | ------------------ | ------------------ | ------------------ | ------------------ |
| Anthropic | :heavy_check_mark: | | | | | | |
+| Avian | :heavy_check_mark: | | | | | | |
| Azure-OpenAI | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | | :heavy_check_mark: | | |
| BaiChuan | :heavy_check_mark: | | | | :heavy_check_mark: | | |
| BaiduYiyan | :heavy_check_mark: | :heavy_check_mark: | | | :heavy_check_mark: | :heavy_check_mark: | |
@@ -27,7 +28,7 @@ A complete list of models supported by RAGFlow, which will continue to expand.
| Fish Audio | | | | :heavy_check_mark: | | | |
| Gemini | :heavy_check_mark: | :heavy_check_mark: | | | :heavy_check_mark: | | |
| Google Cloud | :heavy_check_mark: | | | | | | |
-| GPUStack | :heavy_check_mark: | | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | |
+| GPUStack | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | |
| Groq | :heavy_check_mark: | | | | | | |
| HuggingFace | :heavy_check_mark: | | | | :heavy_check_mark: | | |
| Jina | | | | | :heavy_check_mark: | :heavy_check_mark: | |
@@ -45,6 +46,7 @@ A complete list of models supported by RAGFlow, which will continue to expand.
| OpenAI | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | :heavy_check_mark: | | |
| OpenAI-API-Compatible | :heavy_check_mark: | :heavy_check_mark: | | | :heavy_check_mark: | :heavy_check_mark: | |
| OpenRouter | :heavy_check_mark: | :heavy_check_mark: | | | | | |
+| Perplexity | | :heavy_check_mark: | | | | | |
| Replicate | :heavy_check_mark: | | | | :heavy_check_mark: | | |
| PPIO | :heavy_check_mark: | | | | | | |
| SILICONFLOW | :heavy_check_mark: | :heavy_check_mark: | | | :heavy_check_mark: | :heavy_check_mark: | |
diff --git a/docs/quickstart.mdx b/docs/quickstart.mdx
index e1de5fe184a..82b97a7e5a7 100644
--- a/docs/quickstart.mdx
+++ b/docs/quickstart.mdx
@@ -1,11 +1,11 @@
---
-sidebar_position: 0
+sidebar_position: 2
slug: /
sidebar_custom_props: {
sidebarIcon: LucideRocket
}
---
-# Get started
+# Quickstart
import Tabs from '@theme/Tabs';
import TabItem from '@theme/TabItem';
import APITable from '@site/src/components/APITable';
@@ -48,7 +48,7 @@ This section provides instructions on setting up the RAGFlow server on Linux. If
`vm.max_map_count`. This value sets the maximum number of memory map areas a process may have. Its default value is 65530. While most applications require fewer than a thousand maps, reducing this value can result in abnormal behaviors, and the system will throw out-of-memory errors when a process reaches the limitation.
- RAGFlow v0.24.0 uses Elasticsearch or [Infinity](https://github.com/infiniflow/infinity) for multiple recall. Setting the value of `vm.max_map_count` correctly is crucial to the proper functioning of the Elasticsearch component.
+ RAGFlow v0.25.0 uses Elasticsearch or [Infinity](https://github.com/infiniflow/infinity) for multiple recall. Setting the value of `vm.max_map_count` correctly is crucial to the proper functioning of the Elasticsearch component.
'`
- - Body:
- - `"ids"`: `list[string]` or `null`
+- Body:
+ - `"ids"`: `list[string]` or `null`
+ - `"delete_all"`: `boolean`
##### Request example
@@ -672,13 +676,24 @@ curl --request DELETE \
}'
```
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/datasets \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "delete_all": true
+ }'
+```
+
##### Request parameters
-- `"ids"`: (*Body parameter*), `list[string]` or `null`, *Required*
+- `"ids"`: (*Body parameter*), `list[string]` or `null`
Specifies the datasets to delete:
- - If `null`, all datasets will be deleted.
- - If an array of IDs, only the specified datasets will be deleted.
- - If an empty array, no datasets will be deleted.
+ - If omitted, or set to `null` or an empty array, no datasets are deleted.
+ - If an array of IDs is provided, only the datasets matching those IDs are deleted.
+- `"delete_all"`: (*Body parameter*), `boolean`
+ Whether to delete all datasets owned by the current user when`"ids"` is omitted, or set to `null` or an empty array. Defaults to `false`.
#### Response
@@ -808,6 +823,9 @@ curl --request PUT \
- Defaults to: `{"use_raptor": false}`
- `"graphrag"`: `object` GRAPHRAG-specific settings.
- Defaults to: `{"use_graphrag": false}`
+ - `"parent_child"`: `object` Parent-child chunking settings. When enabled, each chunk is further split into smaller child chunks using `children_delimiter`. At retrieval time, matched child chunks are replaced by their parent's full text before being passed to the LLM, giving precise vector matching with broader context.
+ - `"use_parent_child"`: `bool` Whether to enable parent-child chunking. Defaults to `false`.
+ - `"children_delimiter"`: `string` The delimiter used to split a parent chunk into child chunks. Only takes effect when `"use_parent_child"` is `true`. Defaults to `"\n"`.
- If `"chunk_method"` is `"qa"`, `"manuel"`, `"paper"`, `"book"`, `"laws"`, or `"presentation"`, the `"parser_config"` object contains the following attribute:
- `"raptor"`: `object` RAPTOR-specific settings.
- Defaults to: `{"use_raptor": false}`.
@@ -836,14 +854,14 @@ Failure:
### List datasets
-**GET** `/api/v1/datasets?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={dataset_name}&id={dataset_id}`
+**GET** `/api/v1/datasets?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={dataset_name}&id={dataset_id}&include_parsing_status={include_parsing_status}`
Lists datasets.
#### Request
- Method: GET
-- URL: `/api/v1/datasets?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={dataset_name}&id={dataset_id}`
+- URL: `/api/v1/datasets?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={dataset_name}&id={dataset_id}&include_parsing_status={include_parsing_status}`
- Headers:
- `'Authorization: Bearer '`
@@ -855,6 +873,13 @@ curl --request GET \
--header 'Authorization: Bearer '
```
+```bash
+# List datasets with parsing status
+curl --request GET \
+ --url 'http://{address}/api/v1/datasets?include_parsing_status=true' \
+ --header 'Authorization: Bearer '
+```
+
##### Request parameters
- `page`: (*Filter parameter*)
@@ -871,6 +896,13 @@ curl --request GET \
The name of the dataset to retrieve.
- `id`: (*Filter parameter*)
The ID of the dataset to retrieve.
+- `include_parsing_status`: (*Filter parameter*)
+ Whether to include document parsing status counts in the response. Defaults to `false`. When set to `true`, each dataset object in the response will include the following additional fields:
+ - `unstart_count`: Number of documents not yet started parsing.
+ - `running_count`: Number of documents currently being parsed.
+ - `cancel_count`: Number of documents whose parsing was cancelled.
+ - `done_count`: Number of documents that have been successfully parsed.
+ - `fail_count`: Number of documents whose parsing failed.
#### Response
@@ -918,6 +950,49 @@ Success:
}
```
+Success (with `include_parsing_status=true`):
+
+```json
+{
+ "code": 0,
+ "data": [
+ {
+ "avatar": null,
+ "cancel_count": 0,
+ "chunk_count": 30,
+ "chunk_method": "qa",
+ "create_date": "2026-03-09T18:57:13",
+ "create_time": 1773053833094,
+ "created_by": "928f92a210b911f1ac4cc39e0b8fa3ad",
+ "description": null,
+ "document_count": 1,
+ "done_count": 1,
+ "embedding_model": "text-embedding-v2@Tongyi-Qianwen",
+ "fail_count": 0,
+ "id": "ba6586c21ba611f1a3dc476f0709e75e",
+ "language": "English",
+ "name": "Test Dataset",
+ "parser_config": {
+ "graphrag": { "use_graphrag": false },
+ "llm_id": "deepseek-chat@DeepSeek",
+ "raptor": { "use_raptor": false }
+ },
+ "permission": "me",
+ "running_count": 0,
+ "similarity_threshold": 0.2,
+ "status": "1",
+ "tenant_id": "928f92a210b911f1ac4cc39e0b8fa3ad",
+ "token_num": 1746,
+ "unstart_count": 0,
+ "update_date": "2026-03-09T18:59:32",
+ "update_time": 1773053972723,
+ "vector_similarity_weight": 0.3
+ }
+ ],
+ "total_datasets": 1
+}
+```
+
Failure:
```json
@@ -1745,6 +1820,7 @@ Deletes documents by ID.
- `'Authorization: Bearer '`
- Body:
- `"ids"`: `list[string]`
+ - `"delete_all"`: `boolean`
##### Request example
@@ -1759,12 +1835,26 @@ curl --request DELETE \
}'
```
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/datasets/{dataset_id}/documents \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "delete_all": true
+ }'
+```
+
##### Request parameters
- `dataset_id`: (*Path parameter*)
The associated dataset ID.
- `"ids"`: (*Body parameter*), `list[string]`
- The IDs of the documents to delete. If it is not specified, all documents in the specified dataset will be deleted.
+ The IDs of the documents to delete.
+ - If omitted, or set to `null` or an empty array, no documents are deleted.
+ - If an array of IDs is provided, only the documents matching those IDs are deleted.
+- `"delete_all"`: (*Body parameter*), `boolean`
+ Whether to delete all documents in the specified dataset when `"ids"` is omitted, or set to `null` or an empty array. Defaults to `false`.
#### Response
@@ -1921,6 +2011,8 @@ Adds a chunk to a specified document in a specified dataset.
- Body:
- `"content"`: `string`
- `"important_keywords"`: `list[string]`
+ - `"tag_kwd"`: `list[string]`
+ - `"image_base64"`: `string`
##### Request example
@@ -1931,22 +2023,27 @@ curl --request POST \
--header 'Authorization: Bearer ' \
--data '
{
- "content": ""
+ "content": "",
+ "image_base64": ""
}'
```
##### Request parameters
-- `dataset_id`: (*Path parameter*)
+- `dataset_id`: (*Path parameter*)
The associated dataset ID.
-- `document_ids`: (*Path parameter*)
+- `document_ids`: (*Path parameter*)
The associated document ID.
-- `"content"`: (*Body parameter*), `string`, *Required*
+- `"content"`: (*Body parameter*), `string`, *Required*
The text content of the chunk.
-- `"important_keywords`(*Body parameter*), `list[string]`
+- `"important_keywords`(*Body parameter*), `list[string]`
The key terms or phrases to tag with the chunk.
+- `"tag_kwd"`: (*Body parameter*), `list[string]`
+ Tag keywords to associate with the chunk.
- `"questions"`(*Body parameter*), `list[string]`
If there is a given question, the embedded chunks will be based on them
+- `"image_base64"`: (*Body parameter*), `string`
+ A base64-encoded image to associate with the chunk. If the chunk already has an image, the new image will be vertically concatenated below the existing one.
#### Response
@@ -1963,7 +2060,9 @@ Success:
"dataset_id": "72f36e1ebdf411efb7250242ac120006",
"document_id": "61d68474be0111ef98dd0242ac120006",
"id": "12ccdc56e59837e5",
+ "image_id": "",
"important_keywords": [],
+ "tag_kwd": [],
"questions": []
}
}
@@ -2034,6 +2133,7 @@ Success:
"id": "b48c170e90f70af998485c1065490726",
"image_id": "",
"important_keywords": "",
+ "tag_kwd": [],
"positions": [
""
]
@@ -2103,6 +2203,7 @@ Deletes chunks by ID.
- `'Authorization: Bearer '`
- Body:
- `"chunk_ids"`: `list[string]`
+ - `"delete_all"`: `boolean`
##### Request example
@@ -2117,6 +2218,16 @@ curl --request DELETE \
}'
```
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/datasets/{dataset_id}/documents/{document_id}/chunks \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "delete_all": true
+ }'
+```
+
##### Request parameters
- `dataset_id`: (*Path parameter*)
@@ -2124,7 +2235,11 @@ curl --request DELETE \
- `document_ids`: (*Path parameter*)
The associated document ID.
- `"chunk_ids"`: (*Body parameter*), `list[string]`
- The IDs of the chunks to delete. If it is not specified, all chunks of the specified document will be deleted.
+ The IDs of the chunks to delete.
+ - If omitted, or set to `null` or an empty array, no chunks are deleted.
+ - If an array of IDs is provided, only the chunks matching those IDs are deleted.
+- `"delete_all"`: (*Body parameter*), `boolean`
+ Whether to delete all chunks of the specified documen when `"chunk_ids"` is omitted, or set to`null` or an empty array. Defaults to `false`.
#### Response
@@ -2163,6 +2278,7 @@ Updates content or configurations for a specified chunk.
- Body:
- `"content"`: `string`
- `"important_keywords"`: `list[string]`
+ - `"tag_kwd"`: `list[string]`
- `"available"`: `boolean`
##### Request example
@@ -2191,6 +2307,8 @@ curl --request PUT \
The text content of the chunk.
- `"important_keywords"`: (*Body parameter*), `list[string]`
A list of key terms or phrases to tag with the chunk.
+- `"tag_kwd"`: (*Body parameter*), `list[string]`
+ Updated tag keywords.
- `"available"`: (*Body parameter*) `boolean`
The chunk's availability status in the dataset. Value options:
- `true`: Available (default)
@@ -2217,6 +2335,105 @@ Failure:
---
+### Update chunk availability
+
+**POST** `/api/v1/datasets/{dataset_id}/documents/{document_id}/chunks/switch`
+
+Updates or switches the availability status of specified chunks, controlling whether they are available for retrieval.
+
+#### Request
+
+- Method: POST
+- URL: `/api/v1/datasets/{dataset_id}/documents/{document_id}/chunks/switch`
+- Headers:
+ - `'Content-Type: application/json'`
+ - `'Authorization: Bearer '`
+- Body:
+ - `"chunk_ids"`: `list[string]` (*Required*)
+ - `"available_int"`: `integer` (*Optional*)
+ - `"available"`: `boolean` (*Optional*)
+
+##### Request example
+
+```bash
+curl --request POST \
+ --url http://{address}/api/v1/datasets/{dataset_id}/documents/{document_id}/chunks/switch \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '
+ {
+ "chunk_ids": ["chunk_id_1", "chunk_id_2"],
+ "available_int": 1
+ }'
+```
+
+##### Request parameters
+
+- `dataset_id`: (*Path parameter*)
+ The ID of the dataset.
+- `document_id`: (*Path parameter*)
+ The ID of the document.
+- `"chunk_ids"`: (*Body parameter*), `list[string]` (*Required*)
+ IDs of the chunks whose availability status is to be updated.
+- `"available_int"`: (*Body parameter*), `integer` (*Optional*)
+ Availability status for the specified chunks. Mutually exclusive with `"available"`. You must provide either `available_int` or `available`, *not* both.
+ - `1`: Available,
+ - `0`: Unavailable.
+- `"available"`: (*Body parameter*), `boolean` (*Optional*)
+ Availability status of the specified chunks. Mutually exclusive with `"available_int"`. You must provide either `available` or `available_int`, *not* both.
+ - `true`: Available,
+ - `false`: Unavailable.
+
+#### Response
+
+Success:
+
+```json
+{
+ "code": 0,
+ "data": true
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 101,
+ "message": "You don't own the dataset {dataset_id}."
+}
+```
+
+```json
+{
+ "code": 101,
+ "message": "`chunk_ids` is required."
+}
+```
+
+```json
+{
+ "code": 101,
+ "message": "`available_int` or `available` is required."
+}
+```
+
+```json
+{
+ "code": 101,
+ "message": "Document not found!"
+}
+```
+
+```json
+{
+ "code": 101,
+ "message": "Index updating failure"
+}
+```
+
+---
+
### Retrieve a metadata summary from a dataset
**GET** `/api/v1/datasets/{dataset_id}/metadata/summary`
@@ -2436,7 +2653,7 @@ curl --request POST \
- `"top_k"`: (*Body parameter*), `integer`
The number of chunks engaged in vector cosine computation. Defaults to `1024`.
- `"use_kg"`: (*Body parameter*), `boolean`
- Whether to search chunks related to the generated knowledge graph for multi-hop queries. Defaults to `False`. Before enabling this, ensure you have successfully constructed a knowledge graph for the specified datasets. See [here](https://ragflow.io/docs/dev/construct_knowledge_graph) for details.
+ Whether to search chunks related to the generated knowledge graph for multi-hop queries. Defaults to `False`. Before enabling this, ensure you have successfully constructed a knowledge graph for the specified datasets. See [here](../guides/dataset/advanced/construct_knowledge_graph.md) for details.
- `"toc_enhance"`: (*Body parameter*), `boolean`
Whether to search chunks with extracted table of content. Defaults to `False`. Before enabling this, ensure you have enabled `TOC_Enhance` and successfully extracted table of contents for the specified datasets. See [here](https://ragflow.io/docs/dev/enable_table_of_contents) for details.
- `"rerank_id"`: (*Body parameter*), `integer`
@@ -2493,6 +2710,7 @@ Success:
"important_keywords": [
""
],
+ "tag_kwd": [],
"kb_id": "c7ee74067a2c11efb21c0242ac120006",
"positions": [
""
@@ -2544,10 +2762,11 @@ Creates a chat assistant.
- `'Authorization: Bearer '`
- Body:
- `"name"`: `string`
- - `"avatar"`: `string`
+ - `"icon"`: `string`
- `"dataset_ids"`: `list[string]`
- - `"llm"`: `object`
- - `"prompt"`: `object`
+ - `"llm_id"`: `string`
+ - `"llm_setting"`: `object`
+ - `"prompt_config"`: `object`
##### Request example
@@ -2566,27 +2785,16 @@ curl --request POST \
- `"name"`: (*Body parameter*), `string`, *Required*
The name of the chat assistant.
-- `"avatar"`: (*Body parameter*), `string`
+- `"icon"`: (*Body parameter*), `string`
Base64 encoding of the avatar.
-- `"dataset_ids"`: (*Body parameter*), `list[string]`
- The IDs of the associated datasets.
-- `"llm"`: (*Body parameter*), `object`
- The LLM settings for the chat assistant to create. If it is not explicitly set, a JSON object with the following values will be generated as the default. An `llm` JSON object contains the following attributes:
- - `"model_name"`, `string`
- The chat model name. If not set, the user's default chat model will be used.
-
- :::caution WARNING
- `model_type` is an *internal* parameter, serving solely as a temporary workaround for the current model-configuration design limitations.
-
- Its main purpose is to let *multimodal* models (stored in the database as `"image2text"`) pass backend validation/dispatching. Be mindful that:
-
- - Do *not* treat it as a stable public API.
- - It is subject to change or removal in future releases.
- :::
-
+- `"dataset_ids"`: (*Body parameter*), `list[string]`
+ The unique identifiers for the associated datasets. If omitted or set to `[]`, an empty chat assistant is created; datasets can be attached at a later time.
+- `"llm_id"`: (*Body parameter*), `string`
+ The identifier of the chat model. If not specified, the system defaults to the user's pre-configured chat model.
+- `"llm_setting"`: (*Body parameter*), `object`
+ A configuration object defining the LLM parameters for the assistant. The `llm_setting` object may contain the following attributes:
- `"model_type"`: `string`
A model type specifier. Only `"chat"` and `"image2text"` are recognized; any other inputs, or when omitted, are treated as `"chat"`.
- - `"model_name"`, `string`
- `"temperature"`: `float`
Controls the randomness of the model's predictions. A lower temperature results in more conservative responses, while a higher temperature yields more creative and diverse responses. Defaults to `0.1`.
- `"top_p"`: `float`
@@ -2595,21 +2803,27 @@ curl --request POST \
This discourages the model from repeating the same information by penalizing words that have already appeared in the conversation. Defaults to `0.4`.
- `"frequency penalty"`: `float`
Similar to the presence penalty, this reduces the model’s tendency to repeat the same words frequently. Defaults to `0.7`.
-- `"prompt"`: (*Body parameter*), `object`
- Instructions for the LLM to follow. If it is not explicitly set, a JSON object with the following values will be generated as the default. A `prompt` JSON object contains the following attributes:
- - `"similarity_threshold"`: `float` RAGFlow employs either a combination of weighted keyword similarity and weighted vector cosine similarity, or a combination of weighted keyword similarity and weighted reranking score during retrieval. This argument sets the threshold for similarities between the user query and chunks. If a similarity score falls below this threshold, the corresponding chunk will be excluded from the results. The default value is `0.2`.
- - `"keywords_similarity_weight"`: `float` This argument sets the weight of keyword similarity in the hybrid similarity score with vector cosine similarity or reranking model similarity. By adjusting this weight, you can control the influence of keyword similarity in relation to other similarity measures. The default value is `0.7`.
- - `"top_n"`: `int` This argument specifies the number of top chunks with similarity scores above the `similarity_threshold` that are fed to the LLM. The LLM will *only* access these 'top N' chunks. The default value is `6`.
- - `"variables"`: `object[]` This argument lists the variables to use in the 'System' field of **Chat Configurations**. Note that:
+- `"prompt_config"`: (*Body parameter*), `object`
+ Instructions for the LLM to follow. A `prompt_config` object may contain the following attributes:
+ - `"system"`: `string` The prompt content.
+ - `"prologue"`: `string` The opening greeting for the user.
+ - `"parameters"`: `object[]` This argument lists the variables to use in the system prompt. Note that:
- `"knowledge"` is a reserved variable, which represents the retrieved chunks.
- - All the variables in 'System' should be curly bracketed.
- - The default value is `[{"key": "knowledge", "optional": true}]`.
- - `"rerank_model"`: `string` If it is not specified, vector cosine similarity will be used; otherwise, reranking score will be used.
- - `top_k`: `int` Refers to the process of reordering or selecting the top-k items from a list or set based on a specific ranking criterion. Default to 1024.
+ - All the variables in `"system"` should be curly bracketed.
- `"empty_response"`: `string` If nothing is retrieved in the dataset for the user's question, this will be used as the response. To allow the LLM to improvise when nothing is found, leave this blank.
- - `"opener"`: `string` The opening greeting for the user. Defaults to `"Hi! I am your assistant, can I help you?"`.
- - `"show_quote`: `boolean` Indicates whether the source of text should be displayed. Defaults to `true`.
- - `"prompt"`: `string` The prompt content.
+ - `"quote"`: `boolean` Whether the source of text should be displayed. Defaults to `true`.
+ - `"tts"`: `boolean`
+ - `"refine_multiturn"`: `boolean`
+ - `"use_kg"`: `boolean`
+ - `"reasoning"`: `boolean`
+ - `"cross_languages"`: `list[string]`
+ - `"tavily_api_key"`: `string`
+ - `"toc_enhance"`: `boolean`
+- `"similarity_threshold"`: (*Body parameter*), `float`
+- `"vector_similarity_weight"`: (*Body parameter*), `float`
+- `"top_n"`: (*Body parameter*), `int`
+- `"top_k"`: (*Body parameter*), `int`
+- `"rerank_id"`: (*Body parameter*), `string`
#### Response
@@ -2619,39 +2833,42 @@ Success:
{
"code": 0,
"data": {
- "avatar": "",
+ "icon": "",
"create_date": "Thu, 24 Oct 2024 11:18:29 GMT",
"create_time": 1729768709023,
"dataset_ids": [
"527fa74891e811ef9c650242ac120006"
],
+ "kb_names": [
+ "dataset_1"
+ ],
"description": "A helpful Assistant",
- "do_refer": "1",
"id": "b1f2f15691f911ef81180242ac120003",
"language": "English",
- "llm": {
+ "llm_id": "qwen-plus@Tongyi-Qianwen",
+ "llm_setting": {
"frequency_penalty": 0.7,
- "model_name": "qwen-plus@Tongyi-Qianwen",
"presence_penalty": 0.4,
"temperature": 0.1,
"top_p": 0.3
},
"name": "12234",
- "prompt": {
+ "prompt_config": {
"empty_response": "Sorry! No relevant content was found in the knowledge base!",
- "keywords_similarity_weight": 0.3,
- "opener": "Hi! I'm your assistant. What can I do for you?",
- "prompt": "You are an intelligent assistant. Please summarize the content of the knowledge base to answer the question. Please list the data in the knowledge base and answer in detail. When all knowledge base content is irrelevant to the question, your answer must include the sentence \"The answer you are looking for is not found in the knowledge base!\" Answers need to consider chat history.\n ",
- "rerank_model": "",
- "similarity_threshold": 0.2,
- "top_n": 6,
- "variables": [
+ "prologue": "Hi! I'm your assistant. What can I do for you?",
+ "quote": true,
+ "system": "You are an intelligent assistant...",
+ "parameters": [
{
"key": "knowledge",
"optional": false
}
]
},
+ "rerank_id": "",
+ "similarity_threshold": 0.2,
+ "vector_similarity_weight": 0.3,
+ "top_n": 6,
"prompt_type": "simple",
"status": "1",
"tenant_id": "69736c5e723611efb51b0242ac120007",
@@ -2667,7 +2884,7 @@ Failure:
```json
{
"code": 102,
- "message": "Duplicated chat name in creating dataset."
+ "message": "Duplicated chat name."
}
```
@@ -2677,7 +2894,9 @@ Failure:
**PUT** `/api/v1/chats/{chat_id}`
-Updates configurations for a specified chat assistant.
+Overwrites the existing configuration for a specified chat assistant.
+
+Use this endpoint only when providing a complete configuration. Any fields omitted from the request will be reset to their server-side default values. For partial updates, use `PATCH /api/v1/chats/{chat_id}` instead.
#### Request
@@ -2688,10 +2907,11 @@ Updates configurations for a specified chat assistant.
- `'Authorization: Bearer '`
- Body:
- `"name"`: `string`
- - `"avatar"`: `string`
+ - `"icon"`: `string`
- `"dataset_ids"`: `list[string]`
- - `"llm"`: `object`
- - `"prompt"`: `object`
+ - `"llm_id"`: `string`
+ - `"llm_setting"`: `object`
+ - `"prompt_config"`: `object`
##### Request example
@@ -2702,7 +2922,23 @@ curl --request PUT \
--header 'Authorization: Bearer ' \
--data '
{
- "name":"Test"
+ "name":"Test",
+ "icon":"",
+ "dataset_ids":["0b2cbc8c877f11ef89070242ac120005"],
+ "llm_id":"qwen-plus@Tongyi-Qianwen",
+ "llm_setting":{"temperature":0.1,"top_p":0.3,"presence_penalty":0.4,"frequency_penalty":0.7},
+ "prompt_config":{
+ "system":"You are an intelligent assistant...",
+ "prologue":"Hi! I'\''m your assistant. What can I do for you?",
+ "parameters":[{"key":"knowledge","optional":false}],
+ "empty_response":"Sorry! No relevant content was found in the knowledge base!",
+ "quote":true
+ },
+ "similarity_threshold":0.2,
+ "vector_similarity_weight":0.3,
+ "top_n":6,
+ "top_k":1024,
+ "rerank_id":""
}'
```
@@ -2712,44 +2948,71 @@ curl --request PUT \
The ID of the chat assistant to update.
- `"name"`: (*Body parameter*), `string`, *Required*
The revised name of the chat assistant.
-- `"avatar"`: (*Body parameter*), `string`
+- `"icon"`: (*Body parameter*), `string`
Base64 encoding of the avatar.
-- `"dataset_ids"`: (*Body parameter*), `list[string]`
+- `"dataset_ids"`: (*Body parameter*), `list[string]`
The IDs of the associated datasets.
-- `"llm"`: (*Body parameter*), `object`
- The LLM settings for the chat assistant to create. If it is not explicitly set, a dictionary with the following values will be generated as the default. An `llm` object contains the following attributes:
- - `"model_name"`, `string`
- The chat model name. If not set, the user's default chat model will be used.
+- `"llm_id"`: (*Body parameter*), `string`
+ The chat model name. If not set, the user's default chat model is used.
+- `"llm_setting"`: (*Body parameter*), `object`
+ The LLM settings for the chat assistant. An `llm_setting` object contains the following attributes:
+ - `"model_type"`: `string`
+ A model type specifier. Supported values are `"chat"` and `"image2text"`. If the field is omitted or an unrecognized value is provided, it defaults to `"chat"`.
- `"temperature"`: `float`
Controls the randomness of the model's predictions. A lower temperature results in more conservative responses, while a higher temperature yields more creative and diverse responses. Defaults to `0.1`.
- `"top_p"`: `float`
Also known as “nucleus sampling”, this parameter sets a threshold to select a smaller set of words to sample from. It focuses on the most likely words, cutting off the less probable ones. Defaults to `0.3`
- `"presence_penalty"`: `float`
- This discourages the model from repeating the same information by penalizing words that have already appeared in the conversation. Defaults to `0.2`.
+ This discourages the model from repeating the same information by penalizing words that have already appeared in the conversation. Defaults to `0.4`.
- `"frequency penalty"`: `float`
Similar to the presence penalty, this reduces the model’s tendency to repeat the same words frequently. Defaults to `0.7`.
-- `"prompt"`: (*Body parameter*), `object`
- Instructions for the LLM to follow. A `prompt` object contains the following attributes:
- - `"similarity_threshold"`: `float` RAGFlow employs either a combination of weighted keyword similarity and weighted vector cosine similarity, or a combination of weighted keyword similarity and weighted rerank score during retrieval. This argument sets the threshold for similarities between the user query and chunks. If a similarity score falls below this threshold, the corresponding chunk will be excluded from the results. The default value is `0.2`.
- - `"keywords_similarity_weight"`: `float` This argument sets the weight of keyword similarity in the hybrid similarity score with vector cosine similarity or reranking model similarity. By adjusting this weight, you can control the influence of keyword similarity in relation to other similarity measures. The default value is `0.7`.
- - `"top_n"`: `int` This argument specifies the number of top chunks with similarity scores above the `similarity_threshold` that are fed to the LLM. The LLM will *only* access these 'top N' chunks. The default value is `8`.
- - `"variables"`: `object[]` This argument lists the variables to use in the 'System' field of **Chat Configurations**. Note that:
- - `"knowledge"` is a reserved variable, which represents the retrieved chunks.
- - All the variables in 'System' should be curly bracketed.
- - The default value is `[{"key": "knowledge", "optional": true}]`
- - `"rerank_model"`: `string` If it is not specified, vector cosine similarity will be used; otherwise, reranking score will be used.
- - `"empty_response"`: `string` If nothing is retrieved in the dataset for the user's question, this will be used as the response. To allow the LLM to improvise when nothing is found, leave this blank.
- - `"opener"`: `string` The opening greeting for the user. Defaults to `"Hi! I am your assistant, can I help you?"`.
- - `"show_quote`: `boolean` Indicates whether the source of text should be displayed. Defaults to `true`.
- - `"prompt"`: `string` The prompt content.
+- `"prompt_config"`: (*Body parameter*), `object`
+- `"similarity_threshold"`: (*Body parameter*), `float`
+- `"vector_similarity_weight"`: (*Body parameter*), `float`
+- `"top_n"`: (*Body parameter*), `int`
+- `"top_k"`: (*Body parameter*), `int`
+- `"rerank_id"`: (*Body parameter*), `string`
+
+For `PUT` requests, any fields omitted from the request body are reset to their server-side default values.
#### Response
-Success:
+Success: returns the full updated chat assistant object.
```json
{
- "code": 0
+ "code": 0,
+ "data": {
+ "id": "04d0d8e28d1911efa3630242ac120006",
+ "name": "Test",
+ "description": "A helpful Assistant",
+ "icon": "",
+ "dataset_ids": ["527fa74891e811ef9c650242ac120006"],
+ "kb_names": ["dataset_1"],
+ "llm_id": "qwen-plus@Tongyi-Qianwen",
+ "llm_setting": {
+ "frequency_penalty": 0.7,
+ "presence_penalty": 0.4,
+ "temperature": 0.1,
+ "top_p": 0.3
+ },
+ "prompt_config": {
+ "empty_response": "Sorry! No relevant content was found in the knowledge base!",
+ "prologue": "Hi! I'm your assistant. What can I do for you?",
+ "quote": true,
+ "system": "You are an intelligent assistant...",
+ "parameters": [{"key": "knowledge", "optional": false}]
+ },
+ "similarity_threshold": 0.2,
+ "vector_similarity_weight": 0.3,
+ "top_n": 6,
+ "top_k": 1024,
+ "rerank_id": "",
+ "status": "1",
+ "tenant_id": "69736c5e723611efb51b0242ac120007",
+ "create_time": 1729232406637,
+ "update_time": 1729232406638
+ }
}
```
@@ -2758,45 +3021,37 @@ Failure:
```json
{
"code": 102,
- "message": "Duplicated chat name in updating dataset."
+ "message": "Duplicated chat name."
}
```
---
-### Delete chat assistants
+### Get chat assistant
-**DELETE** `/api/v1/chats`
+**GET** `/api/v1/chats/{chat_id}`
-Deletes chat assistants by ID.
+Retrieves a specified chat assistant.
#### Request
-- Method: DELETE
-- URL: `/api/v1/chats`
+- Method: GET
+- URL: `/api/v1/chats/{chat_id}`
- Headers:
- - `'content-Type: application/json'`
- `'Authorization: Bearer '`
-- Body:
- - `"ids"`: `list[string]`
##### Request example
```bash
-curl --request DELETE \
- --url http://{address}/api/v1/chats \
- --header 'Content-Type: application/json' \
- --header 'Authorization: Bearer ' \
- --data '
- {
- "ids": ["test_1", "test_2"]
- }'
+curl --request GET \
+ --url http://{address}/api/v1/chats/{chat_id} \
+ --header 'Authorization: Bearer '
```
##### Request parameters
-- `"ids"`: (*Body parameter*), `list[string]`
- The IDs of the chat assistants to delete. If it is not specified, all chat assistants in the system will be deleted.
+- `chat_id`: (*Path parameter*)
+ The ID of the chat assistant to retrieve.
#### Response
@@ -2804,7 +3059,38 @@ Success:
```json
{
- "code": 0
+ "code": 0,
+ "data": {
+ "icon": "",
+ "create_date": "Fri, 18 Oct 2024 06:20:06 GMT",
+ "create_time": 1729232406637,
+ "description": "A helpful Assistant",
+ "id": "04d0d8e28d1911efa3630242ac120006",
+ "dataset_ids": ["527fa74891e811ef9c650242ac120006"],
+ "kb_names": ["dataset_1"],
+ "language": "English",
+ "llm_id": "qwen-plus@Tongyi-Qianwen",
+ "llm_setting": {
+ "temperature": 0.1,
+ "top_p": 0.3
+ },
+ "name": "my_chat",
+ "prompt_config": {
+ "empty_response": "Sorry! No relevant content was found in the knowledge base!",
+ "prologue": "Hi! I'm your assistant. What can I do for you?",
+ "quote": true,
+ "system": "You are an intelligent assistant...",
+ "parameters": [{"key": "knowledge", "optional": false}]
+ },
+ "rerank_id": "",
+ "similarity_threshold": 0.2,
+ "vector_similarity_weight": 0.3,
+ "top_n": 6,
+ "status": "1",
+ "tenant_id": "69736c5e723611efb51b0242ac120007",
+ "update_date": "Fri, 18 Oct 2024 06:20:06 GMT",
+ "update_time": 1729232406638
+ }
}
```
@@ -2813,99 +3099,283 @@ Failure:
```json
{
"code": 102,
- "message": "ids are required"
+ "message": "No authorization."
}
```
---
-### List chat assistants
+### Partially update chat assistant
-**GET** `/api/v1/chats?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={chat_name}&id={chat_id}`
+**PATCH** `/api/v1/chats/{chat_id}`
-Lists chat assistants.
+Performs a partial update on a specified chat assistant.
+
+Unspecified fields are preserved, while nested objects, such as `llm_setting` and `prompt_config`, are deep-merged with the existing configuration. This is the recommended endpoint for renaming an assistant or modifying a specific subset of settings.
#### Request
-- Method: GET
-- URL: `/api/v1/chats?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={chat_name}&id={chat_id}`
+- Method: PATCH
+- URL: `/api/v1/chats/{chat_id}`
- Headers:
+ - `'content-Type: application/json'`
- `'Authorization: Bearer '`
+- Body: any subset of the fields accepted by `PUT /api/v1/chats/{chat_id}`
##### Request example
```bash
-curl --request GET \
- --url http://{address}/api/v1/chats?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={chat_name}&id={chat_id} \
- --header 'Authorization: Bearer '
+curl --request PATCH \
+ --url http://{address}/api/v1/chats/{chat_id} \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "llm_id": "gpt-4o",
+ "llm_setting": {"temperature": 0.5}
+}'
```
-##### Request parameters
-
-- `page`: (*Filter parameter*), `integer`
- Specifies the page on which the chat assistants will be displayed. Defaults to `1`.
-- `page_size`: (*Filter parameter*), `integer`
- The number of chat assistants on each page. Defaults to `30`.
-- `orderby`: (*Filter parameter*), `string`
- The attribute by which the results are sorted. Available options:
- - `create_time` (default)
- - `update_time`
-- `desc`: (*Filter parameter*), `boolean`
- Indicates whether the retrieved chat assistants should be sorted in descending order. Defaults to `true`.
-- `id`: (*Filter parameter*), `string`
- The ID of the chat assistant to retrieve.
-- `name`: (*Filter parameter*), `string`
- The name of the chat assistant to retrieve.
-
#### Response
-Success:
+Success: returns the full updated chat assistant object (same structure as `PUT /api/v1/chats/{chat_id}`).
```json
{
"code": 0,
- "data": [
- {
- "avatar": "",
- "create_date": "Fri, 18 Oct 2024 06:20:06 GMT",
- "create_time": 1729232406637,
- "description": "A helpful Assistant",
- "do_refer": "1",
- "id": "04d0d8e28d1911efa3630242ac120006",
- "dataset_ids": ["527fa74891e811ef9c650242ac120006"],
- "language": "English",
- "llm": {
- "frequency_penalty": 0.7,
- "model_name": "qwen-plus@Tongyi-Qianwen",
- "presence_penalty": 0.4,
- "temperature": 0.1,
- "top_p": 0.3
- },
- "name": "13243",
- "prompt": {
- "empty_response": "Sorry! No relevant content was found in the knowledge base!",
- "keywords_similarity_weight": 0.3,
- "opener": "Hi! I'm your assistant. What can I do for you?",
- "prompt": "You are an intelligent assistant. Please summarize the content of the knowledge base to answer the question. Please list the data in the knowledge base and answer in detail. When all knowledge base content is irrelevant to the question, your answer must include the sentence \"The answer you are looking for is not found in the knowledge base!\" Answers need to consider chat history.\n",
- "rerank_model": "",
- "similarity_threshold": 0.2,
- "top_n": 6,
- "variables": [
- {
- "key": "knowledge",
- "optional": false
- }
- ]
- },
- "prompt_type": "simple",
- "status": "1",
- "tenant_id": "69736c5e723611efb51b0242ac120007",
- "top_k": 1024,
- "update_date": "Fri, 18 Oct 2024 06:20:06 GMT",
- "update_time": 1729232406638
- }
- ]
-}
+ "data": {
+ "id": "04d0d8e28d1911efa3630242ac120006",
+ "name": "Renamed assistant",
+ "llm_id": "qwen-plus@Tongyi-Qianwen",
+ "..." : "..."
+ }
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "No authorization."
+}
+```
+
+---
+
+### Delete chat assistant
+
+**DELETE** `/api/v1/chats/{chat_id}`
+
+Deletes a chat assistant by ID.
+
+#### Request
+
+- Method: DELETE
+- URL: `/api/v1/chats/{chat_id}`
+- Headers:
+ - `'Authorization: Bearer '`
+
+##### Request example
+
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/chats/{chat_id} \
+ --header 'Authorization: Bearer '
+```
+
+##### Request parameters
+
+- `chat_id`: (*Path parameter*)
+ The ID of the chat assistant to delete.
+
+#### Response
+
+Success:
+
+```json
+{
+ "code": 0,
+ "data": true
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "No authorization."
+}
+```
+
+---
+
+### Delete chat assistants
+
+**DELETE** `/api/v1/chats`
+
+Deletes chat assistants by ID.
+
+#### Request
+
+- Method: DELETE
+- URL: `/api/v1/chats`
+- Headers:
+ - `'content-Type: application/json'`
+ - `'Authorization: Bearer '`
+- Body:
+ - `"ids"`: `list[string]`
+ - `"delete_all"`: `boolean`
+
+##### Request example
+
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/chats \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '
+ {
+ "ids": ["test_1", "test_2"]
+ }'
+```
+
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/chats \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "delete_all": true
+ }'
+```
+
+##### Request parameters
+
+- `"ids"`: (*Body parameter*), `list[string]`
+ The IDs of the chat assistants to delete.
+ - If omitted, or set to `null` or an empty array, no chat assistants are deleted.
+ - If an array of IDs is provided, only the chat assistants matching those IDs are deleted.
+- `"delete_all"`: (*Body parameter*), `boolean`
+ Whether to delete all chat assistants owned by the current user when `"ids"` is omitted, or set to`null` or an empty array. Defaults to `false`.
+
+#### Response
+
+Success:
+
+```json
+{
+ "code": 0
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "ids are required"
+}
+```
+
+---
+
+### List chat assistants
+
+**GET** `/api/v1/chats?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&keywords={keywords}&owner_ids={owner_id}&name={chat_name}&id={chat_id}`
+
+Lists chat assistants.
+
+#### Request
+
+- Method: GET
+- URL: `/api/v1/chats?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&keywords={keywords}&owner_ids={owner_id}&name={chat_name}&id={chat_id}`
+- Headers:
+ - `'Authorization: Bearer '`
+
+##### Request example
+
+```bash
+curl --request GET \
+ --url http://{address}/api/v1/chats?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&keywords={keywords}&owner_ids={owner_id}&name={chat_name}&id={chat_id} \
+ --header 'Authorization: Bearer '
+```
+
+##### Request parameters
+
+- `page`: (*Filter parameter*), `integer`
+ Specifies the page on which the chat assistants will be displayed. Defaults to `1`.
+- `page_size`: (*Filter parameter*), `integer`
+ The number of chat assistants on each page. Defaults to `30`.
+- `orderby`: (*Filter parameter*), `string`
+ The attribute by which the results are sorted. Available options:
+ - `create_time` (default)
+ - `update_time`
+- `desc`: (*Filter parameter*), `boolean`
+ Indicates whether the retrieved chat assistants should be sorted in descending order. Defaults to `true`.
+- `keywords`: (*Filter parameter*), `string`
+ Case-insensitive fuzzy match against chat assistant names.
+- `owner_ids`: (*Filter parameter*), `string` (repeatable)
+ Filter by owner tenant IDs. Can be specified multiple times: `?owner_ids=id1&owner_ids=id2`.
+- `id`: (*Filter parameter*), `string`
+ The ID of the chat assistant to retrieve with exact match.
+- `name`: (*Filter parameter*), `string`
+ The name of the chat assistant to retrieve with exact match.
+
+When `id` or `name` is provided, exact filtering takes precedence over `keywords`.
+
+#### Response
+
+Success:
+
+```json
+{
+ "code": 0,
+ "data": {
+ "chats": [
+ {
+ "icon": "",
+ "create_date": "Fri, 18 Oct 2024 06:20:06 GMT",
+ "create_time": 1729232406637,
+ "description": "A helpful Assistant",
+ "id": "04d0d8e28d1911efa3630242ac120006",
+ "dataset_ids": ["527fa74891e811ef9c650242ac120006"],
+ "kb_names": ["dataset_1"],
+ "language": "English",
+ "llm_id": "qwen-plus@Tongyi-Qianwen",
+ "llm_setting": {
+ "frequency_penalty": 0.7,
+ "presence_penalty": 0.4,
+ "temperature": 0.1,
+ "top_p": 0.3
+ },
+ "name": "13243",
+ "prompt_config": {
+ "empty_response": "Sorry! No relevant content was found in the knowledge base!",
+ "prologue": "Hi! I'm your assistant. What can I do for you?",
+ "quote": true,
+ "system": "You are an intelligent assistant...",
+ "parameters": [
+ {
+ "key": "knowledge",
+ "optional": false
+ }
+ ]
+ },
+ "rerank_id": "",
+ "similarity_threshold": 0.2,
+ "vector_similarity_weight": 0.3,
+ "top_n": 6,
+ "prompt_type": "simple",
+ "status": "1",
+ "tenant_id": "69736c5e723611efb51b0242ac120007",
+ "update_date": "Fri, 18 Oct 2024 06:20:06 GMT",
+ "update_time": 1729232406638
+ }
+ ],
+ "total": 1
+ }
+}
```
Failure:
@@ -2992,7 +3462,7 @@ Failure:
```json
{
"code": 102,
- "message": "Name cannot be empty."
+ "message": "`name` can not be empty."
}
```
@@ -3012,8 +3482,7 @@ Updates a session of a specified chat assistant.
- `'content-Type: application/json'`
- `'Authorization: Bearer '`
- Body:
- - `"name`: `string`
- - `"user_id`: `string` (optional)
+ - `"name"`: `string`
##### Request example
@@ -3030,14 +3499,12 @@ curl --request PUT \
##### Request Parameter
-- `chat_id`: (*Path parameter*)
+- `chat_id`: (*Path parameter*)
The ID of the associated chat assistant.
-- `session_id`: (*Path parameter*)
+- `session_id`: (*Path parameter*)
The ID of the session to update.
-- `"name"`: (*Body Parameter*), `string`
+- `"name"`: (*Body Parameter*), `string`
The revised name of the session.
-- `"user_id"`: (*Body parameter*), `string`
- Optional user-defined ID.
#### Response
@@ -3045,7 +3512,23 @@ Success:
```json
{
- "code": 0
+ "code": 0,
+ "data": {
+ "chat_id": "2ca4b22e878011ef88fe0242ac120005",
+ "create_date": "Fri, 11 Oct 2024 08:46:14 GMT",
+ "create_time": 1728636374571,
+ "id": "4606b4ec87ad11efbc4f0242ac120006",
+ "messages": [
+ {
+ "content": "Hi! I am your assistant, can I help you?",
+ "role": "assistant"
+ }
+ ],
+ "name": "updated session name",
+ "update_date": "Fri, 11 Oct 2024 08:46:14 GMT",
+ "update_time": 1728636374571,
+ "user_id": ""
+ }
}
```
@@ -3054,7 +3537,7 @@ Failure:
```json
{
"code": 102,
- "message": "Name cannot be empty."
+ "message": "`name` can not be empty."
}
```
@@ -3062,7 +3545,7 @@ Failure:
### List chat assistant's sessions
-**GET** `/api/v1/chats/{chat_id}/sessions?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={session_name}&id={session_id}`
+**GET** `/api/v1/chats/{chat_id}/sessions?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={session_name}&id={session_id}&user_id={user_id}`
Lists sessions associated with a specified chat assistant.
@@ -3077,7 +3560,7 @@ Lists sessions associated with a specified chat assistant.
```bash
curl --request GET \
- --url http://{address}/api/v1/chats/{chat_id}/sessions?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={session_name}&id={session_id} \
+ --url http://{address}/api/v1/chats/{chat_id}/sessions?page={page}&page_size={page_size}&orderby={orderby}&desc={desc}&name={session_name}&id={session_id}&user_id={user_id} \
--header 'Authorization: Bearer '
```
@@ -3088,7 +3571,7 @@ curl --request GET \
- `page`: (*Filter parameter*), `integer`
Specifies the page on which the sessions will be displayed. Defaults to `1`.
- `page_size`: (*Filter parameter*), `integer`
- The number of sessions on each page. Defaults to `30`.
+ The number of sessions on each page. Defaults to `30`. If set to `0`, an empty list is returned.
- `orderby`: (*Filter parameter*), `string`
The field by which sessions should be sorted. Available options:
- `create_time` (default)
@@ -3111,7 +3594,7 @@ Success:
"code": 0,
"data": [
{
- "chat": "2ca4b22e878011ef88fe0242ac120005",
+ "chat_id": "2ca4b22e878011ef88fe0242ac120005",
"create_date": "Fri, 11 Oct 2024 08:46:43 GMT",
"create_time": 1728636403974,
"id": "578d541e87ad11ef96b90242ac120006",
@@ -3122,8 +3605,10 @@ Success:
}
],
"name": "new session",
+ "reference": [],
"update_date": "Fri, 11 Oct 2024 08:46:43 GMT",
- "update_time": 1728636403974
+ "update_time": 1728636403974,
+ "user_id": ""
}
]
}
@@ -3140,6 +3625,202 @@ Failure:
---
+### Get chat assistant's session
+
+**GET** `/api/v1/chats/{chat_id}/sessions/{session_id}`
+
+Gets a specific session of a specified chat assistant, including its messages, references, and avatar.
+
+#### Request
+
+- Method: GET
+- URL: `/api/v1/chats/{chat_id}/sessions/{session_id}`
+- Headers:
+ - `'Authorization: Bearer '`
+
+##### Request example
+
+```bash
+curl --request GET \
+ --url http://{address}/api/v1/chats/{chat_id}/sessions/{session_id} \
+ --header 'Authorization: Bearer '
+```
+
+##### Request Parameters
+
+- `chat_id`: (*Path parameter*)
+ The ID of the associated chat assistant.
+- `session_id`: (*Path parameter*)
+ The ID of the session to retrieve.
+
+#### Response
+
+Success:
+
+```json
+{
+ "code": 0,
+ "data": {
+ "chat_id": "2ca4b22e878011ef88fe0242ac120005",
+ "id": "4606b4ec87ad11efbc4f0242ac120006",
+ "name": "new session",
+ "avatar": "data:image/png;base64,...",
+ "messages": [
+ {
+ "content": "Hi! I am your assistant, can I help you?",
+ "role": "assistant"
+ }
+ ],
+ "reference": []
+ }
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "Session not found!"
+}
+```
+
+---
+
+### Delete a message from a chat assistant's session
+
+**DELETE** `/api/v1/chats/{chat_id}/sessions/{session_id}/messages/{msg_id}`
+
+Deletes a user message and its paired assistant reply from a specified chat assistant session.
+
+#### Request
+
+- Method: DELETE
+- URL: `/api/v1/chats/{chat_id}/sessions/{session_id}/messages/{msg_id}`
+- Headers:
+ - `'Authorization: Bearer '`
+
+##### Request example
+
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/chats/{chat_id}/sessions/{session_id}/messages/{msg_id} \
+ --header 'Authorization: Bearer '
+```
+
+##### Request Parameters
+
+- `chat_id`: (*Path parameter*)
+ The ID of the associated chat assistant.
+- `session_id`: (*Path parameter*)
+ The ID of the session that owns the message.
+- `msg_id`: (*Path parameter*)
+ The ID of the message to delete.
+
+#### Response
+
+Success: returns the updated session object.
+
+```json
+{
+ "code": 0,
+ "data": {
+ "chat_id": "2ca4b22e878011ef88fe0242ac120005",
+ "id": "4606b4ec87ad11efbc4f0242ac120006",
+ "messages": [],
+ "reference": []
+ }
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "Session not found!"
+}
+```
+
+---
+
+### Update message feedback in a chat assistant's session
+
+**PUT** `/api/v1/chats/{chat_id}/sessions/{session_id}/messages/{msg_id}/feedback`
+
+Updates feedback for an assistant message in a specified chat assistant session.
+
+#### Request
+
+- Method: PUT
+- URL: `/api/v1/chats/{chat_id}/sessions/{session_id}/messages/{msg_id}/feedback`
+- Headers:
+ - `'Content-Type: application/json'`
+ - `'Authorization: Bearer '`
+- Body:
+ - `"thumbup"`: `boolean`
+ - `"feedback"`: `string` (optional)
+
+##### Request example
+
+```bash
+curl --request PUT \
+ --url http://{address}/api/v1/chats/{chat_id}/sessions/{session_id}/messages/{msg_id}/feedback \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "thumbup": false,
+ "feedback": "The answer missed the cited document."
+ }'
+```
+
+##### Request Parameters
+
+- `chat_id`: (*Path parameter*)
+ The ID of the associated chat assistant.
+- `session_id`: (*Path parameter*)
+ The ID of the session that owns the message.
+- `msg_id`: (*Path parameter*)
+ The ID of the assistant message to update.
+- `"thumbup"`: (*Body parameter*), `boolean`
+ Whether the assistant message is marked as positive feedback.
+- `"feedback"`: (*Body parameter*), `string`
+ Optional feedback text, typically used when `"thumbup"` is `false`.
+
+#### Response
+
+Success: returns the updated session object.
+
+```json
+{
+ "code": 0,
+ "data": {
+ "chat_id": "2ca4b22e878011ef88fe0242ac120005",
+ "id": "4606b4ec87ad11efbc4f0242ac120006",
+ "messages": [
+ {
+ "id": "message-id",
+ "role": "assistant",
+ "content": "Here is the answer.",
+ "thumbup": false,
+ "feedback": "The answer missed the cited document."
+ }
+ ]
+ }
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "Session not found!"
+}
+```
+
+---
+
### Delete chat assistant's sessions
**DELETE** `/api/v1/chats/{chat_id}/sessions`
@@ -3155,6 +3836,7 @@ Deletes sessions of a chat assistant by ID.
- `'Authorization: Bearer '`
- Body:
- `"ids"`: `list[string]`
+ - `"delete_all"`: `boolean`
##### Request example
@@ -3169,12 +3851,26 @@ curl --request DELETE \
}'
```
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/chats/{chat_id}/sessions \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "delete_all": true
+ }'
+```
+
##### Request Parameters
- `chat_id`: (*Path parameter*)
The ID of the associated chat assistant.
- `"ids"`: (*Body Parameter*), `list[string]`
- The IDs of the sessions to delete. If it is not specified, all sessions associated with the specified chat assistant will be deleted.
+ The IDs of the sessions to delete.
+ - If omitted, or set to `null` or an empty array, no sessions are deleted.
+ - If an array of IDs is provided, only the sessions matching those IDs are deleted.
+- `"delete_all"`: (*Body Parameter*), `boolean`
+ Whether to delete all sessions of the specified chat assistant when `"ids"` is omitted, or set to `null` or an empty array. Defaults to `false`.
#### Response
@@ -3681,15 +4377,16 @@ Asks a specified agent a question to start an AI-powered conversation.
- `"session_id"`: `string` (optional)
- `"inputs"`: `object` (optional)
- `"user_id"`: `string` (optional)
- - `"return_trace"`: `boolean` (optional, default `false`) — include execution trace logs.
+ - `"return_trace"`: `boolean` (optional, default `false`) — whether to include execution trace logs. See the `node_finished` event.
+ - `"release"`: `boolean` (optional, default `false`) - whether to visit the latest published canvas.
#### Streaming events to handle
-When `stream=true`, the server sends Server-Sent Events (SSE). Clients should handle these `event` types:
+When `stream=true`, the server sends Server-Sent Events (SSE). A client should handle these events:
-- `message`: streaming content from Message components.
-- `message_end`: end of a Message component; may include `reference`/`attachment`.
-- `node_finished`: a component finishes; `data.inputs/outputs/error/elapsed_time` describe the node result. If `return_trace=true`, the trace is attached inside the same `node_finished` event (`data.trace`).
+- `message`: Streaming content from the **Message** components.
+- `message_end`: End of a **Message** component, which may include `reference`/`attachment`.
+- `node_finished`: A component finishes; `data.inputs/outputs/error/elapsed_time` describes the node result. If a component produces structured output, read it from that component's `data.outputs.structured`. If `return_trace=true`, the trace is attached inside the same `node_finished` event (`data.trace`).
The stream terminates with `[DONE]`.
@@ -3969,6 +4666,8 @@ When `extra_body.reference_metadata.include` is `true`, each reference chunk may
Non-stream:
+If one or more components produce structured output, ensure you set `return_trace=true` and check each component's structured output via `trace`. The top-level `data.structured` field is a shortcut aggregated by `component_id`.
+
```json
{
"code": 0,
@@ -4497,55 +5196,220 @@ Failure:
```json
{
"code": 102,
- "message": "You don't own the agent ccd2f856b12311ef94ca0242ac1200052."
+ "message": "You don't own the agent ccd2f856b12311ef94ca0242ac1200052."
+}
+```
+
+---
+
+### Delete agent's sessions
+
+**DELETE** `/api/v1/agents/{agent_id}/sessions`
+
+Deletes sessions of an agent by ID.
+
+#### Request
+
+- Method: DELETE
+- URL: `/api/v1/agents/{agent_id}/sessions`
+- Headers:
+ - `'content-Type: application/json'`
+ - `'Authorization: Bearer '`
+- Body:
+ - `"ids"`: `list[string]`
+ - `"delete_all"`: `boolean`
+
+##### Request example
+
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/agents/{agent_id}/sessions \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '
+ {
+ "ids": ["test_1", "test_2"]
+ }'
+```
+
+```bash
+curl --request DELETE \
+ --url http://{address}/api/v1/agents/{agent_id}/sessions \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "delete_all": true
+ }'
+```
+
+##### Request Parameters
+
+- `agent_id`: (*Path parameter*)
+ The ID of the associated agent.
+- `"ids"`: (*Body Parameter*), `list[string]`
+ The IDs of the sessions to delete.
+ - If omitted, or set to `null` or an empty array, no sessions are deleted.
+ - If an array of IDs is provided, only the sessions matching those IDs are deleted.
+- `"delete_all"`: (*Body Parameter*), `boolean`
+ Whether to delete all sessions of the specified agent when `"ids"` is omitted, or set to `null` or an empty array. Defaults to `false`.
+
+#### Response
+
+Success:
+
+```json
+{
+ "code": 0
+}
+```
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "The agent doesn't own the session cbd31e52f73911ef93b232903b842af6"
+}
+```
+
+---
+
+### Text-to-speech
+
+**POST** `/api/v1/chats/tts`
+
+Converts text to speech audio using the tenant's default TTS model, returning a streaming audio response.
+
+#### Request
+
+- Method: POST
+- URL: `/api/v1/chats/tts`
+- Headers:
+ - `'Content-Type: application/json'`
+ - `'Authorization: Bearer '`
+- Body:
+ - `"text"`: `string` *(Required)* The text to synthesize.
+
+##### Request example
+
+```bash
+curl --request POST \
+ --url http://{address}/api/v1/chats/tts \
+ --header 'Content-Type: application/json' \
+ --header 'Authorization: Bearer ' \
+ --output audio.mp3 \
+ --data '{"text": "Hello, how can I help you today?"}'
+```
+
+#### Response
+
+Success: binary `audio/mpeg` stream with headers `Cache-Control: no-cache`, `Connection: keep-alive`, `X-Accel-Buffering: no`.
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "No default TTS model is set"
+}
+```
+
+---
+
+### Speech-to-text
+
+**POST** `/api/v1/chats/transcriptions`
+
+Transcribes an audio file using the tenant's default ASR (automatic speech recognition) model.
+
+#### Request
+
+- Method: POST
+- URL: `/api/v1/chats/transcriptions`
+- Headers:
+ - `'Authorization: Bearer '`
+- Body (multipart/form-data):
+ - `"file"`: audio file (`.wav`, `.mp3`, `.m4a`, `.aac`, `.flac`, `.ogg`, `.webm`, `.opus`, `.wma`)
+ - `"stream"`: `string` `"true"` for SSE streaming, `"false"` (default) for a single JSON response.
+
+##### Request example
+
+```bash
+curl --request POST \
+ --url http://{address}/api/v1/chats/transcriptions \
+ --header 'Authorization: Bearer ' \
+ --form file=@recording.wav \
+ --form stream=false
+```
+
+#### Response
+
+Success (non-streaming):
+
+```json
+{
+ "code": 0,
+ "data": {
+ "text": "Hello, how can I help you today?"
+ }
+}
+```
+
+Success (streaming): SSE events with `data: {"event": "partial", "text": "..."}`.
+
+Failure:
+
+```json
+{
+ "code": 102,
+ "message": "Unsupported audio format: .mp4. Allowed: .aac, .flac, .m4a, .mp3, .ogg, .opus, .wav, .webm, .wma"
}
```
---
-### Delete agent's sessions
+### Generate mind map
-**DELETE** `/api/v1/agents/{agent_id}/sessions`
+**POST** `/api/v1/chats/mindmap`
-Deletes sessions of an agent by ID.
+Generates a mind map from a question and a set of knowledge base IDs.
#### Request
-- Method: DELETE
-- URL: `/api/v1/agents/{agent_id}/sessions`
+- Method: POST
+- URL: `/api/v1/chats/mindmap`
- Headers:
- - `'content-Type: application/json'`
- - `'Authorization: Bearer '`
+ - `'Content-Type: application/json'`
+ - `'Authorization: Bearer '`
- Body:
- - `"ids"`: `list[string]`
+ - `"question"`: `string` *(Required)* The central question or topic.
+ - `"kb_ids"`: `list[string]` *(Required)* Knowledge base IDs to search.
+ - `"search_id"`: `string` *(Optional)* ID of a saved search configuration to merge additional `kb_ids` and settings.
##### Request example
```bash
-curl --request DELETE \
- --url http://{address}/api/v1/agents/{agent_id}/sessions \
+curl --request POST \
+ --url http://{address}/api/v1/chats/mindmap \
--header 'Content-Type: application/json' \
- --header 'Authorization: Bearer ' \
- --data '
- {
- "ids": ["test_1", "test_2"]
+ --header 'Authorization: Bearer ' \
+ --data '{
+ "question": "What is retrieval-augmented generation?",
+ "kb_ids": ["kb-abc123"]
}'
```
-##### Request Parameters
-
-- `agent_id`: (*Path parameter*)
- The ID of the associated agent.
-- `"ids"`: (*Body Parameter*), `list[string]`
- The IDs of the sessions to delete. If it is not specified, all sessions associated with the specified agent will be deleted.
-
#### Response
Success:
```json
{
- "code": 0
+ "code": 0,
+ "data": {
+ "name": "Retrieval-Augmented Generation",
+ "children": [...]
+ }
}
```
@@ -4553,8 +5417,8 @@ Failure:
```json
{
- "code": 102,
- "message": "The agent doesn't own the session cbd31e52f73911ef93b232903b842af6"
+ "code": 500,
+ "message": "..."
}
```
@@ -4562,7 +5426,7 @@ Failure:
### Generate related questions
-**POST** `/api/v1/sessions/related_questions`
+**POST** `/api/v1/chats/related_questions`
Generates five to ten alternative question strings from the user's original query to retrieve more relevant search results.
@@ -4577,25 +5441,23 @@ The chat model autonomously determines the number of questions to generate based
#### Request
- Method: POST
-- URL: `/api/v1/sessions/related_questions`
+- URL: `/api/v1/chats/related_questions`
- Headers:
- `'content-Type: application/json'`
- `'Authorization: Bearer


-DFE0E5) +
+  +
+  +
+ 





 @@ -85,6 +88,7 @@ Try our demo at [https://demo.ragflow.io](https://demo.ragflow.io).
## 🔥 Latest Updates
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — Provides an official skill for accessing RAGFlow datasets via OpenClaw.
- 2025-12-26 Supports 'Memory' for AI agent.
- 2025-11-19 Supports Gemini 3 Pro.
- 2025-11-12 Supports data synchronization from Confluence, S3, Notion, Discord, Google Drive.
@@ -188,12 +192,12 @@ releases! 🌟
> All Docker images are built for x86 platforms. We don't currently offer Docker images for ARM64.
> If you are on an ARM64 platform, follow [this guide](https://ragflow.io/docs/dev/build_docker_image) to build a Docker image compatible with your system.
-> The command below downloads the `v0.24.0` edition of the RAGFlow Docker image. See the following table for descriptions of different RAGFlow editions. To download a RAGFlow edition different from `v0.24.0`, update the `RAGFLOW_IMAGE` variable accordingly in **docker/.env** before using `docker compose` to start the server.
+> The command below downloads the `v0.25.0` edition of the RAGFlow Docker image. See the following table for descriptions of different RAGFlow editions. To download a RAGFlow edition different from `v0.25.0`, update the `RAGFLOW_IMAGE` variable accordingly in **docker/.env** before using `docker compose` to start the server.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Optional: use a stable tag (see releases: https://github.com/infiniflow/ragflow/releases)
# This step ensures the **entrypoint.sh** file in the code matches the Docker image version.
@@ -325,7 +329,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Launch the dependent services (MinIO, Elasticsearch, Redis, and MySQL) using Docker Compose:
@@ -389,8 +393,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_ar.md b/README_ar.md
new file mode 100644
index 00000000000..d03fa2a1eee
--- /dev/null
+++ b/README_ar.md
@@ -0,0 +1,414 @@
+
@@ -85,6 +88,7 @@ Try our demo at [https://demo.ragflow.io](https://demo.ragflow.io).
## 🔥 Latest Updates
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — Provides an official skill for accessing RAGFlow datasets via OpenClaw.
- 2025-12-26 Supports 'Memory' for AI agent.
- 2025-11-19 Supports Gemini 3 Pro.
- 2025-11-12 Supports data synchronization from Confluence, S3, Notion, Discord, Google Drive.
@@ -188,12 +192,12 @@ releases! 🌟
> All Docker images are built for x86 platforms. We don't currently offer Docker images for ARM64.
> If you are on an ARM64 platform, follow [this guide](https://ragflow.io/docs/dev/build_docker_image) to build a Docker image compatible with your system.
-> The command below downloads the `v0.24.0` edition of the RAGFlow Docker image. See the following table for descriptions of different RAGFlow editions. To download a RAGFlow edition different from `v0.24.0`, update the `RAGFLOW_IMAGE` variable accordingly in **docker/.env** before using `docker compose` to start the server.
+> The command below downloads the `v0.25.0` edition of the RAGFlow Docker image. See the following table for descriptions of different RAGFlow editions. To download a RAGFlow edition different from `v0.25.0`, update the `RAGFLOW_IMAGE` variable accordingly in **docker/.env** before using `docker compose` to start the server.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Optional: use a stable tag (see releases: https://github.com/infiniflow/ragflow/releases)
# This step ensures the **entrypoint.sh** file in the code matches the Docker image version.
@@ -325,7 +329,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Launch the dependent services (MinIO, Elasticsearch, Redis, and MySQL) using Docker Compose:
@@ -389,8 +393,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_ar.md b/README_ar.md
new file mode 100644
index 00000000000..d03fa2a1eee
--- /dev/null
+++ b/README_ar.md
@@ -0,0 +1,414 @@
+



+
+
+
+
+
 +
+ +
+ +
+



+
+
+
+
+
 +
+ +
+ +
+

-DFE0E5) +
+  +
+  +
+ 





 @@ -85,6 +88,7 @@ Coba demo kami di [https://demo.ragflow.io](https://demo.ragflow.io).
## 🔥 Pembaruan Terbaru
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — Menyediakan skill resmi untuk mengakses dataset RAGFlow melalui OpenClaw.
- 2025-12-26 Mendukung 'Memori' untuk agen AI.
- 2025-11-19 Mendukung Gemini 3 Pro.
- 2025-11-12 Mendukung sinkronisasi data dari Confluence, S3, Notion, Discord, Google Drive.
@@ -188,12 +192,12 @@ Coba demo kami di [https://demo.ragflow.io](https://demo.ragflow.io).
> Semua gambar Docker dibangun untuk platform x86. Saat ini, kami tidak menawarkan gambar Docker untuk ARM64.
> Jika Anda menggunakan platform ARM64, [silakan gunakan panduan ini untuk membangun gambar Docker yang kompatibel dengan sistem Anda](https://ragflow.io/docs/dev/build_docker_image).
-> Perintah di bawah ini mengunduh edisi v0.24.0 dari gambar Docker RAGFlow. Silakan merujuk ke tabel berikut untuk deskripsi berbagai edisi RAGFlow. Untuk mengunduh edisi RAGFlow yang berbeda dari v0.24.0, perbarui variabel RAGFLOW_IMAGE di docker/.env sebelum menggunakan docker compose untuk memulai server.
+> Perintah di bawah ini mengunduh edisi v0.25.0 dari gambar Docker RAGFlow. Silakan merujuk ke tabel berikut untuk deskripsi berbagai edisi RAGFlow. Untuk mengunduh edisi RAGFlow yang berbeda dari v0.25.0, perbarui variabel RAGFLOW_IMAGE di docker/.env sebelum menggunakan docker compose untuk memulai server.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Opsional: gunakan tag stabil (lihat releases: https://github.com/infiniflow/ragflow/releases)
# This steps ensures the **entrypoint.sh** file in the code matches the Docker image version.
@@ -299,7 +303,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Jalankan aplikasi yang diperlukan (MinIO, Elasticsearch, Redis, dan MySQL) menggunakan Docker Compose:
@@ -361,8 +365,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_ja.md b/README_ja.md
index afff19bc8fd..84f42b05876 100644
--- a/README_ja.md
+++ b/README_ja.md
@@ -1,5 +1,5 @@
@@ -85,6 +88,7 @@ Coba demo kami di [https://demo.ragflow.io](https://demo.ragflow.io).
## 🔥 Pembaruan Terbaru
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — Menyediakan skill resmi untuk mengakses dataset RAGFlow melalui OpenClaw.
- 2025-12-26 Mendukung 'Memori' untuk agen AI.
- 2025-11-19 Mendukung Gemini 3 Pro.
- 2025-11-12 Mendukung sinkronisasi data dari Confluence, S3, Notion, Discord, Google Drive.
@@ -188,12 +192,12 @@ Coba demo kami di [https://demo.ragflow.io](https://demo.ragflow.io).
> Semua gambar Docker dibangun untuk platform x86. Saat ini, kami tidak menawarkan gambar Docker untuk ARM64.
> Jika Anda menggunakan platform ARM64, [silakan gunakan panduan ini untuk membangun gambar Docker yang kompatibel dengan sistem Anda](https://ragflow.io/docs/dev/build_docker_image).
-> Perintah di bawah ini mengunduh edisi v0.24.0 dari gambar Docker RAGFlow. Silakan merujuk ke tabel berikut untuk deskripsi berbagai edisi RAGFlow. Untuk mengunduh edisi RAGFlow yang berbeda dari v0.24.0, perbarui variabel RAGFLOW_IMAGE di docker/.env sebelum menggunakan docker compose untuk memulai server.
+> Perintah di bawah ini mengunduh edisi v0.25.0 dari gambar Docker RAGFlow. Silakan merujuk ke tabel berikut untuk deskripsi berbagai edisi RAGFlow. Untuk mengunduh edisi RAGFlow yang berbeda dari v0.25.0, perbarui variabel RAGFLOW_IMAGE di docker/.env sebelum menggunakan docker compose untuk memulai server.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Opsional: gunakan tag stabil (lihat releases: https://github.com/infiniflow/ragflow/releases)
# This steps ensures the **entrypoint.sh** file in the code matches the Docker image version.
@@ -299,7 +303,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Jalankan aplikasi yang diperlukan (MinIO, Elasticsearch, Redis, dan MySQL) menggunakan Docker Compose:
@@ -361,8 +365,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_ja.md b/README_ja.md
index afff19bc8fd..84f42b05876 100644
--- a/README_ja.md
+++ b/README_ja.md
@@ -1,5 +1,5 @@


-DFE0E5) +
+  +
+  +
+ 





 @@ -66,6 +69,7 @@
## 🔥 最新情報
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — OpenClaw経由でRAGFlowデータセットにアクセスする公式スキルを提供。
- 2025-12-26 AIエージェントの「メモリ」機能をサポート。
- 2025-11-19 Gemini 3 Proをサポートしています。
- 2025-11-12 Confluence、S3、Notion、Discord、Google Drive からのデータ同期をサポートします。
@@ -168,12 +172,12 @@
> 現在、公式に提供されているすべての Docker イメージは x86 アーキテクチャ向けにビルドされており、ARM64 用の Docker イメージは提供されていません。
> ARM64 アーキテクチャのオペレーティングシステムを使用している場合は、[このドキュメント](https://ragflow.io/docs/dev/build_docker_image)を参照して Docker イメージを自分でビルドしてください。
-> 以下のコマンドは、RAGFlow Docker イメージの v0.24.0 エディションをダウンロードします。異なる RAGFlow エディションの説明については、以下の表を参照してください。v0.24.0 とは異なるエディションをダウンロードするには、docker/.env ファイルの RAGFLOW_IMAGE 変数を適宜更新し、docker compose を使用してサーバーを起動してください。
+> 以下のコマンドは、RAGFlow Docker イメージの v0.25.0 エディションをダウンロードします。異なる RAGFlow エディションの説明については、以下の表を参照してください。v0.25.0 とは異なるエディションをダウンロードするには、docker/.env ファイルの RAGFLOW_IMAGE 変数を適宜更新し、docker compose を使用してサーバーを起動してください。
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# 任意: 安定版タグを利用 (一覧: https://github.com/infiniflow/ragflow/releases)
# この手順は、コード内の entrypoint.sh ファイルが Docker イメージのバージョンと一致していることを確認します。
@@ -299,7 +303,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Docker Compose を使用して依存サービス(MinIO、Elasticsearch、Redis、MySQL)を起動する:
@@ -361,8 +365,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_ko.md b/README_ko.md
index 91978a72a5d..578e247e9fa 100644
--- a/README_ko.md
+++ b/README_ko.md
@@ -1,5 +1,5 @@
@@ -66,6 +69,7 @@
## 🔥 最新情報
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — OpenClaw経由でRAGFlowデータセットにアクセスする公式スキルを提供。
- 2025-12-26 AIエージェントの「メモリ」機能をサポート。
- 2025-11-19 Gemini 3 Proをサポートしています。
- 2025-11-12 Confluence、S3、Notion、Discord、Google Drive からのデータ同期をサポートします。
@@ -168,12 +172,12 @@
> 現在、公式に提供されているすべての Docker イメージは x86 アーキテクチャ向けにビルドされており、ARM64 用の Docker イメージは提供されていません。
> ARM64 アーキテクチャのオペレーティングシステムを使用している場合は、[このドキュメント](https://ragflow.io/docs/dev/build_docker_image)を参照して Docker イメージを自分でビルドしてください。
-> 以下のコマンドは、RAGFlow Docker イメージの v0.24.0 エディションをダウンロードします。異なる RAGFlow エディションの説明については、以下の表を参照してください。v0.24.0 とは異なるエディションをダウンロードするには、docker/.env ファイルの RAGFLOW_IMAGE 変数を適宜更新し、docker compose を使用してサーバーを起動してください。
+> 以下のコマンドは、RAGFlow Docker イメージの v0.25.0 エディションをダウンロードします。異なる RAGFlow エディションの説明については、以下の表を参照してください。v0.25.0 とは異なるエディションをダウンロードするには、docker/.env ファイルの RAGFLOW_IMAGE 変数を適宜更新し、docker compose を使用してサーバーを起動してください。
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# 任意: 安定版タグを利用 (一覧: https://github.com/infiniflow/ragflow/releases)
# この手順は、コード内の entrypoint.sh ファイルが Docker イメージのバージョンと一致していることを確認します。
@@ -299,7 +303,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Docker Compose を使用して依存サービス(MinIO、Elasticsearch、Redis、MySQL)を起動する:
@@ -361,8 +365,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_ko.md b/README_ko.md
index 91978a72a5d..578e247e9fa 100644
--- a/README_ko.md
+++ b/README_ko.md
@@ -1,5 +1,5 @@


-DFE0E5) +
+  +
+  +
+ 





 @@ -67,6 +70,7 @@
## 🔥 업데이트
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — OpenClaw를 통해 RAGFlow 데이터셋에 접근하는 공식 스킬 제공.
- 2025-12-26 AI 에이전트의 '메모리' 기능 지원.
- 2025-11-19 Gemini 3 Pro를 지원합니다.
- 2025-11-12 Confluence, S3, Notion, Discord, Google Drive에서 데이터 동기화를 지원합니다.
@@ -170,12 +174,12 @@
> 모든 Docker 이미지는 x86 플랫폼을 위해 빌드되었습니다. 우리는 현재 ARM64 플랫폼을 위한 Docker 이미지를 제공하지 않습니다.
> ARM64 플랫폼을 사용 중이라면, [시스템과 호환되는 Docker 이미지를 빌드하려면 이 가이드를 사용해 주세요](https://ragflow.io/docs/dev/build_docker_image).
- > 아래 명령어는 RAGFlow Docker 이미지의 v0.24.0 버전을 다운로드합니다. 다양한 RAGFlow 버전에 대한 설명은 다음 표를 참조하십시오. v0.24.0과 다른 RAGFlow 버전을 다운로드하려면, docker/.env 파일에서 RAGFLOW_IMAGE 변수를 적절히 업데이트한 후 docker compose를 사용하여 서버를 시작하십시오.
+ > 아래 명령어는 RAGFlow Docker 이미지의 v0.25.0 버전을 다운로드합니다. 다양한 RAGFlow 버전에 대한 설명은 다음 표를 참조하십시오. v0.25.0과 다른 RAGFlow 버전을 다운로드하려면, docker/.env 파일에서 RAGFLOW_IMAGE 변수를 적절히 업데이트한 후 docker compose를 사용하여 서버를 시작하십시오.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Optional: use a stable tag (see releases: https://github.com/infiniflow/ragflow/releases)
# 이 단계는 코드의 entrypoint.sh 파일이 Docker 이미지 버전과 일치하도록 보장합니다.
@@ -294,7 +298,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
@@ -365,8 +369,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_pt_br.md b/README_pt_br.md
index 8fa5b6692e1..88f34b19532 100644
--- a/README_pt_br.md
+++ b/README_pt_br.md
@@ -1,5 +1,5 @@
@@ -67,6 +70,7 @@
## 🔥 업데이트
+- 2026-03-24 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — OpenClaw를 통해 RAGFlow 데이터셋에 접근하는 공식 스킬 제공.
- 2025-12-26 AI 에이전트의 '메모리' 기능 지원.
- 2025-11-19 Gemini 3 Pro를 지원합니다.
- 2025-11-12 Confluence, S3, Notion, Discord, Google Drive에서 데이터 동기화를 지원합니다.
@@ -170,12 +174,12 @@
> 모든 Docker 이미지는 x86 플랫폼을 위해 빌드되었습니다. 우리는 현재 ARM64 플랫폼을 위한 Docker 이미지를 제공하지 않습니다.
> ARM64 플랫폼을 사용 중이라면, [시스템과 호환되는 Docker 이미지를 빌드하려면 이 가이드를 사용해 주세요](https://ragflow.io/docs/dev/build_docker_image).
- > 아래 명령어는 RAGFlow Docker 이미지의 v0.24.0 버전을 다운로드합니다. 다양한 RAGFlow 버전에 대한 설명은 다음 표를 참조하십시오. v0.24.0과 다른 RAGFlow 버전을 다운로드하려면, docker/.env 파일에서 RAGFLOW_IMAGE 변수를 적절히 업데이트한 후 docker compose를 사용하여 서버를 시작하십시오.
+ > 아래 명령어는 RAGFlow Docker 이미지의 v0.25.0 버전을 다운로드합니다. 다양한 RAGFlow 버전에 대한 설명은 다음 표를 참조하십시오. v0.25.0과 다른 RAGFlow 버전을 다운로드하려면, docker/.env 파일에서 RAGFLOW_IMAGE 변수를 적절히 업데이트한 후 docker compose를 사용하여 서버를 시작하십시오.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Optional: use a stable tag (see releases: https://github.com/infiniflow/ragflow/releases)
# 이 단계는 코드의 entrypoint.sh 파일이 Docker 이미지 버전과 일치하도록 보장합니다.
@@ -294,7 +298,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
@@ -365,8 +369,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_pt_br.md b/README_pt_br.md
index 8fa5b6692e1..88f34b19532 100644
--- a/README_pt_br.md
+++ b/README_pt_br.md
@@ -1,5 +1,5 @@


-DBEDFA) +
+  +
+  +
+ 





 @@ -86,6 +89,7 @@ Experimente nossa demo em [https://demo.ragflow.io](https://demo.ragflow.io).
## 🔥 Últimas Atualizações
+- 24-03-2026 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — Fornece um skill oficial para acessar datasets do RAGFlow via OpenClaw.
- 26-12-2025 Suporte à função 'Memória' para agentes de IA.
- 19-11-2025 Suporta Gemini 3 Pro.
- 12-11-2025 Suporta a sincronização de dados do Confluence, S3, Notion, Discord e Google Drive.
@@ -188,12 +192,12 @@ Experimente nossa demo em [https://demo.ragflow.io](https://demo.ragflow.io).
> Todas as imagens Docker são construídas para plataformas x86. Atualmente, não oferecemos imagens Docker para ARM64.
> Se você estiver usando uma plataforma ARM64, por favor, utilize [este guia](https://ragflow.io/docs/dev/build_docker_image) para construir uma imagem Docker compatível com o seu sistema.
- > O comando abaixo baixa a edição`v0.24.0` da imagem Docker do RAGFlow. Consulte a tabela a seguir para descrições de diferentes edições do RAGFlow. Para baixar uma edição do RAGFlow diferente da `v0.24.0`, atualize a variável `RAGFLOW_IMAGE` conforme necessário no **docker/.env** antes de usar `docker compose` para iniciar o servidor.
+ > O comando abaixo baixa a edição`v0.25.0` da imagem Docker do RAGFlow. Consulte a tabela a seguir para descrições de diferentes edições do RAGFlow. Para baixar uma edição do RAGFlow diferente da `v0.25.0`, atualize a variável `RAGFLOW_IMAGE` conforme necessário no **docker/.env** antes de usar `docker compose` para iniciar o servidor.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Opcional: use uma tag estável (veja releases: https://github.com/infiniflow/ragflow/releases)
# Esta etapa garante que o arquivo entrypoint.sh no código corresponda à versão da imagem do Docker.
@@ -316,7 +320,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # instala os módulos Python dependentes do RAGFlow
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Inicie os serviços dependentes (MinIO, Elasticsearch, Redis e MySQL) usando Docker Compose:
@@ -378,8 +382,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_tr.md b/README_tr.md
new file mode 100644
index 00000000000..89be2c0d790
--- /dev/null
+++ b/README_tr.md
@@ -0,0 +1,409 @@
+
@@ -86,6 +89,7 @@ Experimente nossa demo em [https://demo.ragflow.io](https://demo.ragflow.io).
## 🔥 Últimas Atualizações
+- 24-03-2026 [RAGFlow Skill on OpenClaw](https://clawhub.ai/yingfeng/ragflow-skill) — Fornece um skill oficial para acessar datasets do RAGFlow via OpenClaw.
- 26-12-2025 Suporte à função 'Memória' para agentes de IA.
- 19-11-2025 Suporta Gemini 3 Pro.
- 12-11-2025 Suporta a sincronização de dados do Confluence, S3, Notion, Discord e Google Drive.
@@ -188,12 +192,12 @@ Experimente nossa demo em [https://demo.ragflow.io](https://demo.ragflow.io).
> Todas as imagens Docker são construídas para plataformas x86. Atualmente, não oferecemos imagens Docker para ARM64.
> Se você estiver usando uma plataforma ARM64, por favor, utilize [este guia](https://ragflow.io/docs/dev/build_docker_image) para construir uma imagem Docker compatível com o seu sistema.
- > O comando abaixo baixa a edição`v0.24.0` da imagem Docker do RAGFlow. Consulte a tabela a seguir para descrições de diferentes edições do RAGFlow. Para baixar uma edição do RAGFlow diferente da `v0.24.0`, atualize a variável `RAGFLOW_IMAGE` conforme necessário no **docker/.env** antes de usar `docker compose` para iniciar o servidor.
+ > O comando abaixo baixa a edição`v0.25.0` da imagem Docker do RAGFlow. Consulte a tabela a seguir para descrições de diferentes edições do RAGFlow. Para baixar uma edição do RAGFlow diferente da `v0.25.0`, atualize a variável `RAGFLOW_IMAGE` conforme necessário no **docker/.env** antes de usar `docker compose` para iniciar o servidor.
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# Opcional: use uma tag estável (veja releases: https://github.com/infiniflow/ragflow/releases)
# Esta etapa garante que o arquivo entrypoint.sh no código corresponda à versão da imagem do Docker.
@@ -316,7 +320,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # instala os módulos Python dependentes do RAGFlow
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. Inicie os serviços dependentes (MinIO, Elasticsearch, Redis e MySQL) usando Docker Compose:
@@ -378,8 +382,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_tr.md b/README_tr.md
new file mode 100644
index 00000000000..89be2c0d790
--- /dev/null
+++ b/README_tr.md
@@ -0,0 +1,409 @@
+



+
+
+
+
+
 +
+ +
+ +
+

-DFE0E5) +
+  +
+  +
+ 





 @@ -85,6 +88,7 @@
## 🔥 近期更新
+- 2026-03-24 發布 [RAGFlow 官方 Skill](https://clawhub.ai/yingfeng/ragflow-skill) — 提供官方 Skill 以透過 OpenClaw 訪問 RAGFlow 數據集。
- 2025-12-26 支援AI代理的「記憶」功能。
- 2025-11-19 支援 Gemini 3 Pro。
- 2025-11-12 支援從 Confluence、S3、Notion、Discord、Google Drive 進行資料同步。
@@ -187,12 +191,12 @@
> 所有 Docker 映像檔都是為 x86 平台建置的。目前,我們不提供 ARM64 平台的 Docker 映像檔。
> 如果您使用的是 ARM64 平台,請使用 [這份指南](https://ragflow.io/docs/dev/build_docker_image) 來建置適合您系統的 Docker 映像檔。
-> 執行以下指令會自動下載 RAGFlow Docker 映像 `v0.24.0`。請參考下表查看不同 Docker 發行版的說明。如需下載不同於 `v0.24.0` 的 Docker 映像,請在執行 `docker compose` 啟動服務之前先更新 **docker/.env** 檔案內的 `RAGFLOW_IMAGE` 變數。
+> 執行以下指令會自動下載 RAGFlow Docker 映像 `v0.25.0`。請參考下表查看不同 Docker 發行版的說明。如需下載不同於 `v0.25.0` 的 Docker 映像,請在執行 `docker compose` 啟動服務之前先更新 **docker/.env** 檔案內的 `RAGFLOW_IMAGE` 變數。
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# 可選:使用穩定版標籤(查看發佈:https://github.com/infiniflow/ragflow/releases)
# 此步驟確保程式碼中的 entrypoint.sh 檔案與 Docker 映像版本一致。
@@ -326,7 +330,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. 透過 Docker Compose 啟動依賴的服務(MinIO, Elasticsearch, Redis, and MySQL):
@@ -392,8 +396,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_zh.md b/README_zh.md
index 5b194daa0ff..473794a934f 100644
--- a/README_zh.md
+++ b/README_zh.md
@@ -1,5 +1,5 @@
@@ -85,6 +88,7 @@
## 🔥 近期更新
+- 2026-03-24 發布 [RAGFlow 官方 Skill](https://clawhub.ai/yingfeng/ragflow-skill) — 提供官方 Skill 以透過 OpenClaw 訪問 RAGFlow 數據集。
- 2025-12-26 支援AI代理的「記憶」功能。
- 2025-11-19 支援 Gemini 3 Pro。
- 2025-11-12 支援從 Confluence、S3、Notion、Discord、Google Drive 進行資料同步。
@@ -187,12 +191,12 @@
> 所有 Docker 映像檔都是為 x86 平台建置的。目前,我們不提供 ARM64 平台的 Docker 映像檔。
> 如果您使用的是 ARM64 平台,請使用 [這份指南](https://ragflow.io/docs/dev/build_docker_image) 來建置適合您系統的 Docker 映像檔。
-> 執行以下指令會自動下載 RAGFlow Docker 映像 `v0.24.0`。請參考下表查看不同 Docker 發行版的說明。如需下載不同於 `v0.24.0` 的 Docker 映像,請在執行 `docker compose` 啟動服務之前先更新 **docker/.env** 檔案內的 `RAGFLOW_IMAGE` 變數。
+> 執行以下指令會自動下載 RAGFlow Docker 映像 `v0.25.0`。請參考下表查看不同 Docker 發行版的說明。如需下載不同於 `v0.25.0` 的 Docker 映像,請在執行 `docker compose` 啟動服務之前先更新 **docker/.env** 檔案內的 `RAGFLOW_IMAGE` 變數。
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# 可選:使用穩定版標籤(查看發佈:https://github.com/infiniflow/ragflow/releases)
# 此步驟確保程式碼中的 entrypoint.sh 檔案與 Docker 映像版本一致。
@@ -326,7 +330,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
3. 透過 Docker Compose 啟動依賴的服務(MinIO, Elasticsearch, Redis, and MySQL):
@@ -392,8 +396,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/README_zh.md b/README_zh.md
index 5b194daa0ff..473794a934f 100644
--- a/README_zh.md
+++ b/README_zh.md
@@ -1,5 +1,5 @@


-DFE0E5) +
+  +
+  +
+ 





 @@ -85,7 +88,8 @@
## 🔥 近期更新
-- 2025-12-26 支持AI代理的“记忆”功能。
+- 2026-03-24 发布 [RAGFlow 官方 Skill](https://clawhub.ai/yingfeng/ragflow-skill) — 提供官方 Skill 以通过 OpenClaw 访问 RAGFlow 数据集。
+- 2025-12-26 支持AI代理的"记忆"功能。
- 2025-11-19 支持 Gemini 3 Pro。
- 2025-11-12 支持从 Confluence、S3、Notion、Discord、Google Drive 进行数据同步。
- 2025-10-23 支持 MinerU 和 Docling 作为文档解析方法。
@@ -188,12 +192,12 @@
> 请注意,目前官方提供的所有 Docker 镜像均基于 x86 架构构建,并不提供基于 ARM64 的 Docker 镜像。
> 如果你的操作系统是 ARM64 架构,请参考[这篇文档](https://ragflow.io/docs/dev/build_docker_image)自行构建 Docker 镜像。
- > 运行以下命令会自动下载 RAGFlow Docker 镜像 `v0.24.0`。请参考下表查看不同 Docker 发行版的描述。如需下载不同于 `v0.24.0` 的 Docker 镜像,请在运行 `docker compose` 启动服务之前先更新 **docker/.env** 文件内的 `RAGFLOW_IMAGE` 变量。
+ > 运行以下命令会自动下载 RAGFlow Docker 镜像 `v0.25.0`。请参考下表查看不同 Docker 发行版的描述。如需下载不同于 `v0.25.0` 的 Docker 镜像,请在运行 `docker compose` 启动服务之前先更新 **docker/.env** 文件内的 `RAGFLOW_IMAGE` 变量。
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# 可选:使用稳定版本标签(查看发布:https://github.com/infiniflow/ragflow/releases)
# 这一步确保代码中的 entrypoint.sh 文件与 Docker 镜像的版本保持一致。
@@ -326,7 +330,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
@@ -395,8 +399,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/admin/client/COMMAND.md b/admin/client/COMMAND.md
new file mode 100644
index 00000000000..cd8e376c4db
--- /dev/null
+++ b/admin/client/COMMAND.md
@@ -0,0 +1,779 @@
+# RAGFlow CLI User Command Reference
+
+This document describes the user commands available in RAGFlow CLI. All commands must end with a semicolon (`;`).
+
+## Command List
+
+### ping_server
+
+**Description**
+Tests the connection status to the server.
+
+**Usage**
+```
+PING;
+```
+
+**Parameters**
+No parameters.
+
+**Example**
+```
+ragflow> PING;
+```
+
+**Display Effect**
+(Sample output will be provided by the user)
+
+---
+
+### show_current_user
+
+**Description**
+Displays information about the currently logged-in user.
+
+**Usage**
+```
+SHOW CURRENT USER;
+```
+
+**Parameters**
+No parameters.
+
+**Example**
+```
+ragflow> SHOW CURRENT USER;
+```
+
+**Display Effect**
+(Sample output will be provided by the user)
+
+---
+
+### create_model_provider
+
+**Description**
+Creates a new model provider.
+
+**Usage**
+```
+CREATE MODEL PROVIDER
@@ -85,7 +88,8 @@
## 🔥 近期更新
-- 2025-12-26 支持AI代理的“记忆”功能。
+- 2026-03-24 发布 [RAGFlow 官方 Skill](https://clawhub.ai/yingfeng/ragflow-skill) — 提供官方 Skill 以通过 OpenClaw 访问 RAGFlow 数据集。
+- 2025-12-26 支持AI代理的"记忆"功能。
- 2025-11-19 支持 Gemini 3 Pro。
- 2025-11-12 支持从 Confluence、S3、Notion、Discord、Google Drive 进行数据同步。
- 2025-10-23 支持 MinerU 和 Docling 作为文档解析方法。
@@ -188,12 +192,12 @@
> 请注意,目前官方提供的所有 Docker 镜像均基于 x86 架构构建,并不提供基于 ARM64 的 Docker 镜像。
> 如果你的操作系统是 ARM64 架构,请参考[这篇文档](https://ragflow.io/docs/dev/build_docker_image)自行构建 Docker 镜像。
- > 运行以下命令会自动下载 RAGFlow Docker 镜像 `v0.24.0`。请参考下表查看不同 Docker 发行版的描述。如需下载不同于 `v0.24.0` 的 Docker 镜像,请在运行 `docker compose` 启动服务之前先更新 **docker/.env** 文件内的 `RAGFLOW_IMAGE` 变量。
+ > 运行以下命令会自动下载 RAGFlow Docker 镜像 `v0.25.0`。请参考下表查看不同 Docker 发行版的描述。如需下载不同于 `v0.25.0` 的 Docker 镜像,请在运行 `docker compose` 启动服务之前先更新 **docker/.env** 文件内的 `RAGFLOW_IMAGE` 变量。
```bash
$ cd ragflow/docker
- # git checkout v0.24.0
+ # git checkout v0.25.0
# 可选:使用稳定版本标签(查看发布:https://github.com/infiniflow/ragflow/releases)
# 这一步确保代码中的 entrypoint.sh 文件与 Docker 镜像的版本保持一致。
@@ -326,7 +330,7 @@ docker build --platform linux/amd64 \
git clone https://github.com/infiniflow/ragflow.git
cd ragflow/
uv sync --python 3.12 # install RAGFlow dependent python modules
- uv run download_deps.py
+ uv run python3 download_deps.py
pre-commit install
```
@@ -395,8 +399,8 @@ docker build --platform linux/amd64 \
- [Quickstart](https://ragflow.io/docs/dev/)
- [Configuration](https://ragflow.io/docs/dev/configurations)
- [Release notes](https://ragflow.io/docs/dev/release_notes)
-- [User guides](https://ragflow.io/docs/dev/category/guides)
-- [Developer guides](https://ragflow.io/docs/dev/category/developers)
+- [User guides](https://ragflow.io/docs/category/user-guides)
+- [Developer guides](https://ragflow.io/docs/category/developer-guides)
- [References](https://ragflow.io/docs/dev/category/references)
- [FAQs](https://ragflow.io/docs/dev/faq)
diff --git a/admin/client/COMMAND.md b/admin/client/COMMAND.md
new file mode 100644
index 00000000000..cd8e376c4db
--- /dev/null
+++ b/admin/client/COMMAND.md
@@ -0,0 +1,779 @@
+# RAGFlow CLI User Command Reference
+
+This document describes the user commands available in RAGFlow CLI. All commands must end with a semicolon (`;`).
+
+## Command List
+
+### ping_server
+
+**Description**
+Tests the connection status to the server.
+
+**Usage**
+```
+PING;
+```
+
+**Parameters**
+No parameters.
+
+**Example**
+```
+ragflow> PING;
+```
+
+**Display Effect**
+(Sample output will be provided by the user)
+
+---
+
+### show_current_user
+
+**Description**
+Displays information about the currently logged-in user.
+
+**Usage**
+```
+SHOW CURRENT USER;
+```
+
+**Parameters**
+No parameters.
+
+**Example**
+```
+ragflow> SHOW CURRENT USER;
+```
+
+**Display Effect**
+(Sample output will be provided by the user)
+
+---
+
+### create_model_provider
+
+**Description**
+Creates a new model provider.
+
+**Usage**
+```
+CREATE MODEL PROVIDER