diff --git a/projects/website-angular/content/about/digital-preservation.mdx b/projects/website-angular/content/about/digital-preservation.mdx
index 1f42eb4..6422c58 100644
--- a/projects/website-angular/content/about/digital-preservation.mdx
+++ b/projects/website-angular/content/about/digital-preservation.mdx
@@ -1,5 +1,5 @@
---
-title: Digital Preservation Plan
+title: Untitled
category: "about"
---
diff --git a/projects/website-angular/content/about/index.mdx b/projects/website-angular/content/about/index.mdx
deleted file mode 100644
index 291d08b..0000000
--- a/projects/website-angular/content/about/index.mdx
+++ /dev/null
@@ -1,30 +0,0 @@
----
-title: About
-category: about
----
-
-
-## What is Reactome ?
-
-### Mission Statement
-
-REACTOME is an open-source, open access, manually curated and peer-reviewed pathway database. Our goal is to provide intuitive bioinformatics tools for the visualization, interpretation and analysis of pathway knowledge to support basic and clinical research, genome analysis, modeling, systems biology and education. Founded in 2003, the Reactome project is led by Lincoln Stein of [OICR](), Peter D’Eustachio of [NYU Langone Health](), Henning Hermjakob of [EMBL-EBI](), and Guanming Wu of [OHSU]().
-
-### The Reactome Project
-
-Biological information has become so abundant and complex in recent years that it is difficult, if not impossible, even for expert individuals to manage in traditional publication formats and with existing knowledge management tools. It is an ongoing challenge for researchers to keep up-to-date on research developments in their fields, and identify relevant research to support their own studies without devoting too much time collecting unconnected information. The Reactome group has recognized this challenge and is developing a set of novel online resources that use features of the electronic media to organize biological pathway information in ways that provide for more efficient access and that allow new forms of analysis that were not possible with information stored in the traditional printed literature.
-
-The cornerstone of Reactome is a freely available, open source relational database of signaling and metabolic molecules and their relations organized into biological pathways and processes. The core unit of the Reactome data model is the reaction. Entities (nucleic acids, proteins, complexes, vaccines, anti-cancer therapeutics and small molecules) participating in reactions form a network of biological interactions and are grouped into pathways. Examples of biological pathways in Reactome include [classical intermediary metabolism](), [signaling](), [transcriptional regulation](), [apoptosis]() and [disease](). The Reactome curation process for a pathway is similar to the editing of a scientific review. An external domain expert provides his or her expertise, a curator formalizes it into the database structure, and an external domain expert reviews the representation. A system of evidence tracking ensures that all assertions are backed up by the primary literature.
-
-The Reactome website is designed to literally give the user a graphical map of known biological processes and pathways that is also an interface which the user can ‘click through’ to authoritative detailed information on components and their relations. The Reactome database and website enable scientists, researchers, students, and educators to find, organize, and utilize biological information to support [data visualization](), [integration]() and [analysis](). Reactome pathway, reaction and molecules pages extensively cross-reference to over 100 different online bioinformatics resources, including NCBI Gene, Ensembl and UniProt databases, the UCSC Genome Browser, ChEBI small molecule databases, and the PubMed literature database.
-
-Reactome is used by clinicians, geneticists, genomics researchers, and molecular biologists to interpret the results of high-throughput experimental studies, by bioinformaticians seeking to develop novel algorithms for mining knowledge from genomic studies, and by systems biologists building predictive models of normal and disease variant pathways.
-
-All data and software are freely available for [download](). Interaction, reaction and pathway data are provided as downloadable flat, [Neo4j GraphDB](), [MySQL](), [BioPAX](), [SBML ]()and [PSI-MITAB]() files and are also accessible through our Web Services APIs. Software and instructions for local installation of the Reactome database, website, and data entry tools are also available to support independent pathway curation.
-
-### For more information:
-
-Visit our Youtube Video for an [Introduction to Reactome]()!
-
-If you have any feedback or questions, please contact us at the Reactome This email address is being protected from spambots. You need JavaScript enabled to view it..
-
diff --git a/projects/website-angular/content/about/logo.mdx b/projects/website-angular/content/about/logo.mdx
index e8a4311..3cfb84a 100644
--- a/projects/website-angular/content/about/logo.mdx
+++ b/projects/website-angular/content/about/logo.mdx
@@ -19,29 +19,29 @@ We invite you to use the brand ([imagotype](<#imagotype>)) or the logo ([isotype
PNG
-[Small [297x70]]( "Click to Download our logo in PNG format")
+[Small [297x70]]( "Click to Download our logo in PNG format")
-[Medium [591x138]]( "Click to Download our logo in PNG format")
+[Medium [591x138]]( "Click to Download our logo in PNG format")
-[Large [1183x276]]( "Click to Download our logo in PNG format")
+[Large [1183x276]]( "Click to Download our logo in PNG format")
-[SVG]( "Click to Download our logo in SVG format")
+[SVG]( "Click to Download our logo in SVG format")
-[EMF]( "Click to Download our logo in EMF format")
+[EMF]( "Click to Download our logo in EMF format")

PNG
-[Small [297x70]]( "Click to Download our logo in PNG format")
+[Small [297x70]]( "Click to Download our logo in PNG format")
-[Medium [591x138]]( "Click to Download our logo in PNG format")
+[Medium [591x138]]( "Click to Download our logo in PNG format")
-[Large [1183x276]]( "Click to Download our logo in PNG format")
+[Large [1183x276]]( "Click to Download our logo in PNG format")
-[SVG]( "Click to Download our logo in SVG format")
+[SVG]( "Click to Download our logo in SVG format")
-[EMF]( "Click to Download our logo in EMF format")
+[EMF]( "Click to Download our logo in EMF format")
### [__Isotype](<#isotype>)
@@ -49,26 +49,26 @@ PNG
PNG
-[Small [119x138]]( "Click to Download our logo in PNG format")
+[Small [119x138]]( "Click to Download our logo in PNG format")
-[Medium [297x342]]( "Click to Download our logo in PNG format")
+[Medium [297x342]]( "Click to Download our logo in PNG format")
-[Large [591x682]]( "Click to Download our logo in PNG format")
+[Large [591x682]]( "Click to Download our logo in PNG format")
-[SVG]( "Click to Download our logo in SVG format")
+[SVG]( "Click to Download our logo in SVG format")
-[EMF]( "Click to Download our logo in EMF format")
+[EMF]( "Click to Download our logo in EMF format")

PNG
-[Small [119x138]]( "Click to Download our logo in PNG format")
+[Small [119x138]]( "Click to Download our logo in PNG format")
-[Medium [297x342]]( "Click to Download our logo in PNG format")
+[Medium [297x342]]( "Click to Download our logo in PNG format")
-[Large [591x682]]( "Click to Download our logo in PNG format")
+[Large [591x682]]( "Click to Download our logo in PNG format")
-[SVG]( "Click to Download our logo in SVG format")
+[SVG]( "Click to Download our logo in SVG format")
-[EMF]( "Click to Download our logo in EMF format")
+[EMF]( "Click to Download our logo in EMF format")
diff --git a/projects/website-angular/content/about/news/article-71.mdx b/projects/website-angular/content/about/news/102-version-64-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-71.mdx
rename to projects/website-angular/content/about/news/102-version-64-released.mdx
index f55f42a..46da049 100644
--- a/projects/website-angular/content/about/news/article-71.mdx
+++ b/projects/website-angular/content/about/news/102-version-64-released.mdx
@@ -2,10 +2,10 @@
title: Version 64 Released
category: "about"
date: "2018-03-26T16:00:21-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "102-version-64-released"]
---
-## [ Version 64 Released ]()
+## Version 64 Released
[]()
diff --git a/projects/website-angular/content/about/news/article-70.mdx b/projects/website-angular/content/about/news/103-version-65-released.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-70.mdx
rename to projects/website-angular/content/about/news/103-version-65-released.mdx
index 0dc1078..d64041c 100644
--- a/projects/website-angular/content/about/news/article-70.mdx
+++ b/projects/website-angular/content/about/news/103-version-65-released.mdx
@@ -2,10 +2,10 @@
title: Version 65 Released
category: "about"
date: "2018-06-12T16:19:48-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "103-version-65-released"]
---
-## [ Version 65 Released ]()
+## Version 65 Released
[]()
diff --git a/projects/website-angular/content/about/news/article-68.mdx b/projects/website-angular/content/about/news/105-our-data-now-available-through-google-dataset-search-engine.mdx
similarity index 86%
rename from projects/website-angular/content/about/news/article-68.mdx
rename to projects/website-angular/content/about/news/105-our-data-now-available-through-google-dataset-search-engine.mdx
index dbb8438..d98a4bf 100644
--- a/projects/website-angular/content/about/news/article-68.mdx
+++ b/projects/website-angular/content/about/news/105-our-data-now-available-through-google-dataset-search-engine.mdx
@@ -2,10 +2,10 @@
title: Our data now available through Google Dataset Search
category: "about"
date: "2018-09-06T13:49:49-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "105-our-data-now-available-through-google-dataset-search-engine"]
---
-## [ Our data now available through Google Dataset Search ]()
+## Our data now available through Google Dataset Search

diff --git a/projects/website-angular/content/about/news/article-67.mdx b/projects/website-angular/content/about/news/106-version-66-release.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-67.mdx
rename to projects/website-angular/content/about/news/106-version-66-release.mdx
index a6357c8..3e71b7f 100644
--- a/projects/website-angular/content/about/news/article-67.mdx
+++ b/projects/website-angular/content/about/news/106-version-66-release.mdx
@@ -2,10 +2,10 @@
title: Version 66 Released
category: "about"
date: "2018-09-27T08:56:24-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "106-version-66-release"]
---
-## [ Version 66 Released ]()
+## Version 66 Released
[]()
diff --git a/projects/website-angular/content/about/news/article-69.mdx b/projects/website-angular/content/about/news/107-new-tool-to-generate-analysis-pdf-reports.mdx
similarity index 87%
rename from projects/website-angular/content/about/news/article-69.mdx
rename to projects/website-angular/content/about/news/107-new-tool-to-generate-analysis-pdf-reports.mdx
index 183ed03..1a20346 100644
--- a/projects/website-angular/content/about/news/article-69.mdx
+++ b/projects/website-angular/content/about/news/107-new-tool-to-generate-analysis-pdf-reports.mdx
@@ -2,10 +2,10 @@
title: New tool to generate analysis PDF reports
category: "about"
date: "2018-07-12T06:51:41-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "107-new-tool-to-generate-analysis-pdf-reports"]
---
-## [ New tool to generate analysis PDF reports ]()
+## New tool to generate analysis PDF reports

diff --git a/projects/website-angular/content/about/news/article-66.mdx b/projects/website-angular/content/about/news/109-new-advanced-search-available-in-our-pathway-browser.mdx
similarity index 90%
rename from projects/website-angular/content/about/news/article-66.mdx
rename to projects/website-angular/content/about/news/109-new-advanced-search-available-in-our-pathway-browser.mdx
index 84d1f4a..081f55b 100644
--- a/projects/website-angular/content/about/news/article-66.mdx
+++ b/projects/website-angular/content/about/news/109-new-advanced-search-available-in-our-pathway-browser.mdx
@@ -2,10 +2,10 @@
title: New advanced search available in our PathwayBrowser
category: "about"
date: "2018-10-01T05:05:45-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "109-new-advanced-search-available-in-our-pathway-browser"]
---
-## [ New advanced search available in our PathwayBrowser ]()
+## New advanced search available in our PathwayBrowser
Your browser does not support the video tag.
diff --git a/projects/website-angular/content/about/news/article-65.mdx b/projects/website-angular/content/about/news/110-sbgn-files-revamp.mdx
similarity index 90%
rename from projects/website-angular/content/about/news/article-65.mdx
rename to projects/website-angular/content/about/news/110-sbgn-files-revamp.mdx
index 521b0b7..866dac7 100644
--- a/projects/website-angular/content/about/news/article-65.mdx
+++ b/projects/website-angular/content/about/news/110-sbgn-files-revamp.mdx
@@ -2,10 +2,10 @@
title: SBGN files revamp
category: "about"
date: "2018-10-03T08:53:19-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "110-sbgn-files-revamp"]
---
-## [ SBGN files revamp ]()
+## SBGN files revamp

diff --git a/projects/website-angular/content/about/news/article-64.mdx b/projects/website-angular/content/about/news/111-international-open-access-week-analysis-tools.mdx
similarity index 90%
rename from projects/website-angular/content/about/news/article-64.mdx
rename to projects/website-angular/content/about/news/111-international-open-access-week-analysis-tools.mdx
index a65b9e0..88e969d 100644
--- a/projects/website-angular/content/about/news/article-64.mdx
+++ b/projects/website-angular/content/about/news/111-international-open-access-week-analysis-tools.mdx
@@ -2,10 +2,10 @@
title: "International Open Access Week: Analysis Tools"
category: "about"
date: "2018-10-23T04:25:02-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "111-international-open-access-week-analysis-tools"]
---
-## [ International Open Access Week: Analysis Tools ]()
+## International Open Access Week: Analysis Tools

diff --git a/projects/website-angular/content/about/news/article-63.mdx b/projects/website-angular/content/about/news/112-international-open-access-week-network-analysis-tools.mdx
similarity index 92%
rename from projects/website-angular/content/about/news/article-63.mdx
rename to projects/website-angular/content/about/news/112-international-open-access-week-network-analysis-tools.mdx
index dcdbd8c..3dcf026 100644
--- a/projects/website-angular/content/about/news/article-63.mdx
+++ b/projects/website-angular/content/about/news/112-international-open-access-week-network-analysis-tools.mdx
@@ -2,10 +2,10 @@
title: "International Open Access Week: Network Visualization & Analysis Tool"
category: "about"
date: "2018-10-23T12:54:11-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "112-international-open-access-week-network-analysis-tools"]
---
-## [ International Open Access Week: Network Visualization & Analysis Tool ]()
+## International Open Access Week: Network Visualization & Analysis Tool

diff --git a/projects/website-angular/content/about/news/article-62.mdx b/projects/website-angular/content/about/news/113-oaweek-interactive-illustrations-icons.mdx
similarity index 93%
rename from projects/website-angular/content/about/news/article-62.mdx
rename to projects/website-angular/content/about/news/113-oaweek-interactive-illustrations-icons.mdx
index d7ee546..c29db2a 100644
--- a/projects/website-angular/content/about/news/article-62.mdx
+++ b/projects/website-angular/content/about/news/113-oaweek-interactive-illustrations-icons.mdx
@@ -2,10 +2,10 @@
title: "International Open Access Week: Interactive illustrations and icon library"
category: "about"
date: "2018-10-25T04:19:18-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "113-oaweek-interactive-illustrations-icons"]
---
-## [ International Open Access Week: Interactive illustrations and icon library ]()
+## International Open Access Week: Interactive illustrations and icon library

diff --git a/projects/website-angular/content/about/news/article-61.mdx b/projects/website-angular/content/about/news/114-graphdb-content-service.mdx
similarity index 93%
rename from projects/website-angular/content/about/news/article-61.mdx
rename to projects/website-angular/content/about/news/114-graphdb-content-service.mdx
index ddacd6f..86a417b 100644
--- a/projects/website-angular/content/about/news/article-61.mdx
+++ b/projects/website-angular/content/about/news/114-graphdb-content-service.mdx
@@ -2,10 +2,10 @@
title: "International Open Access Week: GraphDB and Content Service"
category: "about"
date: "2018-10-26T04:21:29-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "114-graphdb-content-service"]
---
-## [ International Open Access Week: GraphDB and Content Service ]()
+## International Open Access Week: GraphDB and Content Service

diff --git a/projects/website-angular/content/about/news/article-60.mdx b/projects/website-angular/content/about/news/115-performing-gsea-analysis-with-reactomefiviz-app.mdx
similarity index 86%
rename from projects/website-angular/content/about/news/article-60.mdx
rename to projects/website-angular/content/about/news/115-performing-gsea-analysis-with-reactomefiviz-app.mdx
index 5db10dc..f0d0283 100644
--- a/projects/website-angular/content/about/news/article-60.mdx
+++ b/projects/website-angular/content/about/news/115-performing-gsea-analysis-with-reactomefiviz-app.mdx
@@ -2,10 +2,10 @@
title: Performing GSEA analysis with ReactomeFIViz app
category: "about"
date: "2018-11-01T10:00:04-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "115-performing-gsea-analysis-with-reactomefiviz-app"]
---
-## [ Performing GSEA analysis with ReactomeFIViz app ]()
+## Performing GSEA analysis with ReactomeFIViz app

diff --git a/projects/website-angular/content/about/news/article-59.mdx b/projects/website-angular/content/about/news/116-version-67-released.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-59.mdx
rename to projects/website-angular/content/about/news/116-version-67-released.mdx
index b0ab90d..2afbe39 100644
--- a/projects/website-angular/content/about/news/article-59.mdx
+++ b/projects/website-angular/content/about/news/116-version-67-released.mdx
@@ -2,10 +2,10 @@
title: Version 67 Released
category: "about"
date: "2018-12-10T15:18:07-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "116-version-67-released"]
---
-## [ Version 67 Released ]()
+## Version 67 Released

diff --git a/projects/website-angular/content/about/news/article-58.mdx b/projects/website-angular/content/about/news/119-new-reactions-layout-with-nested-compartments.mdx
similarity index 94%
rename from projects/website-angular/content/about/news/article-58.mdx
rename to projects/website-angular/content/about/news/119-new-reactions-layout-with-nested-compartments.mdx
index a3cdd89..b5efc04 100644
--- a/projects/website-angular/content/about/news/article-58.mdx
+++ b/projects/website-angular/content/about/news/119-new-reactions-layout-with-nested-compartments.mdx
@@ -2,10 +2,10 @@
title: New reactions layout with nested compartments
category: "about"
date: "2019-01-09T08:52:43-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "119-new-reactions-layout-with-nested-compartments"]
---
-## [ New reactions layout with nested compartments ]()
+## New reactions layout with nested compartments
[]()
diff --git a/projects/website-angular/content/about/news/article-57.mdx b/projects/website-angular/content/about/news/120-classifying-pathways-into-tissue-types-based-on-protein-expression.mdx
similarity index 87%
rename from projects/website-angular/content/about/news/article-57.mdx
rename to projects/website-angular/content/about/news/120-classifying-pathways-into-tissue-types-based-on-protein-expression.mdx
index 6510498..252968c 100644
--- a/projects/website-angular/content/about/news/article-57.mdx
+++ b/projects/website-angular/content/about/news/120-classifying-pathways-into-tissue-types-based-on-protein-expression.mdx
@@ -2,10 +2,10 @@
title: New tool to classify pathways into tissue types based on protein expression
category: "about"
date: "2019-01-15T05:56:23-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "120-classifying-pathways-into-tissue-types-based-on-protein-expression"]
---
-## [ New tool to classify pathways into tissue types based on protein expression ]()
+## New tool to classify pathways into tissue types based on protein expression

diff --git a/projects/website-angular/content/about/news/article-56.mdx b/projects/website-angular/content/about/news/121-new-pdf-book.mdx
similarity index 93%
rename from projects/website-angular/content/about/news/article-56.mdx
rename to projects/website-angular/content/about/news/121-new-pdf-book.mdx
index b4b1d84..2cbda99 100644
--- a/projects/website-angular/content/about/news/article-56.mdx
+++ b/projects/website-angular/content/about/news/121-new-pdf-book.mdx
@@ -2,10 +2,10 @@
title: Download our biological pathway knowledge in a PDF book
category: "about"
date: "2019-01-30T10:26:26-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "121-new-pdf-book"]
---
-## [ Download our biological pathway knowledge in a PDF book ]()
+## Download our biological pathway knowledge in a PDF book

diff --git a/projects/website-angular/content/about/news/article-55.mdx b/projects/website-angular/content/about/news/124-deprecated-support-for-reactome-restful-api.mdx
similarity index 84%
rename from projects/website-angular/content/about/news/article-55.mdx
rename to projects/website-angular/content/about/news/124-deprecated-support-for-reactome-restful-api.mdx
index cf2f101..44d23a4 100644
--- a/projects/website-angular/content/about/news/article-55.mdx
+++ b/projects/website-angular/content/about/news/124-deprecated-support-for-reactome-restful-api.mdx
@@ -2,10 +2,10 @@
title: Deprecated support for Reactome RESTful API
category: "about"
date: "2019-03-20T13:14:21-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "124-deprecated-support-for-reactome-restful-api"]
---
-## [ Deprecated support for Reactome RESTful API ]()
+## Deprecated support for Reactome RESTful API
As of Version 68 (March 2019), the RESTful API is deprecated and will be superceded by our ContentService. The ContentService is currently available through our website at and is designed for bioinformaticians, computer scientists, and software developers to access our pathway data. See more details about this API at [https://reactome.org/dev/content-service. ]()
diff --git a/projects/website-angular/content/about/news/article-54.mdx b/projects/website-angular/content/about/news/125-version-68-of-reactome-is-now-online.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-54.mdx
rename to projects/website-angular/content/about/news/125-version-68-of-reactome-is-now-online.mdx
index 0993a99..ba1bb05 100644
--- a/projects/website-angular/content/about/news/article-54.mdx
+++ b/projects/website-angular/content/about/news/125-version-68-of-reactome-is-now-online.mdx
@@ -2,10 +2,10 @@
title: Version 68 Released
category: "about"
date: "2019-03-20T13:31:18-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "125-version-68-of-reactome-is-now-online"]
---
-## [ Version 68 Released ]()
+## Version 68 Released

diff --git a/projects/website-angular/content/about/news/article-53.mdx b/projects/website-angular/content/about/news/126-panther-resource-now-used-to-identify-orthologs-of-curated-human-proteins.mdx
similarity index 92%
rename from projects/website-angular/content/about/news/article-53.mdx
rename to projects/website-angular/content/about/news/126-panther-resource-now-used-to-identify-orthologs-of-curated-human-proteins.mdx
index cf2ec4e..30e1075 100644
--- a/projects/website-angular/content/about/news/article-53.mdx
+++ b/projects/website-angular/content/about/news/126-panther-resource-now-used-to-identify-orthologs-of-curated-human-proteins.mdx
@@ -2,10 +2,10 @@
title: PANTHER resource now used to identify orthologs of curated human proteins
category: "about"
date: "2019-03-20T13:39:44-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "126-panther-resource-now-used-to-identify-orthologs-of-curated-human-proteins"]
---
-## [ PANTHER resource now used to identify orthologs of curated human proteins ]()
+## PANTHER resource now used to identify orthologs of curated human proteins

diff --git a/projects/website-angular/content/about/news/article-52.mdx b/projects/website-angular/content/about/news/127-come-to-our-release-68-webinar-to-learn-more-about-reactome.mdx
similarity index 89%
rename from projects/website-angular/content/about/news/article-52.mdx
rename to projects/website-angular/content/about/news/127-come-to-our-release-68-webinar-to-learn-more-about-reactome.mdx
index 1abbd08..2e2f7a5 100644
--- a/projects/website-angular/content/about/news/article-52.mdx
+++ b/projects/website-angular/content/about/news/127-come-to-our-release-68-webinar-to-learn-more-about-reactome.mdx
@@ -2,10 +2,10 @@
title: Come to our release 68 webinar to learn more about Reactome
category: "about"
date: "2019-03-28T14:08:26-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "127-come-to-our-release-68-webinar-to-learn-more-about-reactome"]
---
-## [ Come to our release 68 webinar to learn more about Reactome ]()
+## Come to our release 68 webinar to learn more about Reactome

diff --git a/projects/website-angular/content/about/news/article-51.mdx b/projects/website-angular/content/about/news/133-reactomefiviz-app-version-7-2-0-released.mdx
similarity index 92%
rename from projects/website-angular/content/about/news/article-51.mdx
rename to projects/website-angular/content/about/news/133-reactomefiviz-app-version-7-2-0-released.mdx
index d014654..6913a9e 100644
--- a/projects/website-angular/content/about/news/article-51.mdx
+++ b/projects/website-angular/content/about/news/133-reactomefiviz-app-version-7-2-0-released.mdx
@@ -2,10 +2,10 @@
title: ReactomeFI Network v2018 and FIViz App v2.7.0 Released
category: "about"
date: "2019-04-18T00:09:37-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "133-reactomefiviz-app-version-7-2-0-released"]
---
-## [ ReactomeFI Network v2018 and FIViz App v2.7.0 Released ]()
+## ReactomeFI Network v2018 and FIViz App v2.7.0 Released

diff --git a/projects/website-angular/content/about/news/article-50.mdx b/projects/website-angular/content/about/news/136-season-of-docs.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-50.mdx
rename to projects/website-angular/content/about/news/136-season-of-docs.mdx
index 91a552f..73d1660 100644
--- a/projects/website-angular/content/about/news/article-50.mdx
+++ b/projects/website-angular/content/about/news/136-season-of-docs.mdx
@@ -2,10 +2,10 @@
title: Season of Docs
category: "about"
date: "2019-04-22T15:13:04-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "136-season-of-docs"]
---
-## [ Season of Docs ]()
+## Season of Docs
* 
diff --git a/projects/website-angular/content/about/news/article-49.mdx b/projects/website-angular/content/about/news/137-version-69-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-49.mdx
rename to projects/website-angular/content/about/news/137-version-69-released.mdx
index bfd54d4..d076d97 100644
--- a/projects/website-angular/content/about/news/article-49.mdx
+++ b/projects/website-angular/content/about/news/137-version-69-released.mdx
@@ -2,10 +2,10 @@
title: Version 69 Released
category: "about"
date: "2019-05-28T13:41:55-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "137-version-69-released"]
---
-## [ Version 69 Released ]()
+## Version 69 Released

diff --git a/projects/website-angular/content/about/news/article-47.mdx b/projects/website-angular/content/about/news/138-orcid-claim-your-works.mdx
similarity index 93%
rename from projects/website-angular/content/about/news/article-47.mdx
rename to projects/website-angular/content/about/news/138-orcid-claim-your-works.mdx
index e2e2377..a071e16 100644
--- a/projects/website-angular/content/about/news/article-47.mdx
+++ b/projects/website-angular/content/about/news/138-orcid-claim-your-works.mdx
@@ -2,10 +2,10 @@
title: "ORCID: Claim your works"
category: "about"
date: "2019-06-06T10:16:00-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "138-orcid-claim-your-works"]
---
-## [ ORCID: Claim your works ]()
+## ORCID: Claim your works

diff --git a/projects/website-angular/content/about/news/141-reacfoam-genome-wide-pathway-overview-based-on-voronoi-tessellation.mdx b/projects/website-angular/content/about/news/141-reacfoam-genome-wide-pathway-overview-based-on-voronoi-tessellation.mdx
new file mode 100644
index 0000000..613e272
--- /dev/null
+++ b/projects/website-angular/content/about/news/141-reacfoam-genome-wide-pathway-overview-based-on-voronoi-tessellation.mdx
@@ -0,0 +1,14 @@
+---
+title: "ReacFoam: Genome-wide pathway overview based on Voronoi tessellation"
+category: "about"
+date: "2019-07-02T07:38:13-04:00"
+tags: ["about", "news", "141-reacfoam-genome-wide-pathway-overview-based-on-voronoi-tessellation"]
+---
+
+## ReacFoam: Genome-wide pathway overview based on Voronoi tessellation
+
+[]()
+
+As part of our continuous effort to provide visually attractive and more user-friendly access to our biological pathways, we are pleased to announce our new high level pathways overview visualisation based on Voronoi tessellation.
+
+Following any [pathway analysis](), the [pathway overview]() provides a [new icon]() which leads to a comprehensive, highly visual, interactive overview of pathway analysis results. The functionality is available on both desktop and mobile platforms.
diff --git a/projects/website-angular/content/about/news/article-45.mdx b/projects/website-angular/content/about/news/142-version-70-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-45.mdx
rename to projects/website-angular/content/about/news/142-version-70-released.mdx
index d05a0d3..6b0ba79 100644
--- a/projects/website-angular/content/about/news/article-45.mdx
+++ b/projects/website-angular/content/about/news/142-version-70-released.mdx
@@ -2,10 +2,10 @@
title: Version 70 Released
category: "about"
date: "2019-08-30T12:30:07-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "142-version-70-released"]
---
-## [ Version 70 Released ]()
+## Version 70 Released

diff --git a/projects/website-angular/content/about/news/article-44.mdx b/projects/website-angular/content/about/news/144-new-reactome-publication-published-in-2020-nar-database-issue.mdx
similarity index 80%
rename from projects/website-angular/content/about/news/article-44.mdx
rename to projects/website-angular/content/about/news/144-new-reactome-publication-published-in-2020-nar-database-issue.mdx
index f0ff929..57dc789 100644
--- a/projects/website-angular/content/about/news/article-44.mdx
+++ b/projects/website-angular/content/about/news/144-new-reactome-publication-published-in-2020-nar-database-issue.mdx
@@ -2,10 +2,10 @@
title: New Reactome Publication published in 2020 NAR Database Issue
category: "about"
date: "2019-12-02T00:38:57-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "144-new-reactome-publication-published-in-2020-nar-database-issue"]
---
-## [ New Reactome Publication published in 2020 NAR Database Issue ]()
+## New Reactome Publication published in 2020 NAR Database Issue

diff --git a/projects/website-angular/content/about/news/article-43.mdx b/projects/website-angular/content/about/news/145-version-71-releases.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-43.mdx
rename to projects/website-angular/content/about/news/145-version-71-releases.mdx
index ed8ac86..38400f8 100644
--- a/projects/website-angular/content/about/news/article-43.mdx
+++ b/projects/website-angular/content/about/news/145-version-71-releases.mdx
@@ -2,10 +2,10 @@
title: Version 71 Released
category: "about"
date: "2019-12-02T00:43:55-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "145-version-71-releases"]
---
-## [ Version 71 Released ]()
+## Version 71 Released
****
diff --git a/projects/website-angular/content/about/news/article-42.mdx b/projects/website-angular/content/about/news/146-new-paper-published-in-database-oxford.mdx
similarity index 83%
rename from projects/website-angular/content/about/news/article-42.mdx
rename to projects/website-angular/content/about/news/146-new-paper-published-in-database-oxford.mdx
index b3db1a5..b1feaea 100644
--- a/projects/website-angular/content/about/news/article-42.mdx
+++ b/projects/website-angular/content/about/news/146-new-paper-published-in-database-oxford.mdx
@@ -2,10 +2,10 @@
title: New Paper published in Database (Oxford)
category: "about"
date: "2019-12-18T23:36:50-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "146-new-paper-published-in-database-oxford"]
---
-## [ New Paper published in Database (Oxford) ]()
+## New Paper published in Database (Oxford)

diff --git a/projects/website-angular/content/about/news/article-41.mdx b/projects/website-angular/content/about/news/147-version-72-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-41.mdx
rename to projects/website-angular/content/about/news/147-version-72-released.mdx
index 7991cbd..15511bc 100644
--- a/projects/website-angular/content/about/news/article-41.mdx
+++ b/projects/website-angular/content/about/news/147-version-72-released.mdx
@@ -2,10 +2,10 @@
title: Version 72 Released
category: "about"
date: "2020-03-12T21:38:36-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "147-version-72-released"]
---
-## [ Version 72 Released ]()
+## Version 72 Released

diff --git a/projects/website-angular/content/about/news/article-40.mdx b/projects/website-angular/content/about/news/150-new-paper-published-in-elife.mdx
similarity index 86%
rename from projects/website-angular/content/about/news/article-40.mdx
rename to projects/website-angular/content/about/news/150-new-paper-published-in-elife.mdx
index 116f909..d95a3c5 100644
--- a/projects/website-angular/content/about/news/article-40.mdx
+++ b/projects/website-angular/content/about/news/150-new-paper-published-in-elife.mdx
@@ -2,10 +2,10 @@
title: New Paper published in eLife
category: "about"
date: "2020-04-06T12:48:52-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "150-new-paper-published-in-elife"]
---
-## [ New Paper published in eLife ]()
+## New Paper published in eLife

diff --git a/projects/website-angular/content/about/news/article-39.mdx b/projects/website-angular/content/about/news/151-new-package-facilitates-communication-between-reactome-services-and-data-with-python.mdx
similarity index 89%
rename from projects/website-angular/content/about/news/article-39.mdx
rename to projects/website-angular/content/about/news/151-new-package-facilitates-communication-between-reactome-services-and-data-with-python.mdx
index 3af908c..538fa4a 100644
--- a/projects/website-angular/content/about/news/article-39.mdx
+++ b/projects/website-angular/content/about/news/151-new-package-facilitates-communication-between-reactome-services-and-data-with-python.mdx
@@ -2,10 +2,10 @@
title: New reactome2py package links our web services and data with python.
category: "about"
date: "2020-04-06T13:59:35-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "151-new-package-facilitates-communication-between-reactome-services-and-data-with-python"]
---
-## [ New reactome2py package links our web services and data with python. ]()
+## New reactome2py package links our web services and data with python.

diff --git a/projects/website-angular/content/about/news/article-38.mdx b/projects/website-angular/content/about/news/154-new-paper-published-in-autophagy.mdx
similarity index 85%
rename from projects/website-angular/content/about/news/article-38.mdx
rename to projects/website-angular/content/about/news/154-new-paper-published-in-autophagy.mdx
index 331885b..60d4090 100644
--- a/projects/website-angular/content/about/news/article-38.mdx
+++ b/projects/website-angular/content/about/news/154-new-paper-published-in-autophagy.mdx
@@ -2,9 +2,9 @@
title: New Paper published in Autophagy
category: "about"
date: "2020-06-11T15:40:50-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "154-new-paper-published-in-autophagy"]
---
-## [ New Paper published in Autophagy ]()
+## New Paper published in Autophagy
A new research article entitled “[Using Reactome to build an autophagy mechanism knowledgebase]()” has been published in the [Autophagy]() journal. The paper discusses the curation and annotation of the molecular mechanisms of autophagy. The autophagy pathway can be viewed [here](). More publications from the [Reactome Team]() can be found [here]().
diff --git a/projects/website-angular/content/about/news/article-37.mdx b/projects/website-angular/content/about/news/155-version-73-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-37.mdx
rename to projects/website-angular/content/about/news/155-version-73-released.mdx
index a970524..270773a 100644
--- a/projects/website-angular/content/about/news/article-37.mdx
+++ b/projects/website-angular/content/about/news/155-version-73-released.mdx
@@ -2,10 +2,10 @@
title: Version 73 Released
category: "about"
date: "2020-06-11T16:56:40-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "155-version-73-released"]
---
-## [ Version 73 Released ]()
+## Version 73 Released

diff --git a/projects/website-angular/content/about/news/article-36.mdx b/projects/website-angular/content/about/news/161-version-74-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-36.mdx
rename to projects/website-angular/content/about/news/161-version-74-released.mdx
index 5f0136f..3d7899c 100644
--- a/projects/website-angular/content/about/news/article-36.mdx
+++ b/projects/website-angular/content/about/news/161-version-74-released.mdx
@@ -2,10 +2,10 @@
title: "COVID-19: SARS-CoV-2 infection pathway Released"
category: "about"
date: "2020-09-21T16:18:18-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "161-version-74-released"]
---
-## [ COVID-19: SARS-CoV-2 infection pathway Released ]()
+## COVID-19: SARS-CoV-2 infection pathway Released

diff --git a/projects/website-angular/content/about/news/article-35.mdx b/projects/website-angular/content/about/news/162-new-paper-published-in-molecular-cellular-proteomics.mdx
similarity index 84%
rename from projects/website-angular/content/about/news/article-35.mdx
rename to projects/website-angular/content/about/news/162-new-paper-published-in-molecular-cellular-proteomics.mdx
index 535c832..534a5fc 100644
--- a/projects/website-angular/content/about/news/article-35.mdx
+++ b/projects/website-angular/content/about/news/162-new-paper-published-in-molecular-cellular-proteomics.mdx
@@ -2,10 +2,10 @@
title: "New Paper published in Molecular & Cellular Proteomics"
category: "about"
date: "2020-10-20T11:16:35-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "162-new-paper-published-in-molecular-cellular-proteomics"]
---
-## [ New Paper published in Molecular & Cellular Proteomics ]()
+## New Paper published in Molecular & Cellular Proteomics

diff --git a/projects/website-angular/content/about/news/article-34.mdx b/projects/website-angular/content/about/news/163-version-75-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-34.mdx
rename to projects/website-angular/content/about/news/163-version-75-released.mdx
index cbf016c..22e4037 100644
--- a/projects/website-angular/content/about/news/article-34.mdx
+++ b/projects/website-angular/content/about/news/163-version-75-released.mdx
@@ -2,10 +2,10 @@
title: Version 75 Released
category: "about"
date: "2020-12-02T13:58:30-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "163-version-75-released"]
---
-## [ Version 75 Released ]()
+## Version 75 Released

diff --git a/projects/website-angular/content/about/news/article-33.mdx b/projects/website-angular/content/about/news/164-version-76-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-33.mdx
rename to projects/website-angular/content/about/news/164-version-76-released.mdx
index 4a5bddf..3b6d7d9 100644
--- a/projects/website-angular/content/about/news/article-33.mdx
+++ b/projects/website-angular/content/about/news/164-version-76-released.mdx
@@ -2,10 +2,10 @@
title: Version 76 Released
category: "about"
date: "2021-03-21T23:18:41-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "164-version-76-released"]
---
-## [ Version 76 Released ]()
+## Version 76 Released

diff --git a/projects/website-angular/content/about/news/article-32.mdx b/projects/website-angular/content/about/news/165-the-reactome-idg-portal-is-released.mdx
similarity index 95%
rename from projects/website-angular/content/about/news/article-32.mdx
rename to projects/website-angular/content/about/news/165-the-reactome-idg-portal-is-released.mdx
index 3b771cb..d733b59 100644
--- a/projects/website-angular/content/about/news/article-32.mdx
+++ b/projects/website-angular/content/about/news/165-the-reactome-idg-portal-is-released.mdx
@@ -2,10 +2,10 @@
title: The Reactome IDG Portal is released
category: "about"
date: "2021-06-07T13:47:37-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "165-the-reactome-idg-portal-is-released"]
---
-## [ The Reactome IDG Portal is released ]()
+## The Reactome IDG Portal is released

diff --git a/projects/website-angular/content/about/news/article-31.mdx b/projects/website-angular/content/about/news/166-version-77-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-31.mdx
rename to projects/website-angular/content/about/news/166-version-77-released.mdx
index b997eb5..5facf04 100644
--- a/projects/website-angular/content/about/news/article-31.mdx
+++ b/projects/website-angular/content/about/news/166-version-77-released.mdx
@@ -2,10 +2,10 @@
title: Version 77 Released
category: "about"
date: "2021-06-09T15:53:12-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "166-version-77-released"]
---
-## [ Version 77 Released ]()
+## Version 77 Released

diff --git a/projects/website-angular/content/about/news/article-30.mdx b/projects/website-angular/content/about/news/167-reactome-multi-omics-pathway-analysis-webinar-reaches-record-attendance.mdx
similarity index 85%
rename from projects/website-angular/content/about/news/article-30.mdx
rename to projects/website-angular/content/about/news/167-reactome-multi-omics-pathway-analysis-webinar-reaches-record-attendance.mdx
index fc6196e..7ff7c49 100644
--- a/projects/website-angular/content/about/news/article-30.mdx
+++ b/projects/website-angular/content/about/news/167-reactome-multi-omics-pathway-analysis-webinar-reaches-record-attendance.mdx
@@ -2,9 +2,9 @@
title: "Reactome Multi-Omics Pathway Analysis webinar reaches record attendance"
category: "about"
date: "2021-06-21T12:32:18-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "167-reactome-multi-omics-pathway-analysis-webinar-reaches-record-attendance"]
---
-## [ Reactome Multi-Omics Pathway Analysis webinar reaches record attendance ]()
+## Reactome Multi-Omics Pathway Analysis webinar reaches record attendance
On June 2, 2021, Dr. Johannes Griss presented "[A guide to multiomics pathway analysis]()" to a record audience of 572 participants as part of the [EMBL-EBI webinar series](). The [seminar recording]() and [training materials](), which are now available online, provides insight into performing comparative multi-omics pathway analyses using the ReactomeGSA platform. We want to take this opportunity to thank all those that took part in the training webinar. If you are interested in learning more about Reactome or if you have any additional questions, please This email address is being protected from spambots. You need JavaScript enabled to view it..
diff --git a/projects/website-angular/content/about/news/article-29.mdx b/projects/website-angular/content/about/news/169-version-78-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-29.mdx
rename to projects/website-angular/content/about/news/169-version-78-released.mdx
index b1b756c..b09f556 100644
--- a/projects/website-angular/content/about/news/article-29.mdx
+++ b/projects/website-angular/content/about/news/169-version-78-released.mdx
@@ -2,10 +2,10 @@
title: Version 78 Released
category: "about"
date: "2021-10-10T23:10:15-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "169-version-78-released"]
---
-## [ Version 78 Released ]()
+## Version 78 Released

diff --git a/projects/website-angular/content/about/news/article-28.mdx b/projects/website-angular/content/about/news/172-we-want-to-hear-your-success-story.mdx
similarity index 94%
rename from projects/website-angular/content/about/news/article-28.mdx
rename to projects/website-angular/content/about/news/172-we-want-to-hear-your-success-story.mdx
index 87b5d74..bb27ddc 100644
--- a/projects/website-angular/content/about/news/article-28.mdx
+++ b/projects/website-angular/content/about/news/172-we-want-to-hear-your-success-story.mdx
@@ -2,10 +2,10 @@
title: "We want to hear your Success Story!"
category: "about"
date: "2021-11-01T13:25:15-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "172-we-want-to-hear-your-success-story"]
---
-## [ We want to hear your Success Story! ]()
+## We want to hear your Success Story!
Our user community is built on success stories! There are thousands of you around the world and many of you have created amazing experiments, tools and resources using Reactome. We're always looking for ways to share interesting discoveries made with Reactome, and have decided to showcase projects on our website to inspire others to create new stories. If you have a novel success story or use case you'd like to share, we would love to hear from you.
diff --git a/projects/website-angular/content/about/news/article-27.mdx b/projects/website-angular/content/about/news/174-version-79-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-27.mdx
rename to projects/website-angular/content/about/news/174-version-79-released.mdx
index f48dbc9..c556653 100644
--- a/projects/website-angular/content/about/news/article-27.mdx
+++ b/projects/website-angular/content/about/news/174-version-79-released.mdx
@@ -2,10 +2,10 @@
title: Version 79 Released
category: "about"
date: "2021-12-13T01:01:33-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "174-version-79-released"]
---
-## [ Version 79 Released ]()
+## Version 79 Released

diff --git a/projects/website-angular/content/about/news/article-26.mdx b/projects/website-angular/content/about/news/175-version-80-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-26.mdx
rename to projects/website-angular/content/about/news/175-version-80-released.mdx
index 254ee71..9589506 100644
--- a/projects/website-angular/content/about/news/article-26.mdx
+++ b/projects/website-angular/content/about/news/175-version-80-released.mdx
@@ -2,10 +2,10 @@
title: Version 80 Released
category: "about"
date: "2022-04-05T13:59:24-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "175-version-80-released"]
---
-## [ Version 80 Released ]()
+## Version 80 Released

diff --git a/projects/website-angular/content/about/news/article-23.mdx b/projects/website-angular/content/about/news/176-version-82-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-23.mdx
rename to projects/website-angular/content/about/news/176-version-82-released.mdx
index b3a2857..8ba4016 100644
--- a/projects/website-angular/content/about/news/article-23.mdx
+++ b/projects/website-angular/content/about/news/176-version-82-released.mdx
@@ -2,10 +2,10 @@
title: Version 82 Released
category: "about"
date: "2022-09-15T14:28:48-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "176-version-82-released"]
---
-## [ Version 82 Released ]()
+## Version 82 Released
[SARS-CoV-2 Infection]()
diff --git a/projects/website-angular/content/about/news/article-25.mdx b/projects/website-angular/content/about/news/177-new-paper-pulished-in-database-oxford.mdx
similarity index 86%
rename from projects/website-angular/content/about/news/article-25.mdx
rename to projects/website-angular/content/about/news/177-new-paper-pulished-in-database-oxford.mdx
index 8ff685d..d0c97dc 100644
--- a/projects/website-angular/content/about/news/article-25.mdx
+++ b/projects/website-angular/content/about/news/177-new-paper-pulished-in-database-oxford.mdx
@@ -2,10 +2,10 @@
title: New Paper pulished in Database (Oxford)
category: "about"
date: "2022-06-16T13:28:48-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "177-new-paper-pulished-in-database-oxford"]
---
-## [ New Paper pulished in Database (Oxford) ]()
+## New Paper pulished in Database (Oxford)

diff --git a/projects/website-angular/content/about/news/article-24.mdx b/projects/website-angular/content/about/news/178-version-81-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-24.mdx
rename to projects/website-angular/content/about/news/178-version-81-released.mdx
index aed8630..c76c48a 100644
--- a/projects/website-angular/content/about/news/article-24.mdx
+++ b/projects/website-angular/content/about/news/178-version-81-released.mdx
@@ -2,10 +2,10 @@
title: Version 81 Released
category: "about"
date: "2022-06-14T13:25:02-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "178-version-81-released"]
---
-## [ Version 81 Released ]()
+## Version 81 Released

diff --git a/projects/website-angular/content/about/news/article-22.mdx b/projects/website-angular/content/about/news/179-collaboration-with-pharmgkb.mdx
similarity index 90%
rename from projects/website-angular/content/about/news/article-22.mdx
rename to projects/website-angular/content/about/news/179-collaboration-with-pharmgkb.mdx
index ceaccd0..20b6abd 100644
--- a/projects/website-angular/content/about/news/article-22.mdx
+++ b/projects/website-angular/content/about/news/179-collaboration-with-pharmgkb.mdx
@@ -2,10 +2,10 @@
title: Collaboration with PharmGKB
category: "about"
date: "2022-09-28T11:02:07-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "179-collaboration-with-pharmgkb"]
---
-## [ Collaboration with PharmGKB ]()
+## Collaboration with PharmGKB

diff --git a/projects/website-angular/content/about/news/article-21.mdx b/projects/website-angular/content/about/news/183-reactome-research-spotlight.mdx
similarity index 92%
rename from projects/website-angular/content/about/news/article-21.mdx
rename to projects/website-angular/content/about/news/183-reactome-research-spotlight.mdx
index eb4946f..34d4838 100644
--- a/projects/website-angular/content/about/news/article-21.mdx
+++ b/projects/website-angular/content/about/news/183-reactome-research-spotlight.mdx
@@ -2,10 +2,10 @@
title: Reactome Research Spotlight
category: "about"
date: "2022-11-10T12:22:32-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "183-reactome-research-spotlight"]
---
-## [ Reactome Research Spotlight ]()
+## Reactome Research Spotlight
Introducing the Reactome Research Spotlight!
diff --git a/projects/website-angular/content/about/news/article-20.mdx b/projects/website-angular/content/about/news/184-version-83-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-20.mdx
rename to projects/website-angular/content/about/news/184-version-83-released.mdx
index 82d9a9d..c20ce6e 100644
--- a/projects/website-angular/content/about/news/article-20.mdx
+++ b/projects/website-angular/content/about/news/184-version-83-released.mdx
@@ -2,10 +2,10 @@
title: Version 83 Released
category: "about"
date: "2022-11-30T11:53:24-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "184-version-83-released"]
---
-## [ Version 83 Released ]()
+## Version 83 Released
[Cytokine Signaling in Immune System]()
diff --git a/projects/website-angular/content/about/news/article-19.mdx b/projects/website-angular/content/about/news/187-reactome-named-as-a-global-core-biodata-resource.mdx
similarity index 84%
rename from projects/website-angular/content/about/news/article-19.mdx
rename to projects/website-angular/content/about/news/187-reactome-named-as-a-global-core-biodata-resource.mdx
index 45506c3..aa37019 100644
--- a/projects/website-angular/content/about/news/article-19.mdx
+++ b/projects/website-angular/content/about/news/187-reactome-named-as-a-global-core-biodata-resource.mdx
@@ -2,10 +2,10 @@
title: Reactome named as a Global Core Biodata Resource
category: "about"
date: "2022-12-21T13:16:35-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "187-reactome-named-as-a-global-core-biodata-resource"]
---
-## [ Reactome named as a Global Core Biodata Resource ]()
+## Reactome named as a Global Core Biodata Resource
On December 15, the Global Biodata Coalition ([]()) of research funders recognized the Reactome Knowledgebase ([www.reactome.org]()) as one of just 37 resources worldwide whose long-term funding and sustainability are critical to life science and biomedical research. See the full announcement [here]().
diff --git a/projects/website-angular/content/about/news/article-18.mdx b/projects/website-angular/content/about/news/191-reactome-is-hiring.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-18.mdx
rename to projects/website-angular/content/about/news/191-reactome-is-hiring.mdx
index d1aa420..1860f6b 100644
--- a/projects/website-angular/content/about/news/article-18.mdx
+++ b/projects/website-angular/content/about/news/191-reactome-is-hiring.mdx
@@ -2,10 +2,10 @@
title: "Reactome is hiring!"
category: "about"
date: "2023-03-20T13:26:04-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "191-reactome-is-hiring"]
---
-## [ Reactome is hiring! ]()
+## Reactome is hiring!
**Outreach Coordinator**
diff --git a/projects/website-angular/content/about/news/article-17.mdx b/projects/website-angular/content/about/news/194-v84-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-17.mdx
rename to projects/website-angular/content/about/news/194-v84-released.mdx
index f7b1814..44c43cb 100644
--- a/projects/website-angular/content/about/news/article-17.mdx
+++ b/projects/website-angular/content/about/news/194-v84-released.mdx
@@ -2,10 +2,10 @@
title: V84 released
category: "about"
date: "2023-03-23T15:07:16-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "194-v84-released"]
---
-## [ V84 released ]()
+## V84 released
[Infectious Disease]()
diff --git a/projects/website-angular/content/about/news/article-16.mdx b/projects/website-angular/content/about/news/225-v85-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-16.mdx
rename to projects/website-angular/content/about/news/225-v85-released.mdx
index 2e00a15..224ed77 100644
--- a/projects/website-angular/content/about/news/article-16.mdx
+++ b/projects/website-angular/content/about/news/225-v85-released.mdx
@@ -2,10 +2,10 @@
title: V85 released
category: "about"
date: "2023-05-25T20:43:26-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "225-v85-released"]
---
-## [ V85 released ]()
+## V85 released

diff --git a/projects/website-angular/content/about/news/article-15.mdx b/projects/website-angular/content/about/news/230-version-86-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-15.mdx
rename to projects/website-angular/content/about/news/230-version-86-released.mdx
index 9dee4ff..1df0dd8 100644
--- a/projects/website-angular/content/about/news/article-15.mdx
+++ b/projects/website-angular/content/about/news/230-version-86-released.mdx
@@ -2,10 +2,10 @@
title: V86 Released
category: "about"
date: "2023-09-07T00:15:08-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "230-version-86-released"]
---
-## [ V86 Released ]()
+## V86 Released

diff --git a/projects/website-angular/content/about/news/article-14.mdx b/projects/website-angular/content/about/news/238-version-87-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-14.mdx
rename to projects/website-angular/content/about/news/238-version-87-released.mdx
index d775428..1ece0b4 100644
--- a/projects/website-angular/content/about/news/article-14.mdx
+++ b/projects/website-angular/content/about/news/238-version-87-released.mdx
@@ -2,10 +2,10 @@
title: V87 released
category: "about"
date: "2023-12-04T22:25:30-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "238-version-87-released"]
---
-## [ V87 released ]()
+## V87 released
**Respiratory Syncytial Virus Infection Pathway**
diff --git a/projects/website-angular/content/about/news/article-13.mdx b/projects/website-angular/content/about/news/246-v88-released-2.mdx
similarity index 99%
rename from projects/website-angular/content/about/news/article-13.mdx
rename to projects/website-angular/content/about/news/246-v88-released-2.mdx
index 8b902e2..8d8c879 100644
--- a/projects/website-angular/content/about/news/article-13.mdx
+++ b/projects/website-angular/content/about/news/246-v88-released-2.mdx
@@ -2,10 +2,10 @@
title: V88 Released
category: "about"
date: "2024-03-20T00:35:16-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "246-v88-released-2"]
---
-## [ V88 Released ]()
+## V88 Released

diff --git a/projects/website-angular/content/about/news/article-12.mdx b/projects/website-angular/content/about/news/250-new-publication-in-database-oxford.mdx
similarity index 94%
rename from projects/website-angular/content/about/news/article-12.mdx
rename to projects/website-angular/content/about/news/250-new-publication-in-database-oxford.mdx
index 0f28095..d237c11 100644
--- a/projects/website-angular/content/about/news/article-12.mdx
+++ b/projects/website-angular/content/about/news/250-new-publication-in-database-oxford.mdx
@@ -2,10 +2,10 @@
title: "New Publication in Database (Oxford)!"
category: "about"
date: "2024-05-14T17:45:16-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "250-new-publication-in-database-oxford"]
---
-## [ New Publication in Database (Oxford)! ]()
+## New Publication in Database (Oxford)!
Introducing a new publication from the Reactome team in the Database (Oxford) journal by Orlic-Milacic et al, titled "[Pathway-based, reaction-specific annotation of disease variants for elucidation of molecular phenotypes]()"!
diff --git a/projects/website-angular/content/about/news/article-11.mdx b/projects/website-angular/content/about/news/251-v89-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-11.mdx
rename to projects/website-angular/content/about/news/251-v89-released.mdx
index a502711..ffbde44 100644
--- a/projects/website-angular/content/about/news/article-11.mdx
+++ b/projects/website-angular/content/about/news/251-v89-released.mdx
@@ -2,10 +2,10 @@
title: V89 Released
category: "about"
date: "2024-06-09T22:59:54-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "251-v89-released"]
---
-## [ V89 Released ]()
+## V89 Released

diff --git a/projects/website-angular/content/about/news/article-10.mdx b/projects/website-angular/content/about/news/261-v90-news.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-10.mdx
rename to projects/website-angular/content/about/news/261-v90-news.mdx
index 13d870b..4177e9d 100644
--- a/projects/website-angular/content/about/news/article-10.mdx
+++ b/projects/website-angular/content/about/news/261-v90-news.mdx
@@ -2,10 +2,10 @@
title: V90 Released
category: "about"
date: "2024-09-17T00:01:08-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "261-v90-news"]
---
-## [ V90 Released ]()
+## V90 Released

diff --git a/projects/website-angular/content/about/news/article-9.mdx b/projects/website-angular/content/about/news/265-v91-news.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-9.mdx
rename to projects/website-angular/content/about/news/265-v91-news.mdx
index e2f0dd4..c427804 100644
--- a/projects/website-angular/content/about/news/article-9.mdx
+++ b/projects/website-angular/content/about/news/265-v91-news.mdx
@@ -2,10 +2,10 @@
title: V91 Released
category: "about"
date: "2024-11-30T10:22:36-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "265-v91-news"]
---
-## [ V91 Released ]()
+## V91 Released

diff --git a/projects/website-angular/content/about/news/article-8.mdx b/projects/website-angular/content/about/news/270-v92-news.mdx
similarity index 99%
rename from projects/website-angular/content/about/news/article-8.mdx
rename to projects/website-angular/content/about/news/270-v92-news.mdx
index c1957fe..da2f426 100644
--- a/projects/website-angular/content/about/news/article-8.mdx
+++ b/projects/website-angular/content/about/news/270-v92-news.mdx
@@ -2,10 +2,10 @@
title: V92 Released
category: "about"
date: "2025-03-20T01:13:44-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "270-v92-news"]
---
-## [ V92 Released ]()
+## V92 Released

diff --git a/projects/website-angular/content/about/news/article-7.mdx b/projects/website-angular/content/about/news/273-coretrustseal-news.mdx
similarity index 91%
rename from projects/website-angular/content/about/news/article-7.mdx
rename to projects/website-angular/content/about/news/273-coretrustseal-news.mdx
index 3f27c35..b8113e2 100644
--- a/projects/website-angular/content/about/news/article-7.mdx
+++ b/projects/website-angular/content/about/news/273-coretrustseal-news.mdx
@@ -2,10 +2,10 @@
title: Reactome Recognized with CoreTrustSeal Certification
category: "about"
date: "2025-05-20T12:48:44-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "273-coretrustseal-news"]
---
-## [ Reactome Recognized with CoreTrustSeal Certification ]()
+## Reactome Recognized with CoreTrustSeal Certification
We are excited to share that the Reactome Knowledgebase has officially received CoreTrustSeal certification, a recognition awarded to data repositories that meet high standards for trustworthiness, sustainability, and open data practices.
diff --git a/projects/website-angular/content/about/news/article-6.mdx b/projects/website-angular/content/about/news/275-v93-released.mdx
similarity index 99%
rename from projects/website-angular/content/about/news/article-6.mdx
rename to projects/website-angular/content/about/news/275-v93-released.mdx
index 25a97fe..d6100d9 100644
--- a/projects/website-angular/content/about/news/article-6.mdx
+++ b/projects/website-angular/content/about/news/275-v93-released.mdx
@@ -2,10 +2,10 @@
title: V93 Released
category: "about"
date: "2025-06-12T13:09:20-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "275-v93-released"]
---
-## [ V93 Released ]()
+## V93 Released

diff --git a/projects/website-angular/content/about/news/article-5.mdx b/projects/website-angular/content/about/news/279-v94-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-5.mdx
rename to projects/website-angular/content/about/news/279-v94-released.mdx
index 924671a..1261124 100644
--- a/projects/website-angular/content/about/news/article-5.mdx
+++ b/projects/website-angular/content/about/news/279-v94-released.mdx
@@ -2,10 +2,10 @@
title: V94 Released
category: "about"
date: "2025-09-11T01:17:50-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "279-v94-released"]
---
-## [ V94 Released ]()
+## V94 Released

diff --git a/projects/website-angular/content/about/news/article-4.mdx b/projects/website-angular/content/about/news/280-reactome-pathway-browser-new-beta-release.mdx
similarity index 95%
rename from projects/website-angular/content/about/news/article-4.mdx
rename to projects/website-angular/content/about/news/280-reactome-pathway-browser-new-beta-release.mdx
index 598020b..e4b38a2 100644
--- a/projects/website-angular/content/about/news/article-4.mdx
+++ b/projects/website-angular/content/about/news/280-reactome-pathway-browser-new-beta-release.mdx
@@ -2,10 +2,10 @@
title: "Reactome Pathway Browser: New Beta Release"
category: "about"
date: "2025-09-18T08:34:09-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "280-reactome-pathway-browser-new-beta-release"]
---
-## [ Reactome Pathway Browser: New Beta Release ]()
+## Reactome Pathway Browser: New Beta Release
Reactome is excited to announce the beta release of our redesigned Pathway Browser, now available at: []()
diff --git a/projects/website-angular/content/about/news/article-3.mdx b/projects/website-angular/content/about/news/284-v95-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-3.mdx
rename to projects/website-angular/content/about/news/284-v95-released.mdx
index ad44be0..022b644 100644
--- a/projects/website-angular/content/about/news/article-3.mdx
+++ b/projects/website-angular/content/about/news/284-v95-released.mdx
@@ -2,10 +2,10 @@
title: V95 Released
category: "about"
date: "2025-12-03T18:22:40-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "284-v95-released"]
---
-## [ V95 Released ]()
+## V95 Released

diff --git a/projects/website-angular/content/about/news/article-2.mdx b/projects/website-angular/content/about/news/286-new-publication-in-nar-2026.mdx
similarity index 89%
rename from projects/website-angular/content/about/news/article-2.mdx
rename to projects/website-angular/content/about/news/286-new-publication-in-nar-2026.mdx
index ed6b156..63cb4c0 100644
--- a/projects/website-angular/content/about/news/article-2.mdx
+++ b/projects/website-angular/content/about/news/286-new-publication-in-nar-2026.mdx
@@ -2,10 +2,10 @@
title: "New Publication in NAR 2026:"
category: "about"
date: "2025-12-08T16:18:39-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "286-new-publication-in-nar-2026"]
---
-## [ New Publication in NAR 2026: ]()
+## New Publication in NAR 2026:
A new research article entitled “The Reactome Knowledgebase 2026” has been published in the Nucleic Acids Research 2026 Databases Issue. The paper highlights Reactome’s redesigned Angular-based interface, enhanced global and entity-level visualizations, multi-omics analysis tools, and innovations such as the React-to-Me chatbot. It also details Reactome’s community resources, FAIR data compliance, and recognition as a CoreTrustSeal-certified and ELIXIR Global Core Biodata resource.
diff --git a/projects/website-angular/content/about/news/article-1.mdx b/projects/website-angular/content/about/news/288-reactome-releases-two-new-ai-focused-preprints.mdx
similarity index 79%
rename from projects/website-angular/content/about/news/article-1.mdx
rename to projects/website-angular/content/about/news/288-reactome-releases-two-new-ai-focused-preprints.mdx
index b08f628..5324880 100644
--- a/projects/website-angular/content/about/news/article-1.mdx
+++ b/projects/website-angular/content/about/news/288-reactome-releases-two-new-ai-focused-preprints.mdx
@@ -2,22 +2,22 @@
title: "Reactome Releases Two New AI-Focused Preprints"
category: "about"
date: "2026-02-05T11:16:48-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "288-reactome-releases-two-new-ai-focused-preprints"]
---
-## [ Reactome Releases Two New AI-Focused Preprints ]()
+## Reactome Releases Two New AI-Focused Preprints
Reactome has released two preprints describing recent advances in applying artificial intelligence to pathway curation and user access.
The preprint, [_Application of Large Language Models for Annotating Genes into Reactome Pathways_](), presents an LLM-assisted workflow that supports expert curators by identifying candidate pathways for genes, retrieving relevant literature, and generating mechanistic summaries. Evaluation shows that the approach can meaningfully support manual curation while preserving curator oversight.
-
+
_Screenshots of the LLM App in the web-based Reactome curation tool for query gene input, configuration and annotated pathways._
The preprint, [_React-to-Me: A Conversational Interface for Interactive Exploration of the Reactome Pathway Knowledgebase_](), introduces a grounded conversational assistant that enables natural language queries over Reactome content. The system enforces source traceability to curated data and avoids speculative responses, improving accessibility without sacrificing factual accuracy.
-
+
_Graphical abstract showing React-to-Me._
diff --git a/projects/website-angular/content/about/news/article-87.mdx b/projects/website-angular/content/about/news/71-version-57-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-87.mdx
rename to projects/website-angular/content/about/news/71-version-57-released.mdx
index 281c526..59fd18d 100644
--- a/projects/website-angular/content/about/news/article-87.mdx
+++ b/projects/website-angular/content/about/news/71-version-57-released.mdx
@@ -2,10 +2,10 @@
title: Version 57 Released
category: "about"
date: "2016-06-27T13:13:06-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "71-version-57-released"]
---
-## [ Version 57 Released ]()
+## Version 57 Released
In version V57, topics with new or revised pathways include: Developmental biology ([RET signaling]()), Disease ([Signaling by FGFR in disease]() and [Defective CFTR causes cystic fibrosis]()), Gene expression ([rRNA modification in the mitochondrion]() and [Transcriptional regulation by the AP-2 (TFAP2) family of transcription factors]()), Immune system ([ER-Phagosome pathway](), [Interleukin-7 signaling](), [Regulation by endogenous TLR ligand](), and [TCR signaling]()), Metabolism ([Lipid digestion, mobilization, and transport]() and [Phosphate bond hydrolysis by NTPDase proteins]()). Metabolism of proteins ([Deubiquitination]()), Neuronal system ([Interactions of neurexins and neuroligins at synapses]() and [SALM protein interactions at synapse]()). Signal transduction ([EGFR downregulation](), [Signaling by FGFR1](), and [FGFRL1 modulation of FGFR1 signaling]()), Transmembrane transport of small molecules ([ABC-family proteins mediated transport]()) and Vesicle-mediated transport ([Clathrin-mediated endocytosis]()).
diff --git a/projects/website-angular/content/about/news/article-86.mdx b/projects/website-angular/content/about/news/72-version-58-released.mdx
similarity index 98%
rename from projects/website-angular/content/about/news/article-86.mdx
rename to projects/website-angular/content/about/news/72-version-58-released.mdx
index 9d90a2a..2449458 100644
--- a/projects/website-angular/content/about/news/article-86.mdx
+++ b/projects/website-angular/content/about/news/72-version-58-released.mdx
@@ -2,10 +2,10 @@
title: Version 58 Released
category: "about"
date: "2016-09-28T13:14:30-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "72-version-58-released"]
---
-## [ Version 58 Released ]()
+## Version 58 Released
**[]()**
diff --git a/projects/website-angular/content/about/news/article-85.mdx b/projects/website-angular/content/about/news/73-reactome-celebrates-release-of-10-000th-annotated-protein.mdx
similarity index 94%
rename from projects/website-angular/content/about/news/article-85.mdx
rename to projects/website-angular/content/about/news/73-reactome-celebrates-release-of-10-000th-annotated-protein.mdx
index fafa914..73e9425 100644
--- a/projects/website-angular/content/about/news/article-85.mdx
+++ b/projects/website-angular/content/about/news/73-reactome-celebrates-release-of-10-000th-annotated-protein.mdx
@@ -2,10 +2,10 @@
title: "Reactome celebrates release of 10,000th annotated protein"
category: "about"
date: "2017-08-20T13:15:27-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "73-reactome-celebrates-release-of-10-000th-annotated-protein"]
---
-## [ Reactome celebrates release of 10,000th annotated protein ]()
+## Reactome celebrates release of 10,000th annotated protein
[]()
diff --git a/projects/website-angular/content/about/news/article-84.mdx b/projects/website-angular/content/about/news/74-version-59-release.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-84.mdx
rename to projects/website-angular/content/about/news/74-version-59-release.mdx
index d6d0bda..a1b513b 100644
--- a/projects/website-angular/content/about/news/article-84.mdx
+++ b/projects/website-angular/content/about/news/74-version-59-release.mdx
@@ -2,10 +2,10 @@
title: Version 59 Released
category: "about"
date: "2016-12-21T13:16:00-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "74-version-59-release"]
---
-## [ Version 59 Released ]()
+## Version 59 Released
**New and Updated Pathways.** In version V59, topics with new or revised pathways include: Disease ([Listeria monocytogenes entry into host cells]()), Hemostasis ([Cell surface interaction at the vascular wall]()), Immune System ([Butyrophilins](), [Interleukin 10 signalling](), and [Interleukin-4 and 13 signaling]()), Metabolism ([Nicotinate metabolism](), [Synthesis of PIPs at the nuclear envelope](), [Vitamin B5 (pantothenate) metabolism](), and [Aryl hydrocarbon receptor signalling]()), Metabolism of proteins ([E3 ubiquitin ligases ubiquitinate target proteins](), [Peptide-ligand binding receptors](), [Protein methylation](), [RAB geranylgeranylation]()), Signal transduction ([Class A/1 (Rhodopsin-like receptors]()), and Vesicle-mediated transport ([TBC RABGAPs]()).
diff --git a/projects/website-angular/content/about/news/article-83.mdx b/projects/website-angular/content/about/news/75-new-reactome-publication.mdx
similarity index 81%
rename from projects/website-angular/content/about/news/article-83.mdx
rename to projects/website-angular/content/about/news/75-new-reactome-publication.mdx
index 704df73..900de01 100644
--- a/projects/website-angular/content/about/news/article-83.mdx
+++ b/projects/website-angular/content/about/news/75-new-reactome-publication.mdx
@@ -2,9 +2,9 @@
title: New Reactome Publication
category: "about"
date: "2017-08-20T13:16:36-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "75-new-reactome-publication"]
---
-## [ New Reactome Publication ]()
+## New Reactome Publication
A new Reactome paper titled “Functional Interaction Network Construction and Analysis for Disease Discovery” has been published in [Methods in Molecular Biology](). More publications from the [Reactome Team]() can be found [here]().
diff --git a/projects/website-angular/content/about/news/article-82.mdx b/projects/website-angular/content/about/news/76-new-reactome-paper-published.mdx
similarity index 79%
rename from projects/website-angular/content/about/news/article-82.mdx
rename to projects/website-angular/content/about/news/76-new-reactome-paper-published.mdx
index a0d0bad..f026028 100644
--- a/projects/website-angular/content/about/news/article-82.mdx
+++ b/projects/website-angular/content/about/news/76-new-reactome-paper-published.mdx
@@ -2,9 +2,9 @@
title: New Reactome Paper published
category: "about"
date: "2017-08-20T13:17:08-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "76-new-reactome-paper-published"]
---
-## [ New Reactome Paper published ]()
+## New Reactome Paper published
A new Reactome paper titled “Reactome pathway analysis: a high-performance in-memory approach” has been published in [BMC Bioinformatics](). More publications from the [Reactome Team]() can be found [here]().
diff --git a/projects/website-angular/content/about/news/article-81.mdx b/projects/website-angular/content/about/news/77-version-60-released.mdx
similarity index 99%
rename from projects/website-angular/content/about/news/article-81.mdx
rename to projects/website-angular/content/about/news/77-version-60-released.mdx
index d256e28..ed28938 100644
--- a/projects/website-angular/content/about/news/article-81.mdx
+++ b/projects/website-angular/content/about/news/77-version-60-released.mdx
@@ -2,10 +2,10 @@
title: Version 60 Released
category: "about"
date: "2017-04-20T13:18:03-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "77-version-60-released"]
---
-## [ Version 60 Released ]()
+## Version 60 Released
**New and Updated Pathways.****** In version V60, topics with new or revised pathways include: Cell cycle ([Cyclin D associated events in G1]()), Cell-cell communication [(SDK interactions]()), Cellular response to external stimuli ([HSP90 chaperone cycle for steroid hormone receptors (SHR)]()), Disease ([Diseases of Mismatch Repair]()), Gene Expression ([TP53 Regulates Transcription of Genes Involved in G1 Cell Cycle Arrest]() and [Transcriptional Regulation by the CBFB:RUNX3 complex]()), Immune System ([Butyrophilins]() and [Regulation of complement cascade]()), Metabolism ([Synthesis of IP2, IP, and Ins in the cytosol](), [Synthesis of PIPs at the early endosome membrane](), [Synthesis of PIPs at the ER membrane](), [Synthesis of PIPs at the late endosome membrane](), and [Synthesis of PIPs at the plasma membrane]()), Metabolism of proteins ([CREB3 factors activate genes](), [Neddylation](), [Protein ubiquitination](), and [SUMOylation of chromatin organizing proteins]()), Mitophagy ([Receptor Mediated Mitophagy]()), Neuronal System ([Receptor protein tyrosine phosphatases interactions]()), Organelle biogenesis and maintenance ([Cristae formation]()), and Transport of small molecules ([Mitochondrial calcium ion transport]()).
diff --git a/projects/website-angular/content/about/news/article-80.mdx b/projects/website-angular/content/about/news/78-version-61-released.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-80.mdx
rename to projects/website-angular/content/about/news/78-version-61-released.mdx
index 45d6d1f..fdc73df 100644
--- a/projects/website-angular/content/about/news/article-80.mdx
+++ b/projects/website-angular/content/about/news/78-version-61-released.mdx
@@ -2,10 +2,10 @@
title: Version 61 Released
category: "about"
date: "2017-06-22T13:20:12-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "78-version-61-released"]
---
-## [ Version 61 Released ]()
+## Version 61 Released
**[]()**
diff --git a/projects/website-angular/content/about/news/article-79.mdx b/projects/website-angular/content/about/news/79-new-protein-protein-interaction-files.mdx
similarity index 95%
rename from projects/website-angular/content/about/news/article-79.mdx
rename to projects/website-angular/content/about/news/79-new-protein-protein-interaction-files.mdx
index 67b7fc5..c785f9a 100644
--- a/projects/website-angular/content/about/news/article-79.mdx
+++ b/projects/website-angular/content/about/news/79-new-protein-protein-interaction-files.mdx
@@ -2,10 +2,10 @@
title: "New Protein-protein Interaction files"
category: "about"
date: "2017-06-28T13:24:21-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "79-new-protein-protein-interaction-files"]
---
-## [ New Protein-protein Interaction files ]()
+## New Protein-protein Interaction files

diff --git a/projects/website-angular/content/about/news/article-78.mdx b/projects/website-angular/content/about/news/80-new-sbml-level-3-version-1-export-is-now-available.mdx
similarity index 91%
rename from projects/website-angular/content/about/news/article-78.mdx
rename to projects/website-angular/content/about/news/80-new-sbml-level-3-version-1-export-is-now-available.mdx
index a5683ca..ce7dfc4 100644
--- a/projects/website-angular/content/about/news/article-78.mdx
+++ b/projects/website-angular/content/about/news/80-new-sbml-level-3-version-1-export-is-now-available.mdx
@@ -2,10 +2,10 @@
title: New SBML Level 3 Version 1 export is now available
category: "about"
date: "2017-07-05T13:25:06-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "80-new-sbml-level-3-version-1-export-is-now-available"]
---
-## [ New SBML Level 3 Version 1 export is now available ]()
+## New SBML Level 3 Version 1 export is now available
[]()
diff --git a/projects/website-angular/content/about/news/article-77.mdx b/projects/website-angular/content/about/news/90-version-62-released.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-77.mdx
rename to projects/website-angular/content/about/news/90-version-62-released.mdx
index c8f5b78..6feed77 100644
--- a/projects/website-angular/content/about/news/article-77.mdx
+++ b/projects/website-angular/content/about/news/90-version-62-released.mdx
@@ -2,10 +2,10 @@
title: Version 62 Released
category: "about"
date: "2017-09-28T11:45:38-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "90-version-62-released"]
---
-## [ Version 62 Released ]()
+## Version 62 Released
**[]()**
diff --git a/projects/website-angular/content/about/news/article-76.mdx b/projects/website-angular/content/about/news/93-reactome-launches-new-website.mdx
similarity index 92%
rename from projects/website-angular/content/about/news/article-76.mdx
rename to projects/website-angular/content/about/news/93-reactome-launches-new-website.mdx
index bebf95c..0f24ff2 100644
--- a/projects/website-angular/content/about/news/article-76.mdx
+++ b/projects/website-angular/content/about/news/93-reactome-launches-new-website.mdx
@@ -2,10 +2,10 @@
title: New responsive website with a fresh look
category: "about"
date: "2017-11-01T11:09:43-04:00"
-tags: ["about", "news"]
+tags: ["about", "news", "93-reactome-launches-new-website"]
---
-## [ New responsive website with a fresh look ]()
+## New responsive website with a fresh look

diff --git a/projects/website-angular/content/about/news/article-75.mdx b/projects/website-angular/content/about/news/94-proudly-introducing-our-new-logo.mdx
similarity index 90%
rename from projects/website-angular/content/about/news/article-75.mdx
rename to projects/website-angular/content/about/news/94-proudly-introducing-our-new-logo.mdx
index 4a559b7..6bdba69 100644
--- a/projects/website-angular/content/about/news/article-75.mdx
+++ b/projects/website-angular/content/about/news/94-proudly-introducing-our-new-logo.mdx
@@ -2,10 +2,10 @@
title: Proudly introducing our new logo
category: "about"
date: "2017-11-07T11:44:47-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "94-proudly-introducing-our-new-logo"]
---
-## [ Proudly introducing our new logo ]()
+## Proudly introducing our new logo

diff --git a/projects/website-angular/content/about/news/article-74.mdx b/projects/website-angular/content/about/news/95-new-reactome-publication-published-in-2017-nar-database-issue.mdx
similarity index 82%
rename from projects/website-angular/content/about/news/article-74.mdx
rename to projects/website-angular/content/about/news/95-new-reactome-publication-published-in-2017-nar-database-issue.mdx
index b0f7a74..44a6124 100644
--- a/projects/website-angular/content/about/news/article-74.mdx
+++ b/projects/website-angular/content/about/news/95-new-reactome-publication-published-in-2017-nar-database-issue.mdx
@@ -2,10 +2,10 @@
title: New Reactome Publication published in 2018 NAR Database Issue
category: "about"
date: "2017-11-14T23:02:20-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "95-new-reactome-publication-published-in-2017-nar-database-issue"]
---
-## [ New Reactome Publication published in 2018 NAR Database Issue ]()
+## New Reactome Publication published in 2018 NAR Database Issue

diff --git a/projects/website-angular/content/about/news/article-73.mdx b/projects/website-angular/content/about/news/97-updated-license-agreement.mdx
similarity index 95%
rename from projects/website-angular/content/about/news/article-73.mdx
rename to projects/website-angular/content/about/news/97-updated-license-agreement.mdx
index 0b6fb3f..4d2496c 100644
--- a/projects/website-angular/content/about/news/article-73.mdx
+++ b/projects/website-angular/content/about/news/97-updated-license-agreement.mdx
@@ -2,10 +2,10 @@
title: Updated License Agreement
category: "about"
date: "2017-12-06T23:09:22-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "97-updated-license-agreement"]
---
-## [ Updated License Agreement ]()
+## Updated License Agreement

diff --git a/projects/website-angular/content/about/news/article-72.mdx b/projects/website-angular/content/about/news/98-version-63-released.mdx
similarity index 97%
rename from projects/website-angular/content/about/news/article-72.mdx
rename to projects/website-angular/content/about/news/98-version-63-released.mdx
index 0cd64ed..5f204ca 100644
--- a/projects/website-angular/content/about/news/article-72.mdx
+++ b/projects/website-angular/content/about/news/98-version-63-released.mdx
@@ -2,10 +2,10 @@
title: Version 63 Released
category: "about"
date: "2017-12-18T15:38:11-05:00"
-tags: ["about", "news"]
+tags: ["about", "news", "98-version-63-released"]
---
-## [ Version 63 Released ]()
+## Version 63 Released
[]()
diff --git a/projects/website-angular/content/about/news/article-46.mdx b/projects/website-angular/content/about/news/article-46.mdx
deleted file mode 100644
index a85a37b..0000000
--- a/projects/website-angular/content/about/news/article-46.mdx
+++ /dev/null
@@ -1,14 +0,0 @@
----
-title: "ReacFoam: Genome-wide pathway overview based on Voronoi tessellation"
-category: "about"
-date: "2019-07-02T07:38:13-04:00"
-tags: ["about", "news"]
----
-
-## [ ReacFoam: Genome-wide pathway overview based on Voronoi tessellation ]()
-
-[]()
-
-As part of our continuous effort to provide visually attractive and more user-friendly access to our biological pathways, we are pleased to announce our new high level pathways overview visualisation based on Voronoi tessellation.
-
-Following any [pathway analysis](), the [pathway overview]() provides a [new icon]() which leads to a comprehensive, highly visual, interactive overview of pathway analysis results. The functionality is available on both desktop and mobile platforms.
diff --git a/projects/website-angular/content/about/news/article-48.mdx b/projects/website-angular/content/about/news/article-48.mdx
deleted file mode 100644
index 7784ddf..0000000
--- a/projects/website-angular/content/about/news/article-48.mdx
+++ /dev/null
@@ -1,7 +0,0 @@
----
-title: Claim Your Work
-category: "about"
-tags: ["about", "news"]
----
-
-Follow this [link]( "Click for instructions on how to claim your work") to learn more about how you can claim your work.
diff --git a/projects/website-angular/content/community/collaboration.mdx b/projects/website-angular/content/community/collaboration/collaboration.mdx
similarity index 100%
rename from projects/website-angular/content/community/collaboration.mdx
rename to projects/website-angular/content/community/collaboration/collaboration.mdx
diff --git a/projects/website-angular/content/community/collaboration/faq-for-prospective-reviewers-and-authors.mdx b/projects/website-angular/content/community/collaboration/faq-for-prospective-reviewers-and-authors.mdx
new file mode 100644
index 0000000..a44f3f5
--- /dev/null
+++ b/projects/website-angular/content/community/collaboration/faq-for-prospective-reviewers-and-authors.mdx
@@ -0,0 +1,50 @@
+---
+title: FAQ for prospective Reactome reviewers and authors
+category: "community"
+---
+
+## FAQ for prospective Reactome reviewers and authors
+
+**What is involved in reviewing a Reactome pathway module?**
+
+Reviewing a pathway module is similar to reviewing a review article and involves evaluating both a pathway report in text format and a corresponding online pathway diagram for completeness and accuracy. A Reactome pathway is a hierarchical representation of a biological process. Pathways are broken down into component subpathways. A subpathway is further subdivided into its component biochemical reactions, and each reaction includes input and output molecules as well as any relevant catalyst or regulators. The text report includes a summary of each event (pathway, subpathway or reaction), a list of its supporting references, and a link to the corresponding event in our pathway browser on our development website. This pathway browser web page, shown below, includes an Event Hierarchy as well as a Pathway Diagram and Event Details section.
+
+
+
+In the pathway diagram, reactions are manually laid out showing their relationships to one another. The details section of this webpage provides more detailed dynamic descriptions of the molecular composition and hierarchical organization of reactions.
+
+Selecting any event in the hierarchy (left panel) will bring you to its location in the pathway diagram, and the corresponding event(s) will be highlighted in blue in the diagram. A text and molecular description of the reaction or event can be viewed in the Details tab in the panel below the diagram. The description of the reaction may also be accessed by clicking on the reaction “node” in the diagram. A text description (and cross-references) for individual reaction component molecules can be displayed by selecting the molecule of interest in the diagram. More detailed instructions for navigating the website can be found [here]().
+
+We are asking reviewers to verify that the pathways and reactions described in the text document are annotated clearly and completely and that the molecular details of the reactions (described in-depth on the webpages) are accurate. Additional instructions for navigating the web pages can be found here. We would appreciate any comments or suggestions on the user interface as well.
+
+**How long does it take to review a module?**
+
+Depending on the size of the module, reviewers take anywhere from a day to a month for their reviews. Typically, reviewers provide their feedback within two or three weeks. Reactome has a rolling quarterly release cycle though, so if your review takes a bit longer than expected, this is not a problem. We can include the revised module in the next release.
+
+**If I’ve decided to review a module, how do I get started?**
+
+If you’re ready to get started reviewing, send us an email at This email address is being protected from spambots. You need JavaScript enabled to view it. and we can put you in contact with the curator of the pathway module. You can communicate with this curator if you have any questions and when you’d like to submit your review.
+
+**When can I see my work published on the Reactome website?**
+
+Reactome has a quarterly database release cycle, generally in March, June, September, and December. For a revised module to be included in a given release, we ask that reviews be submitted 6 weeks (or more) before the planned release date. So for a revised module to be included in a March release, we would need to have the review back in mid-January.
+
+**How can I provide feedback and suggestions to Reactome?**
+
+Comments may be added directly to the word document provided. Please opt to “tracking changes” so that your comments and suggested changes are highlighted. If you prefer, you can send your review as a separate document.
+
+**What if my work (or other important work) has not been included or cited?**
+
+Due to limited resources, we may not have included all of the references that are relevant for a particular pathway or reaction, but we would be happy to add any that you might wish to see included. Likewise, if you would like to expand a pathway module to include events that we have not covered, we would be glad to work with you on this and would cite you as an author for any new content.
+
+**What if I have questions while reviewing a pathway module?**
+
+We will pair you with the curator that has worked on the pathway module that you are reviewing. You can communicate directly with this curator for any questions. You can also contact us at This email address is being protected from spambots. You need JavaScript enabled to view it. for any technical questions.
+
+**How am I acknowledged for my contribution to Reactome?**
+
+Your review will be used by a Reactome curator to revise the pathway module. You will be acknowledged as a reviewer of this pathway on the web pages and in the Reactome table of contents. This pathway module will also be associated with a DOI and may be cited as a publication. We ask that you provide us with an ORCID identifier so that we can associate this with the pathways that you have contributed. Our main search allows our authors and reviewers to [query]() for their pathway and reaction contributions using their names, and claim these contributions directly into ORCID.
+
+**Spread the word!**
+
+Reactome is always looking for new contributors! If you know someone that would be interested in contributing a new pathway or reviewing/revising an existing pathway module, please let them know about us and our mission.
diff --git a/projects/website-angular/content/community/collaboration/index.mdx b/projects/website-angular/content/community/collaboration/index.mdx
deleted file mode 100644
index a1c11a7..0000000
--- a/projects/website-angular/content/community/collaboration/index.mdx
+++ /dev/null
@@ -1,10 +0,0 @@
----
-title: Contribute Pathway Knowledge
-category: "community"
----
-
-## Contribute Pathway Knowledge
-
-Become a Reactome contributor! You will be credited with authorship or reviewership for all of your contributions. Each pathway is associated with a DOI and can be cited as a publication. You can quickly and easily claim your Reactome contributions in ORCID using our new ORCID claiming feature when you search for your name in Reactome as described [here]().
-
-We are currently seeking reviewers for the following [pathways](). If you'd like to contribute a pathway that is not on this list, please contact us. We would be happy to work with you! For more information and guidelines for reviewing a Reactome pathway module, see this [FAQ for prospective Reactome reviewers and authors]().
diff --git a/projects/website-angular/content/community/events.mdx b/projects/website-angular/content/community/events.mdx
index da8153b..a775435 100644
--- a/projects/website-angular/content/community/events.mdx
+++ b/projects/website-angular/content/community/events.mdx
@@ -17,7 +17,7 @@ HUPO, November 9-13th, 2025
Disease Maps Community Meeting April 15-17th , 2025
-International Society of Biocuration, April 5-9th, 2025.[ Poster.]()
EMBL-EBI Training: Introduction to RNA-seq and functional interpretation, Feb 24 2024
-Keystone Symposia: Systems and Engineered Biology. The Reactome Knowledgebase: a resource for systems immunology and innate immunity research April 9, 2024. [Poster]()
@@ -119,7 +119,7 @@ UniAndes: Reactome Introduction
EMBL-EBI network resources
-ISCB RSGDream2024 Conference. [Link](). [Poster](). [Poster](), the [Graph Database]() or any of our widgets ([diagram]() or [pathways overview]()). If your resource is using our services, widgets or content and it is not in the list, please This email address is being protected from spambots. You need JavaScript enabled to view it. and we will add it.
diff --git a/projects/website-angular/content/community/resources.mdx b/projects/website-angular/content/community/resources.mdx
new file mode 100644
index 0000000..f843b4a
--- /dev/null
+++ b/projects/website-angular/content/community/resources.mdx
@@ -0,0 +1,12 @@
+---
+title: Resources Guide
+category: "community"
+---
+
+## Resources Guide
+
+We actively seek collaborations with other data resources and users to improve data integration, share efforts, and make our data maximally useful to biologists and bioinformaticians. Here are a list of software tools, websites, databases and research projects that: i) support the use of Reactome data, ii) have integrated Reactome data, or iii) provide data linkages to the Reactome website.
+
+If we are missing your website, database, software tool and project, please contact our []()This email address is being protected from spambots. You need JavaScript enabled to view it..
+
+
diff --git a/projects/website-angular/content/content/index.mdx b/projects/website-angular/content/content/index.mdx
deleted file mode 100644
index 1b26cb8..0000000
--- a/projects/website-angular/content/content/index.mdx
+++ /dev/null
@@ -1,7 +0,0 @@
----
-title: Content
-description: dewszc
-category: content
----
-
-This is where you land cotnetn
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/180-reactome-spotlight.mdx b/projects/website-angular/content/content/reactome-research-spotlight/180-reactome-spotlight.mdx
new file mode 100644
index 0000000..d52c91b
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/180-reactome-spotlight.mdx
@@ -0,0 +1,10 @@
+---
+title: "Post-infusion CAR TReg cells identify patients resistant to CD19-CAR therapy"
+category: "content"
+date: "2022-10-18T10:26:35-04:00"
+tags: ["content", "reactome-research-spotlight", "180-reactome-spotlight"]
+---
+
+## Post-infusion CAR TReg cells identify patients resistant to CD19-CAR therapy
+
+Reactome pathway enrichment analysis helps to pinpoint expansion of regulatory T cells as a new biomarker of CAR T cell therapy resistance and toxicity in patients with B cell lymphoma. The study was published by [Good et al. in Nature Medicine on September 12, 2022]().
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/186-reactome-pathway-gene-sets-in-the-msigdb-facilitated-identification-of-the-liver-proteasome-transcriptional-switch-that-acts-as-the-fasting-timer-in-intermittent-fasting.mdx b/projects/website-angular/content/content/reactome-research-spotlight/186-reactome-pathway-gene-sets-in-the-msigdb-facilitated-identification-of-the-liver-proteasome-transcriptional-switch-that-acts-as-the-fasting-timer-in-intermittent-fasting.mdx
new file mode 100644
index 0000000..ce1f626
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/186-reactome-pathway-gene-sets-in-the-msigdb-facilitated-identification-of-the-liver-proteasome-transcriptional-switch-that-acts-as-the-fasting-timer-in-intermittent-fasting.mdx
@@ -0,0 +1,10 @@
+---
+title: Circadian transcriptional pathway atlas highlights a proteasome switch in intermittent fasting
+category: "content"
+date: "2022-12-12T11:41:07-05:00"
+tags: ["content", "reactome-research-spotlight", "186-reactome-pathway-gene-sets-in-the-msigdb-facilitated-identification-of-the-liver-proteasome-transcriptional-switch-that-acts-as-the-fasting-timer-in-intermittent-fasting"]
+---
+
+## Circadian transcriptional pathway atlas highlights a proteasome switch in intermittent fasting
+
+Reactome pathway gene sets in the MSigDB facilitated identification of the liver proteasome transcriptional switch that acts as the fasting timer in intermittent fasting in work published by [Wei et al. in Cell Reports on October 25, 2022](). The authors suggest that a 16-hour interval in intermittent fasting may be most beneficial for health.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/188-gene-set-enrichment-analysis-gsea-identifies-the-two-most-frequently-upregulated-carbohydrate-metabolism-pathways-in-tumors-with-high-tumor-specific-total-mrna-expression-tms.mdx b/projects/website-angular/content/content/reactome-research-spotlight/188-gene-set-enrichment-analysis-gsea-identifies-the-two-most-frequently-upregulated-carbohydrate-metabolism-pathways-in-tumors-with-high-tumor-specific-total-mrna-expression-tms.mdx
new file mode 100644
index 0000000..761863c
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/188-gene-set-enrichment-analysis-gsea-identifies-the-two-most-frequently-upregulated-carbohydrate-metabolism-pathways-in-tumors-with-high-tumor-specific-total-mrna-expression-tms.mdx
@@ -0,0 +1,10 @@
+---
+title: "Gene set enrichment analysis (GSEA) identifies upregulated carbohydrate metabolism pathways in tumors with high tumor-specific total mRNA expression (TmS)"
+category: "content"
+date: "2023-01-10T14:47:21-05:00"
+tags: ["content", "reactome-research-spotlight", "188-gene-set-enrichment-analysis-gsea-identifies-the-two-most-frequently-upregulated-carbohydrate-metabolism-pathways-in-tumors-with-high-tumor-specific-total-mrna-expression-tms"]
+---
+
+## Gene set enrichment analysis (GSEA) identifies upregulated carbohydrate metabolism pathways in tumors with high tumor-specific total mRNA expression (TmS)
+
+Gene set enrichment analysis (GSEA) conducted on Reactome’s carbohydrate metabolism pathways identifies the [Pentose phosphate pathway]() and the [Glucose metabolism ]()pathway as the two most frequently upregulated pathways in tumors with high tumor-specific total mRNA expression (TmS) across 15 tumor types; TmS is a novel tumor phenotype-predictive quantitative feature described by [Cao et al. in the November 2022 issue of Nature Biotechnology]().
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/189-common-targetable-inflammatory-pathways-in-brain-transcriptome-of-autism-spectrum-disorders-and-tourette-syndrome.mdx b/projects/website-angular/content/content/reactome-research-spotlight/189-common-targetable-inflammatory-pathways-in-brain-transcriptome-of-autism-spectrum-disorders-and-tourette-syndrome.mdx
new file mode 100644
index 0000000..f3f5f7c
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/189-common-targetable-inflammatory-pathways-in-brain-transcriptome-of-autism-spectrum-disorders-and-tourette-syndrome.mdx
@@ -0,0 +1,10 @@
+---
+title: Common targetable inflammatory pathways in brain transcriptome of autism spectrum disorders and Tourette syndrome
+category: "content"
+date: "2023-02-14T08:23:23-05:00"
+tags: ["content", "reactome-research-spotlight", "189-common-targetable-inflammatory-pathways-in-brain-transcriptome-of-autism-spectrum-disorders-and-tourette-syndrome"]
+---
+
+## Common targetable inflammatory pathways in brain transcriptome of autism spectrum disorders and Tourette syndrome
+
+Reactome overrepresentation analyses of differentially expressed genes common to both Autism Spectrum Disorder and Tourette Syndrome help identify common targetable inflammatory pathways as described by [Alshammeryet al. in the December 2022 issue of Frontiers in Neuroscience.]()
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/190-probable-treatment-targets-for-diabetic-retinopathy-based-on-an-integrated-proteomic-and-genomic-analysis.mdx b/projects/website-angular/content/content/reactome-research-spotlight/190-probable-treatment-targets-for-diabetic-retinopathy-based-on-an-integrated-proteomic-and-genomic-analysis.mdx
new file mode 100644
index 0000000..1dbbfb3
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/190-probable-treatment-targets-for-diabetic-retinopathy-based-on-an-integrated-proteomic-and-genomic-analysis.mdx
@@ -0,0 +1,10 @@
+---
+title: Probable Treatment Targets for Diabetic Retinopathy Based on an Integrated Proteomic and Genomic Analysis
+category: "content"
+date: "2023-03-13T16:29:19-04:00"
+tags: ["content", "reactome-research-spotlight", "190-probable-treatment-targets-for-diabetic-retinopathy-based-on-an-integrated-proteomic-and-genomic-analysis"]
+---
+
+## Probable Treatment Targets for Diabetic Retinopathy Based on an Integrated Proteomic and Genomic Analysis
+
+Analysis of all constituents of entire Reactome pathways identified by the presence of upregulated or mutated genes helped [Valdivia et al. in the February, 2023 issue of Translational Vision Science & Technology]() to identify druggable targets and potential drugs for the treatment of diabetic retinopathy (DR). Drugs affecting MMP13 and LGALS3 in the regulation of myeloid cell differentiation by RUNX2 were notable candidates.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/200-patient-derived-cell-based-pharmacogenomic-assessment-to-unveil-underlying-resistance-mechanisms-and-novel-therapeutics-for-advanced-lung-cancer.mdx b/projects/website-angular/content/content/reactome-research-spotlight/200-patient-derived-cell-based-pharmacogenomic-assessment-to-unveil-underlying-resistance-mechanisms-and-novel-therapeutics-for-advanced-lung-cancer.mdx
new file mode 100644
index 0000000..ef79e19
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/200-patient-derived-cell-based-pharmacogenomic-assessment-to-unveil-underlying-resistance-mechanisms-and-novel-therapeutics-for-advanced-lung-cancer.mdx
@@ -0,0 +1,10 @@
+---
+title: "Patient-derived cell-based pharmacogenomic assessment to unveil underlying resistance mechanisms and novel therapeutics for advanced lung cancer"
+category: "content"
+date: "2023-04-11T13:42:02-04:00"
+tags: ["content", "reactome-research-spotlight", "200-patient-derived-cell-based-pharmacogenomic-assessment-to-unveil-underlying-resistance-mechanisms-and-novel-therapeutics-for-advanced-lung-cancer"]
+---
+
+## Patient-derived cell-based pharmacogenomic assessment to unveil underlying resistance mechanisms and novel therapeutics for advanced lung cancer
+
+The Reactome database helped [Yu et al. in the January 2023 issue of the Journal of Experimental & Clinical Cancer Research]() identify candidate drugs for treatment of four subtypes of lung cancer that were categorized by pharmaco-genomic analysis of patient-derived cells and correlated with drug sensitivity of the patient-derived cells and with Reactome pathways identified by gene set variation analysis.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/226-label-free-mass-spectrometry-proteomics-reveals-different-pathways-modulated-in-thp-1-cells-infected-with-therapeutic-failure-and-drug-resistance-leishmania-infantum-clinical-isolates.mdx b/projects/website-angular/content/content/reactome-research-spotlight/226-label-free-mass-spectrometry-proteomics-reveals-different-pathways-modulated-in-thp-1-cells-infected-with-therapeutic-failure-and-drug-resistance-leishmania-infantum-clinical-isolates.mdx
new file mode 100644
index 0000000..e8daa95
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/226-label-free-mass-spectrometry-proteomics-reveals-different-pathways-modulated-in-thp-1-cells-infected-with-therapeutic-failure-and-drug-resistance-leishmania-infantum-clinical-isolates.mdx
@@ -0,0 +1,10 @@
+---
+title: "Label-Free Mass Spectrometry Proteomics Reveals Different Pathways Modulated in THP‐1 Cells Infected with Therapeutic Failure and Drug Resistance Leishmania infantum Clinical Isolates"
+category: "content"
+date: "2023-06-13T09:48:16-04:00"
+tags: ["content", "reactome-research-spotlight", "226-label-free-mass-spectrometry-proteomics-reveals-different-pathways-modulated-in-thp-1-cells-infected-with-therapeutic-failure-and-drug-resistance-leishmania-infantum-clinical-isolates"]
+---
+
+## Label-Free Mass Spectrometry Proteomics Reveals Different Pathways Modulated in THP‐1 Cells Infected with Therapeutic Failure and Drug Resistance Leishmania infantum Clinical Isolates
+
+[Tagliazucchi L et al. 2023 in the March 2023 issue of ACS Infectious Diseases ]()used the REACTOME overrepresentation and pathway topology analyses to identify [Transport of small molecules](), [Cellular response to stress]() and other pathways associated with drug resistance and therapeutic failure (TF) during _Leishmania infantum_ infection; they also discovered NDK3 and TFRC as potential targets for host-directed anti-Leishmania therapies to overcome drug-resistance.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/227-severe-covid-19-in-pregnancy-has-a-distinct-serum-profile-including-greater-complement-activation-and-dysregulation-of-serum-lipids.mdx b/projects/website-angular/content/content/reactome-research-spotlight/227-severe-covid-19-in-pregnancy-has-a-distinct-serum-profile-including-greater-complement-activation-and-dysregulation-of-serum-lipids.mdx
new file mode 100644
index 0000000..6eb6931
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/227-severe-covid-19-in-pregnancy-has-a-distinct-serum-profile-including-greater-complement-activation-and-dysregulation-of-serum-lipids.mdx
@@ -0,0 +1,10 @@
+---
+title: "Severe COVID-19 in pregnancy has a distinct serum profile, including greater complement activation and dysregulation of serum lipids"
+category: "content"
+date: "2023-06-13T10:02:39-04:00"
+tags: ["content", "reactome-research-spotlight", "227-severe-covid-19-in-pregnancy-has-a-distinct-serum-profile-including-greater-complement-activation-and-dysregulation-of-serum-lipids"]
+---
+
+## Severe COVID-19 in pregnancy has a distinct serum profile, including greater complement activation and dysregulation of serum lipids
+
+Pregnancies complicated by Coronavirus Disease 2019 (COVID-19) are at an increased risk of severe morbidity. In multi-omics analyses investigating the pathophysiology behind severe COVID-19 disease, [Altendahl et al, in the November 2022 issue of PLoS One]() found precipitous changes in maternal serum in those with severe COVID-19 infection. Reactome pathway enrichment analysis revealed upregulated analytes in 4 pathways: [Complement cascade](), [Signaling by the B Cell Receptor (BCR)](), [Fc epsilon receptor (FCERI) signaling](), and [ FCGR activation]().
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/228-in-vitro-zika-virus-infection-of-human-neural-progenitor-cells-meta-analysis-of-rna-seq-assays.mdx b/projects/website-angular/content/content/reactome-research-spotlight/228-in-vitro-zika-virus-infection-of-human-neural-progenitor-cells-meta-analysis-of-rna-seq-assays.mdx
new file mode 100644
index 0000000..cf40acb
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/228-in-vitro-zika-virus-infection-of-human-neural-progenitor-cells-meta-analysis-of-rna-seq-assays.mdx
@@ -0,0 +1,10 @@
+---
+title: "In Vitro Zika Virus Infection of Human Neural Progenitor Cells: Meta-Analysis of RNA-Seq Assays"
+category: "content"
+date: "2023-07-14T17:08:49-04:00"
+tags: ["content", "reactome-research-spotlight", "228-in-vitro-zika-virus-infection-of-human-neural-progenitor-cells-meta-analysis-of-rna-seq-assays"]
+---
+
+## In Vitro Zika Virus Infection of Human Neural Progenitor Cells: Meta-Analysis of RNA-Seq Assays
+
+The Zika virus (ZIKV) is an emergent arthropod-borne virus (arbovirus) responsible for congenital Zika syndrome (CZS) and a range of other congenital malformations. With little known about the pathways involved in CZS, [Gratton et al in the February 2020 issue of Microorganisms]() conducted a meta-analysis of transcriptome studies to identify the genes and pathways altered during Zika infection. Reactome analysis identified interferon, pro-inflammatory, and chemokines signaling as well as apoptosis as key IFN signaling pathways in ZIKV-infected cells with three new candidate genes involved in hNPCs infection identified: APOL6, XAF1, and TNFRSF1.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/229-genetic-networks-of-alzheimer-s-disease-aging-and-longevity-in-humans.mdx b/projects/website-angular/content/content/reactome-research-spotlight/229-genetic-networks-of-alzheimer-s-disease-aging-and-longevity-in-humans.mdx
new file mode 100644
index 0000000..a553cb5
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/229-genetic-networks-of-alzheimer-s-disease-aging-and-longevity-in-humans.mdx
@@ -0,0 +1,10 @@
+---
+title: "Genetic Networks of Alzheimer’s Disease, Aging, and Longevity in Humans"
+category: "content"
+date: "2023-08-11T13:49:31-04:00"
+tags: ["content", "reactome-research-spotlight", "229-genetic-networks-of-alzheimer-s-disease-aging-and-longevity-in-humans"]
+---
+
+## Genetic Networks of Alzheimer’s Disease, Aging, and Longevity in Humans
+
+Using Reactome analysis tools and FIVIz, [Balmorez et al. in the March 2023 issue of _Int. J. Mol. Sci._](), established a commonality between the genes and pathways associated with Alzheimer's disease (AD), Ageing (AR) and Longevity. The pathways shared between AD and AR are [p53-Dependent G1/S DNA damage checkpoint](), [FOXO-mediated transcription](), and [SUMOylation](); between AD and longevity are [Cytokine Signaling in Immune system](), [Plasma lipoprotein assembly, remodeling, and clearance](), [Metabolism of fat-soluble vitamins](), and [NR1H2- and NR1H3-mediated signalling](); and between AR and Longevity are [Immune system]() and [Cytokine signaling]().
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/232-computational-drug-repositioning-of-clopidogrel-as-a-novel-therapeutic-option-for-focal-segmental-glomerulosclerosis.mdx b/projects/website-angular/content/content/reactome-research-spotlight/232-computational-drug-repositioning-of-clopidogrel-as-a-novel-therapeutic-option-for-focal-segmental-glomerulosclerosis.mdx
new file mode 100644
index 0000000..91a022d
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/232-computational-drug-repositioning-of-clopidogrel-as-a-novel-therapeutic-option-for-focal-segmental-glomerulosclerosis.mdx
@@ -0,0 +1,11 @@
+---
+title: Computational drug repositioning of clopidogrel as a novel therapeutic option for focal segmental glomerulosclerosis
+category: "content"
+date: "2023-09-07T15:56:12-04:00"
+tags: ["content", "reactome-research-spotlight", "232-computational-drug-repositioning-of-clopidogrel-as-a-novel-therapeutic-option-for-focal-segmental-glomerulosclerosis"]
+---
+
+## Computational drug repositioning of clopidogrel as a novel therapeutic option for focal segmental glomerulosclerosis
+
+
+With current treatments, focal segmental glomerulosclerosis (FSGS), the largest cause of nephrotic syndrome, frequently progresses to end-stage kidney disease.[ Gebeshuber et al. (2023)]() assembled 376 FSGS-associated proteins into a FSGS pathophysiology model, major components of which were Reactome pathways for[ signal transduction]() and[ hemostasis](). The 39 proteins shared between FSGS model and a 102-protein model for the antiplatelet drug clopidogrel included 20 therapeutic targets of the drug. Tested in an FSGS mouse model, clopidogrel significantly attenuated disease severity, repositioning the drug as an attractive candidate for human clinical trials for FSGS.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/233-dna-methylation-and-28-year-cardiovascular-disease-risk-in-type-1-diabetes-the-epidemiology-of-diabetes-complications-edc-cohort-study.mdx b/projects/website-angular/content/content/reactome-research-spotlight/233-dna-methylation-and-28-year-cardiovascular-disease-risk-in-type-1-diabetes-the-epidemiology-of-diabetes-complications-edc-cohort-study.mdx
new file mode 100644
index 0000000..de10553
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/233-dna-methylation-and-28-year-cardiovascular-disease-risk-in-type-1-diabetes-the-epidemiology-of-diabetes-complications-edc-cohort-study.mdx
@@ -0,0 +1,10 @@
+---
+title: "DNA methylation and 28-year cardiovascular disease risk in type 1 diabetes: the Epidemiology of Diabetes Complications (EDC) cohort study"
+category: "content"
+date: "2023-10-22T23:06:05-04:00"
+tags: ["content", "reactome-research-spotlight", "233-dna-methylation-and-28-year-cardiovascular-disease-risk-in-type-1-diabetes-the-epidemiology-of-diabetes-complications-edc-cohort-study"]
+---
+
+## DNA methylation and 28-year cardiovascular disease risk in type 1 diabetes: the Epidemiology of Diabetes Complications (EDC) cohort study
+
+In the [ August 2, 2023 issue of Clinical Epigenetics](), Miller et al. performed an epigenome-wide association study using Reactome Functional Interaction network analysis and determined that DNA methylation at loci involved in calcium channel activity and development was associated with long-term cardiovascular disease risk beyond known risk factors in type 1 diabetes, particularly in individuals with greater glycemic exposure.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/235-xmr-an-explainable-multimodal-neural-network-for-drug-response-prediction.mdx b/projects/website-angular/content/content/reactome-research-spotlight/235-xmr-an-explainable-multimodal-neural-network-for-drug-response-prediction.mdx
new file mode 100644
index 0000000..0951591
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/235-xmr-an-explainable-multimodal-neural-network-for-drug-response-prediction.mdx
@@ -0,0 +1,10 @@
+---
+title: "XMR: an explainable multimodal neural network for drug response prediction"
+category: "content"
+date: "2023-11-02T12:57:25-04:00"
+tags: ["content", "reactome-research-spotlight", "235-xmr-an-explainable-multimodal-neural-network-for-drug-response-prediction"]
+---
+
+## XMR: an explainable multimodal neural network for drug response prediction
+
+In their paper titled “[XMR: an explainable multimodal neural network for drug response prediction]()” published in Frontiers in Bioinformatics in August 2023, Wang et al. use five Reactome pathways, [Cell Cycle](), [DNA repair](), [Disease](), [Signal transduction](), and [Metabolism](), as an architecture of a visible neural network that is part of a deep learning model for prediction of drug responses in triple negative breast cancer.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/237-new-insights-into-clinical-management-for-sickle-cell-disease-uncovering-the-significant-pathways-affected-by-the-involvement-of-sickle-cell-disease.mdx b/projects/website-angular/content/content/reactome-research-spotlight/237-new-insights-into-clinical-management-for-sickle-cell-disease-uncovering-the-significant-pathways-affected-by-the-involvement-of-sickle-cell-disease.mdx
new file mode 100644
index 0000000..9507302
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/237-new-insights-into-clinical-management-for-sickle-cell-disease-uncovering-the-significant-pathways-affected-by-the-involvement-of-sickle-cell-disease.mdx
@@ -0,0 +1,10 @@
+---
+title: "New Insights into Clinical Management for Sickle Cell Disease: Uncovering the Significant Pathways Affected by the Involvement of Sickle Cell Disease"
+category: "content"
+date: "2023-11-29T15:19:05-05:00"
+tags: ["content", "reactome-research-spotlight", "237-new-insights-into-clinical-management-for-sickle-cell-disease-uncovering-the-significant-pathways-affected-by-the-involvement-of-sickle-cell-disease"]
+---
+
+## New Insights into Clinical Management for Sickle Cell Disease: Uncovering the Significant Pathways Affected by the Involvement of Sickle Cell Disease
+
+In the chapter entitled “[New Insights into Clinical Management for Sickle Cell Disease: Uncovering the Significant Pathways Affected by the Involvement of Sickle Cell Disease]()”, published in Methods in Molecular Biology 2024, Chouhan et al. describe the use of Reactome FIviz Cytoscape plugin to analyze pathway enrichment and construct a functional interaction network for DisGNET-derived sickle cell disease-associated genes, identifying genes involved in “[Glucuronidation]()”, “[Aspirin ADME]()”, “[Phase II-Conjugation of compounds]()”, “[Interleukin-4 and interleukin-13 signaling]()”, “[Interleukin-10 signaling]()”, “[Signaling by interleukins]()”, “[Biological oxidations]()”, and “[Cytokine signaling in immune system”]().
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/240-machine-learning-based-analysis-of-cancer-cell-derived-vesicular-proteins-revealed-significant-tumor-specificity-and-predictive-potential-of-extracellular-vesicles-for-cell-invasion-and-proliferation-a-meta-analysis.mdx b/projects/website-angular/content/content/reactome-research-spotlight/240-machine-learning-based-analysis-of-cancer-cell-derived-vesicular-proteins-revealed-significant-tumor-specificity-and-predictive-potential-of-extracellular-vesicles-for-cell-invasion-and-proliferation-a-meta-analysis.mdx
new file mode 100644
index 0000000..662ce97
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/240-machine-learning-based-analysis-of-cancer-cell-derived-vesicular-proteins-revealed-significant-tumor-specificity-and-predictive-potential-of-extracellular-vesicles-for-cell-invasion-and-proliferation-a-meta-analysis.mdx
@@ -0,0 +1,10 @@
+---
+title: "Machine learning-based analysis of cancer cell-derived vesicular proteins revealed significant tumor-specificity and predictive potential of extracellular vesicles for cell invasion and proliferation – A meta-analysis"
+category: "content"
+date: "2024-01-12T12:00:24-05:00"
+tags: ["content", "reactome-research-spotlight", "240-machine-learning-based-analysis-of-cancer-cell-derived-vesicular-proteins-revealed-significant-tumor-specificity-and-predictive-potential-of-extracellular-vesicles-for-cell-invasion-and-proliferation-a-meta-analysis"]
+---
+
+## Machine learning-based analysis of cancer cell-derived vesicular proteins revealed significant tumor-specificity and predictive potential of extracellular vesicles for cell invasion and proliferation – A meta-analysis
+
+In the November 2023 issue of Cell Communication and Signaling, [Bukva et al. (2023)]() analyzed the proteomes of tumor-produced extracellular vesicles and identified sets of proteins that could discriminate tumor types, invasiveness, and proliferative capacity. In this analysis, 172 most predictive proteins were identified and used to classify nine tumor types with 91.67% efficiency. Reactome Pathway enrichment analysis of these proteins showed that each tumor type had perturbations in a distinct set of pathways. The proteins could be organized and used to discriminate the invasiveness and proliferative capacity of the tumors. High expression of proteins positively associated with invasiveness and proliferation correlated with reduced patient survival times.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/242-discovering-the-anti-cancer-phytochemical-rutin-against-breast-cancer-through-the-methodical-platform-based-on-traditional-medicinal-knowledge.mdx b/projects/website-angular/content/content/reactome-research-spotlight/242-discovering-the-anti-cancer-phytochemical-rutin-against-breast-cancer-through-the-methodical-platform-based-on-traditional-medicinal-knowledge.mdx
new file mode 100644
index 0000000..1e54163
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/242-discovering-the-anti-cancer-phytochemical-rutin-against-breast-cancer-through-the-methodical-platform-based-on-traditional-medicinal-knowledge.mdx
@@ -0,0 +1,10 @@
+---
+title: "Discovering the anti-cancer phytochemical rutin against breast cancer through the methodical platform based on traditional medicinal knowledge"
+category: "content"
+date: "2024-02-09T13:21:53-05:00"
+tags: ["content", "reactome-research-spotlight", "242-discovering-the-anti-cancer-phytochemical-rutin-against-breast-cancer-through-the-methodical-platform-based-on-traditional-medicinal-knowledge"]
+---
+
+## Discovering the anti-cancer phytochemical rutin against breast cancer through the methodical platform based on traditional medicinal knowledge
+
+In the July 2023 issue of BMB Reports, [Lee et al. (2023)]() employed the Reactome pathway database and tools to predict the anti-cancer effects of rutin, a natural phytochemical identified as a lead chemotherapeutic against breast cancer by text mining Korean traditional medicinal compendia from 1596 CE and 1613 CE. Genes that may be associated with rutin's effects were analyzed for pathway enrichment and functional interactions by the Reactome Functional Interaction (FI) plugin app of Cytoscape. Focal adhesion and [Apoptosis]() were among the pathways predicted to be affected by rutin and these effects were confirmed by treatment of breast cancer cells with rutin in vitro and in xenografts in mice.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/244-identification-of-potential-biological-processes-and-key-genes-in-diabetes-related-stroke-through-weighted-gene-co-expression-network-analysis.mdx b/projects/website-angular/content/content/reactome-research-spotlight/244-identification-of-potential-biological-processes-and-key-genes-in-diabetes-related-stroke-through-weighted-gene-co-expression-network-analysis.mdx
new file mode 100644
index 0000000..b3ba775
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/244-identification-of-potential-biological-processes-and-key-genes-in-diabetes-related-stroke-through-weighted-gene-co-expression-network-analysis.mdx
@@ -0,0 +1,10 @@
+---
+title: "Identification of potential biological processes and key genes in diabetes-related stroke through weighted gene co-expression network analysis"
+category: "content"
+date: "2024-03-13T03:23:12-04:00"
+tags: ["content", "reactome-research-spotlight", "244-identification-of-potential-biological-processes-and-key-genes-in-diabetes-related-stroke-through-weighted-gene-co-expression-network-analysis"]
+---
+
+## Identification of potential biological processes and key genes in diabetes-related stroke through weighted gene co-expression network analysis
+
+Using WGCNA, GO and KEGG data analysis tools,[ He Y et al. in the January 2024 issue of BMC Medical Genomics](), established a connection among the genes and pathways associated with type 2 diabetes (T2D) and ischemic stroke (IS) and identified GRN (granulin precursor) as the hub gene in T2D-related stroke. The functional enrichment analysis using Reactome analysis tool for GRN identified [Neutrophil degranulation](), [Toll-like Receptor Cascades](), [DDX58/IFIH1-mediated induction of interferon-alpha/ beta](), and[ NLR signaling pathways]() as shared biological processes in T2D and IS.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/248-nickel-induced-transcriptional-memory-in-lung-epithelial-cells-promotes-interferon-signaling-upon-nicotine-exposure.mdx b/projects/website-angular/content/content/reactome-research-spotlight/248-nickel-induced-transcriptional-memory-in-lung-epithelial-cells-promotes-interferon-signaling-upon-nicotine-exposure.mdx
new file mode 100644
index 0000000..8cf5b94
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/248-nickel-induced-transcriptional-memory-in-lung-epithelial-cells-promotes-interferon-signaling-upon-nicotine-exposure.mdx
@@ -0,0 +1,10 @@
+---
+title: "Nickel-induced transcriptional memory in lung epithelial cells promotes interferon signaling upon nicotine exposure"
+category: "content"
+date: "2024-04-09T00:51:43-04:00"
+tags: ["content", "reactome-research-spotlight", "248-nickel-induced-transcriptional-memory-in-lung-epithelial-cells-promotes-interferon-signaling-upon-nicotine-exposure"]
+---
+
+## Nickel-induced transcriptional memory in lung epithelial cells promotes interferon signaling upon nicotine exposure
+
+In the December 2023 issue of[ ]()Toxicology and Applied Pharmacology, [Zhang et al](). used the R package, ReactomePA [(Yu and He, 2016)](), to identify enriched pathways responding to nickel-induced transcriptional memory changes in response to a second respiratory toxicant, nicotine. Nicotine exposure upregulated a specific subset of genes in the cells previously exposed to nickel, identifying a robust activation of [Interferon (IFN) signaling](), a driver of inflammation associated with many chronic lung diseases.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/249-ibpgnet-lung-adenocarcinoma-recurrence-prediction-based-on-neural-network-interpretability.mdx b/projects/website-angular/content/content/reactome-research-spotlight/249-ibpgnet-lung-adenocarcinoma-recurrence-prediction-based-on-neural-network-interpretability.mdx
new file mode 100644
index 0000000..cf54ba2
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/249-ibpgnet-lung-adenocarcinoma-recurrence-prediction-based-on-neural-network-interpretability.mdx
@@ -0,0 +1,10 @@
+---
+title: "IBPGNET: lung adenocarcinoma recurrence prediction based on neural network interpretability"
+category: "content"
+date: "2024-05-10T12:15:21-04:00"
+tags: ["content", "reactome-research-spotlight", "249-ibpgnet-lung-adenocarcinoma-recurrence-prediction-based-on-neural-network-interpretability"]
+---
+
+## IBPGNET: lung adenocarcinoma recurrence prediction based on neural network interpretability
+
+In the May 2024 issue of Briefings in Bioinformatics, [Xu et al.]() develop an Interpretable Biological Pathway Graph Neural Network (IBPGNET) framework based on Reactome pathway hierarchy to predict regulatory mechanisms that lead to lung adenocarcinoma recurrences. IBPGNET identified two genes of interest and performed in vitro knockdown models for drug sensitivity experimental validation. This study offers an approach for exploring molecular mechanisms underlying recurrence using Reactome’s hierarchical pathway structure.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/252-acquired-resistance-to-immunotherapy-and-chemoradiation-in-myc-amplified-head-and-neck-cancer.mdx b/projects/website-angular/content/content/reactome-research-spotlight/252-acquired-resistance-to-immunotherapy-and-chemoradiation-in-myc-amplified-head-and-neck-cancer.mdx
new file mode 100644
index 0000000..4f07a96
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/252-acquired-resistance-to-immunotherapy-and-chemoradiation-in-myc-amplified-head-and-neck-cancer.mdx
@@ -0,0 +1,10 @@
+---
+title: Acquired resistance to immunotherapy and chemoradiation in MYC amplified head and neck cancer
+category: "content"
+date: "2024-06-18T10:04:25-04:00"
+tags: ["content", "reactome-research-spotlight", "252-acquired-resistance-to-immunotherapy-and-chemoradiation-in-myc-amplified-head-and-neck-cancer"]
+---
+
+## Acquired resistance to immunotherapy and chemoradiation in MYC amplified head and neck cancer
+
+In the May 2024 issue of NPJ Precision Oncology, [Cyberski et al.]() used Reactome’s hierarchically arranged pathways with their in silico Pathway Activation Network Decomposition Analysis (iPANDA) algorithm to identify upregulation of networks associated with [cell cycle ]()progression, [signal transduction](), and [metabolism]() and down-regulation of [immune cellular process ]()and [apoptosis]() in MYC-amplified cases of recurrent/metastatic head and neck squamous cell carcinoma.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/254-drug-target-prediction-through-deep-learning-functional-representation-of-gene-signatures.mdx b/projects/website-angular/content/content/reactome-research-spotlight/254-drug-target-prediction-through-deep-learning-functional-representation-of-gene-signatures.mdx
new file mode 100644
index 0000000..a098b8e
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/254-drug-target-prediction-through-deep-learning-functional-representation-of-gene-signatures.mdx
@@ -0,0 +1,10 @@
+---
+title: Drug target prediction through deep learning functional representation of gene signatures
+category: "content"
+date: "2024-07-04T16:52:17-04:00"
+tags: ["content", "reactome-research-spotlight", "254-drug-target-prediction-through-deep-learning-functional-representation-of-gene-signatures"]
+---
+
+## Drug target prediction through deep learning functional representation of gene signatures
+
+In their May 2024 Nature Communications paper, [Chen et al]() used data simulated based on Reactome pathways to validate their Functional Representation of Gene Signatures (FRoGS) algorithm, a deep learning-based approach that was designed to improve the accuracy of drug target predictions by addressing limitations of gene identity-based pathway analysis.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/255-the-landscape-of-cancer-rewired-gpcr-signaling-axes.mdx b/projects/website-angular/content/content/reactome-research-spotlight/255-the-landscape-of-cancer-rewired-gpcr-signaling-axes.mdx
new file mode 100644
index 0000000..238d36d
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/255-the-landscape-of-cancer-rewired-gpcr-signaling-axes.mdx
@@ -0,0 +1,10 @@
+---
+title: "The landscape of cancer-rewired GPCR signaling axes"
+category: "content"
+date: "2024-07-23T13:34:24-04:00"
+tags: ["content", "reactome-research-spotlight", "255-the-landscape-of-cancer-rewired-gpcr-signaling-axes"]
+---
+
+## The landscape of cancer-rewired GPCR signaling axes
+
+In their May 2024 paper in Cell Genomics,[ Arora et al.]() used a framework of Reactome signaling and metabolism pathways to integrate RHEA metabolic reactions and IUPhAR catalogs of G Protein-Coupled Receptors (GPCRs) and their ligands to define axes that combine[ signaling cascades]() and ligand[ metabolic processes](). Altered expression of the sets of proteins that make up these axes correlate with patient survival cataloged in The Cancer Genome Atlas (TCGA) for many tumor types and suggest novel druggable targets.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/256-engineering-toxoplasma-gondii-secretion-systems-for-intracellular-delivery-of-multiple-large-therapeutic-proteins-to-neurons.mdx b/projects/website-angular/content/content/reactome-research-spotlight/256-engineering-toxoplasma-gondii-secretion-systems-for-intracellular-delivery-of-multiple-large-therapeutic-proteins-to-neurons.mdx
new file mode 100644
index 0000000..b58294a
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/256-engineering-toxoplasma-gondii-secretion-systems-for-intracellular-delivery-of-multiple-large-therapeutic-proteins-to-neurons.mdx
@@ -0,0 +1,10 @@
+---
+title: Engineering Toxoplasma gondii secretion systems for intracellular delivery of multiple large therapeutic proteins to neurons
+category: "content"
+date: "2024-08-29T18:48:21-04:00"
+tags: ["content", "reactome-research-spotlight", "256-engineering-toxoplasma-gondii-secretion-systems-for-intracellular-delivery-of-multiple-large-therapeutic-proteins-to-neurons"]
+---
+
+## Engineering Toxoplasma gondii secretion systems for intracellular delivery of multiple large therapeutic proteins to neurons
+
+In the August 2024 issue of Nature Microbiology, [Bracha et al](). use Reactome expression analysis to confirm that they successfully delivered multiple large (>100 kDa) therapeutic proteins across the blood-brain barrier into target neurons in mice using engineered _Toxoplasma gondii_ secretion systems.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/262-chemical-coverage-of-human-biological-pathways.mdx b/projects/website-angular/content/content/reactome-research-spotlight/262-chemical-coverage-of-human-biological-pathways.mdx
new file mode 100644
index 0000000..65f7f35
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/262-chemical-coverage-of-human-biological-pathways.mdx
@@ -0,0 +1,10 @@
+---
+title: Chemical coverage of human biological pathways
+category: "content"
+date: "2024-10-03T11:25:44-04:00"
+tags: ["content", "reactome-research-spotlight", "262-chemical-coverage-of-human-biological-pathways"]
+---
+
+## Chemical coverage of human biological pathways
+
+In the feature article of the October 2024 issue of [Drug Discovery Today]() titled [“Chemical coverage of human biological pathways]()”, Kwak et al. describe the [Target 2035]() initiative, whose mission is to discover chemical tools for all human proteins by 2035. The authors use Reactome as the reference standard to determine the chemical coverage of human biological pathways and to outline the advantages of adopting the pathway-based rather than the proteome-based approach in guiding Target 2035 efforts.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/263-pathintegrate-multivariate-modelling-approaches-for-pathway-based-multi-omics-data-integration.mdx b/projects/website-angular/content/content/reactome-research-spotlight/263-pathintegrate-multivariate-modelling-approaches-for-pathway-based-multi-omics-data-integration.mdx
new file mode 100644
index 0000000..becaf7d
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/263-pathintegrate-multivariate-modelling-approaches-for-pathway-based-multi-omics-data-integration.mdx
@@ -0,0 +1,10 @@
+---
+title: "PathIntegrate: Multivariate modelling approaches for pathway-based multi-omics data integration"
+category: "content"
+date: "2024-10-22T14:59:45-04:00"
+tags: ["content", "reactome-research-spotlight", "263-pathintegrate-multivariate-modelling-approaches-for-pathway-based-multi-omics-data-integration"]
+---
+
+## PathIntegrate: Multivariate modelling approaches for pathway-based multi-omics data integration
+
+In [PLOS Computational Biology](), [Wieder et al. (2024) ]()employ the Reactome database and PathIntegrate, a pathway-based multi-omics integration tool based on single-sample pathway analysis and machine learning, to translate multi-omics datasets from molecular abundance measurements to pathway activity scores, enabling integration of disparate types of omics data according to a common scale. PathIntegrate provides higher sensitivity at low signal levels and efficiently identifies perturbed pathways from multi-omics datasets in COVID-19 and chronic obstructive pulmonary disease (COPD) examples, providing a readily interpretable predictive model.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/264-rna-editing-regulates-host-immune-response-and-t-cell-homeostasis-in-sars-cov-2-infection.mdx b/projects/website-angular/content/content/reactome-research-spotlight/264-rna-editing-regulates-host-immune-response-and-t-cell-homeostasis-in-sars-cov-2-infection.mdx
new file mode 100644
index 0000000..f473970
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/264-rna-editing-regulates-host-immune-response-and-t-cell-homeostasis-in-sars-cov-2-infection.mdx
@@ -0,0 +1,10 @@
+---
+title: "RNA editing regulates host immune response and T cell homeostasis in SARS-CoV-2 infection"
+category: "content"
+date: "2024-11-21T22:52:31-05:00"
+tags: ["content", "reactome-research-spotlight", "264-rna-editing-regulates-host-immune-response-and-t-cell-homeostasis-in-sars-cov-2-infection"]
+---
+
+## RNA editing regulates host immune response and T cell homeostasis in SARS-CoV-2 infection
+
+In the August 2024 issue of PLoS One, [Huang et al.]() used the Reactome database to analyze the pattern of RNA editing in cells in response to infection by SARS-CoV-2 and found that editing was highest in transcripts of genes related to immune response andcytokine production. Single cell transcriptomics showed that the Reactome [Interferon signaling]() pathway is enriched in plasmacytoid B cells, B cells, and T cell subtypes during SARS-CoV-2 infection.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/266-a-living-organoid-biobank-of-patients-with-crohn-s-disease-reveals-molecular-subtypes-for-personalized-therapeutics.mdx b/projects/website-angular/content/content/reactome-research-spotlight/266-a-living-organoid-biobank-of-patients-with-crohn-s-disease-reveals-molecular-subtypes-for-personalized-therapeutics.mdx
new file mode 100644
index 0000000..0996218
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/266-a-living-organoid-biobank-of-patients-with-crohn-s-disease-reveals-molecular-subtypes-for-personalized-therapeutics.mdx
@@ -0,0 +1,10 @@
+---
+title: A living organoid biobank of patients with Crohn’s disease reveals molecular subtypes for personalized therapeutics
+category: "content"
+date: "2024-12-30T21:19:35-05:00"
+tags: ["content", "reactome-research-spotlight", "266-a-living-organoid-biobank-of-patients-with-crohn-s-disease-reveals-molecular-subtypes-for-personalized-therapeutics"]
+---
+
+## A living organoid biobank of patients with Crohn’s disease reveals molecular subtypes for personalized therapeutics
+
+In the October 2024 issue of Cell Reports Medicine, [Tindle et al.]() identified two Crohn’s disease (CD) molecular subtypes - immune-deficient infectious CD (IDICD) and stress and senescence-induced fibrostenotic CD (S2FCD) - through multi-omics and functional analyses of patient-derived organoids. Reactome pathway enrichment analysis revealed subtype-specific dysregulations. In IDICD, the [Nuclear receptor transcription factor]() pathway, [Butyrophilin family interactions](), and [Intestinal infectious disease]() events were upregulated while [Cytokine signaling in immune system]() events were downregulated. In S2FCD, [Oncogene- ]()and [Oxidative stress-induced senescence]() pathways were upregulated and Signaling by TGF-beta receptor complex events were downregulated suggesting distinct subtype-specific therapeutic strategies.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/267-bpp-a-platform-for-automatic-biochemical-pathway-prediction.mdx b/projects/website-angular/content/content/reactome-research-spotlight/267-bpp-a-platform-for-automatic-biochemical-pathway-prediction.mdx
new file mode 100644
index 0000000..b70fe9f
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/267-bpp-a-platform-for-automatic-biochemical-pathway-prediction.mdx
@@ -0,0 +1,10 @@
+---
+title: "BPP: a platform for automatic biochemical pathway prediction"
+category: "content"
+date: "2025-01-28T20:35:50-05:00"
+tags: ["content", "reactome-research-spotlight", "267-bpp-a-platform-for-automatic-biochemical-pathway-prediction"]
+---
+
+## BPP: a platform for automatic biochemical pathway prediction
+
+In the July 2024 issue of Briefings in Bioinformatics, [Yi et al.]() report on The Biochemical Pathway Prediction (BPP) framework, a predictive analytical tool that utilizes various graph representation learning models to predict attributes and links in biochemical pathways. BPP provides two pieces of information: link prediction, which identifies potential connections between entities and reactions, and attribute prediction, which predicts missing attributes of nodes. The BPP framework was used to evaluate datasets derived from Reactome pathway data (version 75 to version 85), specifically identifying a key receptor, [glycosylated-ACE2](), instrumental in the [SARS-CoV-2 viral invasion]() process.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/269-co-methylation-networks-associated-with-cognition-and-structural-brain-development-during-adolescence.mdx b/projects/website-angular/content/content/reactome-research-spotlight/269-co-methylation-networks-associated-with-cognition-and-structural-brain-development-during-adolescence.mdx
new file mode 100644
index 0000000..1b287f9
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/269-co-methylation-networks-associated-with-cognition-and-structural-brain-development-during-adolescence.mdx
@@ -0,0 +1,10 @@
+---
+title: "Co-methylation networks associated with cognition and structural brain development during adolescence"
+category: "content"
+date: "2025-02-25T23:02:54-05:00"
+tags: ["content", "reactome-research-spotlight", "269-co-methylation-networks-associated-with-cognition-and-structural-brain-development-during-adolescence"]
+---
+
+## Co-methylation networks associated with cognition and structural brain development during adolescence
+
+In the January 2025 issue of Frontiers in Genetics, [Jensen et al](). explored the relationship between DNA methylation patterns and adolescent brain development. By analyzing a cohort of adolescents aged 9 to 14, they identified co-methylation networks linked to cognitive improvements and structural brain changes. Pathway analysis using Reactome revealed that these networks are enriched in neuronal-related pathways, suggesting that epigenetic modifications play a significant role in the maturation of the adolescent brain.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/271-genetically-supported-targets-and-drug-repurposing-for-brain-aging-a-systematic-study-in-the-uk-biobank.mdx b/projects/website-angular/content/content/reactome-research-spotlight/271-genetically-supported-targets-and-drug-repurposing-for-brain-aging-a-systematic-study-in-the-uk-biobank.mdx
new file mode 100644
index 0000000..af666da
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/271-genetically-supported-targets-and-drug-repurposing-for-brain-aging-a-systematic-study-in-the-uk-biobank.mdx
@@ -0,0 +1,10 @@
+---
+title: "Genetically supported targets and drug repurposing for brain aging: A systematic study in the UK Biobank"
+category: "content"
+date: "2025-03-31T02:58:31-04:00"
+tags: ["content", "reactome-research-spotlight", "271-genetically-supported-targets-and-drug-repurposing-for-brain-aging-a-systematic-study-in-the-uk-biobank"]
+---
+
+## Genetically supported targets and drug repurposing for brain aging: A systematic study in the UK Biobank
+
+In the March 2025 issue of Science Advances [Yi et al. ]()reported the development of a brain age estimation model using large-scale genetic and imaging data. Brain age gap (BAG) is a digital phenotype that may reflect associations with various brain disorders. This study aimed to identify potential drug targets causally associated with BAG. A total of 64 genes were identified within five Reactome pathways: [programmed cell death](), [platelet signaling and aggregation]() , [extracellular matrix organization](), [cell surface interactions at the vascular wall](), and [apoptosis]() Of these, seven genes were prioritized as targets due to strong genetic evidence for BAG.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/272-reactome-strengthens-accuracy-by-monitoring-for-retracted-publications.mdx b/projects/website-angular/content/content/reactome-research-spotlight/272-reactome-strengthens-accuracy-by-monitoring-for-retracted-publications.mdx
new file mode 100644
index 0000000..7e54d6f
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/272-reactome-strengthens-accuracy-by-monitoring-for-retracted-publications.mdx
@@ -0,0 +1,10 @@
+---
+title: Reactome Strengthens Accuracy by Monitoring for Retracted Publications
+category: "content"
+date: "2025-04-30T23:03:07-04:00"
+tags: ["content", "reactome-research-spotlight", "272-reactome-strengthens-accuracy-by-monitoring-for-retracted-publications"]
+---
+
+## Reactome Strengthens Accuracy by Monitoring for Retracted Publications
+
+Reactome is committed to maintaining the highest standards of scientific accuracy. To help prevent the circulation of retracted research, we conduct regular, systematic reviews of all literature-backed assertions in our database. If a publication listed in the Retraction Watch database has been used as supporting evidence for any Reactome annotation, we re-evaluate the associated data. Annotations linked to retracted papers are either updated with new, valid references or removed entirely if no suitable replacements can be found. Each removal is documented along with the reason for the change. To date, we have reviewed over 40,000 curator-selected references and identified just 70 retracted articles. All affected annotations have been reviewed and revised accordingly. This ongoing process ensures the continued integrity and reliability of Reactome’s content.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/274-interpreting-biologically-informed-neural-networks-for-enhanced-proteomic-biomarker-discovery-and-pathway-analysis.mdx b/projects/website-angular/content/content/reactome-research-spotlight/274-interpreting-biologically-informed-neural-networks-for-enhanced-proteomic-biomarker-discovery-and-pathway-analysis.mdx
new file mode 100644
index 0000000..cf6ad59
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/274-interpreting-biologically-informed-neural-networks-for-enhanced-proteomic-biomarker-discovery-and-pathway-analysis.mdx
@@ -0,0 +1,10 @@
+---
+title: Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
+category: "content"
+date: "2025-06-01T23:18:05-04:00"
+tags: ["content", "reactome-research-spotlight", "274-interpreting-biologically-informed-neural-networks-for-enhanced-proteomic-biomarker-discovery-and-pathway-analysis"]
+---
+
+## Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
+
+[June 1, 2025] The lack of interpretability in deep neural networks is a challenging issue in biomedical applications. In their 2023 Nature Communications study, [“Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis” ]()Hartman et al. used Reactome’s pathway hierarchical tree directly to develop multi-layered, biologically informed neural networks (BINNs) to address this issue and enhance proteomic biomarker discovery and pathway analysis. Reactome provided critical information on biological entity relationships, enabling the creation of BINNs that achieved high predictive accuracy in the identification of disease-relevant biomarkers and pathways in septic acute kidney injury and COVID-19 datasets. These BINNs outperformed traditional methods and provided experimentally testable molecular mechanistic explanations.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/276-rhythm-profiling-using-cofe-reveals-multi-omic-circadian-rhythms-in-human-cancers-in-vivo.mdx b/projects/website-angular/content/content/reactome-research-spotlight/276-rhythm-profiling-using-cofe-reveals-multi-omic-circadian-rhythms-in-human-cancers-in-vivo.mdx
new file mode 100644
index 0000000..a8b8028
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/276-rhythm-profiling-using-cofe-reveals-multi-omic-circadian-rhythms-in-human-cancers-in-vivo.mdx
@@ -0,0 +1,10 @@
+---
+title: "Rhythm profiling using COFE reveals multi-omic circadian rhythms in human cancers in vivo"
+category: "content"
+date: "2025-06-27T11:33:17-04:00"
+tags: ["content", "reactome-research-spotlight", "276-rhythm-profiling-using-cofe-reveals-multi-omic-circadian-rhythms-in-human-cancers-in-vivo"]
+---
+
+## Rhythm profiling using COFE reveals multi-omic circadian rhythms in human cancers in vivo
+
+[July 1, 2025] Gene expression levels in normal and diseased human tissues show circadian variation, but studying this variation directly is difficult. In their May, 2025 PLoS paper, [Ananthasubramaniam and Venkataramanan]() applied unsupervised machine learning to high-throughput omics data from primary human adenocarcinomas to identify circadian expression rhythms in hundreds of genes. Reactome gene set enrichment analysis identified genes with rhythmic expression patterns in multiple tumor types, significantly overrepresented in pathways of [mitochondrial translation](), [respiratory electron transport](), [mitotic cell cycle](), and [adaptive immune system](). The rhythmic expression of gene / protein targets of many FDA-approved and potential anti-cancer drugs in the adenocarcinomas suggests that timing of anti-tumor drug administration may improve efficacy.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/277-identification-and-targeting-of-regulators-of-sars-cov-2-host-interactions-in-the-airway-epithelium.mdx b/projects/website-angular/content/content/reactome-research-spotlight/277-identification-and-targeting-of-regulators-of-sars-cov-2-host-interactions-in-the-airway-epithelium.mdx
new file mode 100644
index 0000000..a994bb2
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/277-identification-and-targeting-of-regulators-of-sars-cov-2-host-interactions-in-the-airway-epithelium.mdx
@@ -0,0 +1,10 @@
+---
+title: "Identification and targeting of regulators of SARS-CoV-2–host interactions in the airway epithelium"
+category: "content"
+date: "2025-07-28T15:14:13-04:00"
+tags: ["content", "reactome-research-spotlight", "277-identification-and-targeting-of-regulators-of-sars-cov-2-host-interactions-in-the-airway-epithelium"]
+---
+
+## Identification and targeting of regulators of SARS-CoV-2–host interactions in the airway epithelium
+
+In the May 2025 issue of Science Advances, [Dirvin et al](). used single-cell transcriptomics and network-based algorithms on primary human airway cells to identify the key master regulator proteins hijacked by SARS-CoV-2, and then performed a large-scale screen to find drugs capable of reversing these effects. They used Reactome pathway analysis to characterize the biological processes, including [membrane trafficking]() and [ infectious disease pathways](), that were modulated by the eleven most promising drug candidates.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/278-learning-and-actioning-general-principles-of-cancer-cell-drug-sensitivity.mdx b/projects/website-angular/content/content/reactome-research-spotlight/278-learning-and-actioning-general-principles-of-cancer-cell-drug-sensitivity.mdx
new file mode 100644
index 0000000..47de561
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/278-learning-and-actioning-general-principles-of-cancer-cell-drug-sensitivity.mdx
@@ -0,0 +1,10 @@
+---
+title: Learning and actioning general principles of cancer cell drug sensitivity
+category: "content"
+date: "2025-08-26T00:28:13-04:00"
+tags: ["content", "reactome-research-spotlight", "278-learning-and-actioning-general-principles-of-cancer-cell-drug-sensitivity"]
+---
+
+## Learning and actioning general principles of cancer cell drug sensitivity
+
+In the February 2025 issue of Nature Communications, [Carli et al](). reported the development of a predictive model of cell line drug sensitivity from RNA-seq data using machine learning approaches. The model leveraged Reactome pathways in combination with large language models (LLMs) to provide a mechanistic foundation. It demonstrated strong performance and was applied to predict patient drug responses, with predictions supported by experimental validation. This work highlights how Reactome provides a robust framework for enhancing the interpretability of machine learning models in precision medicine.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/281-an-immune-competent-lung-on-a-chip-for-modelling-the-human-severe-influenza-infection-response.mdx b/projects/website-angular/content/content/reactome-research-spotlight/281-an-immune-competent-lung-on-a-chip-for-modelling-the-human-severe-influenza-infection-response.mdx
new file mode 100644
index 0000000..ba6a194
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/281-an-immune-competent-lung-on-a-chip-for-modelling-the-human-severe-influenza-infection-response.mdx
@@ -0,0 +1,10 @@
+---
+title: "An immune-competent lung-on-a-chip for modelling the human severe influenza infection response"
+category: "content"
+date: "2025-09-30T18:37:19-04:00"
+tags: ["content", "reactome-research-spotlight", "281-an-immune-competent-lung-on-a-chip-for-modelling-the-human-severe-influenza-infection-response"]
+---
+
+## An immune-competent lung-on-a-chip for modelling the human severe influenza infection response
+
+In their September 2025 [Nature Biomedical Engineering]() paper, [An immune-competent lung-on-a-chip for modelling the human severe influenza infection response](), Ringquist et al. show the importance of including tissue-resident and circulating immune cells, as well as stromal cells, in lung organoid chips to obtain a more realistic human in vitro model system for studying viral respiratory infections. Human-centric Reactome pathway enrichment analysis employed in this study shows that genes expressed in immune and stromal cells, mediating [immune response]() and [extracellular matrix remodeling](), respectively, are among the top 10% upregulated in influenza H1N1-infected lung organoids, amid a global transcriptional shutdown, showing the value this ex vivo model for studying lung infectious disease.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/283-reprogramming-neuroblastoma-by-diet-enhanced-polyamine-depletion.mdx b/projects/website-angular/content/content/reactome-research-spotlight/283-reprogramming-neuroblastoma-by-diet-enhanced-polyamine-depletion.mdx
new file mode 100644
index 0000000..4b24211
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/283-reprogramming-neuroblastoma-by-diet-enhanced-polyamine-depletion.mdx
@@ -0,0 +1,14 @@
+---
+title: "Reprogramming neuroblastoma by diet-enhanced polyamine depletion"
+category: "content"
+date: "2025-10-28T15:00:42-04:00"
+tags: ["content", "reactome-research-spotlight", "283-reprogramming-neuroblastoma-by-diet-enhanced-polyamine-depletion"]
+---
+
+## Reprogramming neuroblastoma by diet-enhanced polyamine depletion
+
+Neuroblastomas are driven by MYCN hyperactivity, accumulate abnormally high amounts of arginine, proline, and ornithine, and produce abnormally high amounts of polyamines due to MYCN upregulation of ornithine decarboxylase (ODC), the rate limiting enzyme of polyamine synthesis. Difluoromethylornithine (DFMO), an inhibitor of ODC, has therapeutic effect against neuroblastoma. Therefore, Cherkaoui et al (2025) in their October, 2025 Nature article ["Reprogramming neuroblastoma by diet-enhanced polyamine depletion"]() tested whether a diet free of proline and arginine, precursors of ornithine in neuroblastoma, would improve survival further. Although the proline-arginine-free diet alone had no effect on survival, in combination with DFMO it approximately doubled survival in experimental models.
+
+Cherkaoui et al. then investigated the mechanism by which DFMO combined with depletion of proline and arginine inhibited growth of neuroblastomas. The treated tumors exhibited a ten-fold reduction in polyamine content relative to untreated tumors. Because spermidine and other polyamines can enhance translation, the translation efficiency of genes was measured by large scale analysis of RNA (RNA-seq), ribosome-bound RNA (Ribo-seq), and proteins. The surprising finding was that in conditions of low polyamines, ribosomes stalled more frequently at codons with adenosine at the third position. This may be due to the combination of a requirement for highly modified tRNAs to translate these codons, and lower hypusination of the eIF5A translation factor.
+
+Why would stalling at a particular set of codons produce a specific anti-cancer effect? Cherkaoui et al. employed the [Reactome]() database to examine the codon usage in the genes encoding components of biological pathways. Surprisingly, the genes of the [cell cycle]() pathway had a significantly higher proportion of codons ending in adenosine than did the genes of the [neuronal system]() pathway, accounting for the selective effect of DFMO and the arginine-proline-free diet on neuroblastoma cell proliferation. Also surprisingly, all pathways varied significantly in codon usage, suggesting possible new methods of therapeutically regulating them.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/285-anti-progestin-therapy-targets-hallmarks-of-breast-cancer-risk.mdx b/projects/website-angular/content/content/reactome-research-spotlight/285-anti-progestin-therapy-targets-hallmarks-of-breast-cancer-risk.mdx
new file mode 100644
index 0000000..f1b348a
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/285-anti-progestin-therapy-targets-hallmarks-of-breast-cancer-risk.mdx
@@ -0,0 +1,10 @@
+---
+title: "Anti-progestin therapy targets hallmarks of breast cancer risk"
+category: "content"
+date: "2025-12-05T14:03:53-05:00"
+tags: ["content", "reactome-research-spotlight", "285-anti-progestin-therapy-targets-hallmarks-of-breast-cancer-risk"]
+---
+
+## Anti-progestin therapy targets hallmarks of breast cancer risk
+
+Progesterone, a hormone cyclically produced during menstrual cycles and used in hormone replacement therapy (HRT) after menopause, can promote cell proliferation primarily through a paracrine signaling mechanism, where progesterone receptor (PR)-positive 'luminal mature' cells secrete signaling factors that act on neighboring PR-negative 'luminal progenitor' cells. This mechanism plays a crucial role in normal mammary gland development and has been implicated in breast cancer pathogenesis. Anti-progestin therapy has long been regarded as a potential strategy for breast cancer prevention. Simões et al. (2025) in their November 2025 Nature article, [Anti-progestin therapy targets hallmarks of breast cancer risk](), report findings from the single-arm phase II trial (BC-APPS1; [NCT02408770]()), showing the effects of ulipristal acetate (UA), a progesterone receptor antagonist, on breast tissue from women at higher risk of breast cancer. The study combined contrast-enhanced magnetic resonance imaging (MRI) data with multi-OMICs and imaging analyses of paired vacuum-assisted breast biopsies collected before and after 12 weeks of daily ulipristal acetate (UA) treatment. UA treatment reduced epithelial proliferation and depleted luminal progenitor cells, impairing their colony-forming capacity. Multi-omics analysis, including single-cell transcriptomics and laser-capture microdissection (LCM) proteomics, identified the extracellular matrix as the primary target of UA. Pathway enrichment using Reactome data revealed that [extracellular matrix (ECM) ]()processes were downregulated in fibroblasts and basal-myoepithelial cells while luminal hormone-sensing cells (LHS) showed downregulation of “[RNA-processing]()” components and upregulation of matrix metalloproteinases associated with “[collagen degradation]()”. Among the downregulated ECM genes, collagen VI chains (COL6A2, COL6A3) were the most significantly reduced. CellChat and NicheNet analyses showed that UA reduced fibroblast and basal–myoepithelial collagen-signaling outputs and linked LHS-derived ligands (WNT5A, RARRES1, APOD) to the regulation of key collagen genes (COL6A3, COL1A2) in fibroblast subclusters, highlighting progesterone-dependent paracrine control of the breast matrisome. Imaging analyses confirmed decreased abundance of collagen I, collagen VI (COL6A3), and fibronectin (FN1) which correlated with reduced tissue stiffness and MRI-assessed fibroglandular volume following UA treatment. Primary human breast organoids grown in stiff hydrogels showed increased expression of PR target gene TNFSF11 and luminal progenitor markers SOX9 and KIT, along with increased mammosphere formation - effects fully suppressed by UA treatment. Collectively, these findings reveal how ulipristal acetate modulates both epithelial and stromal biology through coordinated suppression of luminal progenitors and increased ECM remodeling to reduce breast cancer risk in premenopausal women.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/287-inhibition-of-type-i-interferon-signaling-is-a-conserved-function-of-gamma-herpesvirus-encoded-micrornas.mdx b/projects/website-angular/content/content/reactome-research-spotlight/287-inhibition-of-type-i-interferon-signaling-is-a-conserved-function-of-gamma-herpesvirus-encoded-micrornas.mdx
new file mode 100644
index 0000000..135ebc8
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/287-inhibition-of-type-i-interferon-signaling-is-a-conserved-function-of-gamma-herpesvirus-encoded-micrornas.mdx
@@ -0,0 +1,10 @@
+---
+title: "Inhibition of type I interferon signaling is a conserved function of gamma-herpesvirus-encoded microRNAs"
+category: "content"
+date: "2026-01-10T01:30:20-05:00"
+tags: ["content", "reactome-research-spotlight", "287-inhibition-of-type-i-interferon-signaling-is-a-conserved-function-of-gamma-herpesvirus-encoded-micrornas"]
+---
+
+## Inhibition of type I interferon signaling is a conserved function of gamma-herpesvirus-encoded microRNAs
+
+Type I interferon (IFN) signaling is one of the body’s earliest and most important defenses against viral infection, rapidly activating hundreds of antiviral genes. In the December 2025 Journal of Virology article, “[Inhibition of type I interferon signaling is a conserved function of gamma-herpesvirus-encoded microRNAs]()”, Fachko and colleagues demonstrate that gamma-herpesviruses, closely related to Epstein–Barr virus and Kaposi’s sarcoma–associated herpesvirus, encode microRNAs that consistently inhibit this pathway. By combining reporter assays, primary cell infections, and genetically engineered viruses lacking specific microRNA clusters, the authors demonstrate that viral microRNAs reduce interferon-stimulated gene expression during early infection and make latently infected cells less responsive to interferon. Importantly, they identify direct targeting of interferon receptors (IFNAR1 and IFNAR2) and central JAK/STAT pathway components (including JAK1, IRF9, and STAT-associated transcriptional regulators), revealing a multi-level strategy by which these viruses suppress antiviral immunity.The researchers use Reactome as they reanalyze Argonaute PAR-CLIP datasets, identifying “[Interferon signaling]()” and “[Interferon alpha/beta signaling]()” pathway host genes bound and regulated by viral microRNAs within the immune response pathway. This pathway-based analysis allowed them to move beyond individual gene hits and show that viral microRNAs converge on multiple nodes of the same antiviral signaling network. The use of high-quality Reactome pathway data strengthened the mechanistic conclusions of the study, highlighting how pathway-level analysis can reveal conserved immune evasion strategies employed by herpesviruses across species.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/289-markerpredict-predicting-clinically-relevant-predictive-biomarkers-with-machine-learning.mdx b/projects/website-angular/content/content/reactome-research-spotlight/289-markerpredict-predicting-clinically-relevant-predictive-biomarkers-with-machine-learning.mdx
new file mode 100644
index 0000000..bad7457
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/289-markerpredict-predicting-clinically-relevant-predictive-biomarkers-with-machine-learning.mdx
@@ -0,0 +1,16 @@
+---
+title: "MarkerPredict: predicting clinically relevant predictive biomarkers with machine learning"
+category: "content"
+date: "2026-02-14T12:19:16-05:00"
+tags: ["content", "reactome-research-spotlight", "289-markerpredict-predicting-clinically-relevant-predictive-biomarkers-with-machine-learning"]
+---
+
+## MarkerPredict: predicting clinically relevant predictive biomarkers with machine learning
+
+In their November 2025 NPJ Systems Biology and Applications paper “[MarkerPredict: predicting clinically relevant predictive biomarkers with machine learning]()”, Veres et al. constructed signaling networks using multiple curated interaction resources, with Reactome Functional Interaction (ReactomeFI) serving as a primary network due to its pathway-informed structure. Within these networks, they identified fully connected three-node motifs (“triangles”) containing known oncologic drug targets and intrinsically disordered proteins (IDPs). ReactomeFI showed the strongest enrichment of IDP–target triangles (enrichment ratio 11.91) relative to alternative networks, including CSN and SIGNOR, supporting its suitability for motif-based biomarker discovery.
+
+Features derived from network topology (e.g., motif participation and connectivity patterns) and protein disorder characteristics were used to train Random Forest and XGBoost classifiers. These models were evaluated for their ability to distinguish protein pairs associated with drug sensitivity from non-informative pairs. Model outputs were integrated into a Biomarker Probability Score (BPS), which ranks proteins by their likelihood of serving as predictive biomarkers.
+
+Applying MarkerPredict across targeted cancer therapies yielded 2,084 candidate predictive biomarkers. Among these, proteins such as LCK and ERK1 were highlighted as high-confidence candidates due to their network positioning and structural features, suggesting relevance for further experimental and clinical validation. The results indicate that proteins embedded in specific signaling motifs and exhibiting intrinsic disorder are more likely to function as effective predictors of therapeutic response.
+
+Overall, the study demonstrates that combining ReactomeFI-derived network topology with protein disorder information and supervised machine learning provides a scalable strategy for predictive biomarker discovery. MarkerPredict offers a complementary approach to existing biomarker identification methods and supports more informed selection of targeted therapies in oncology.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/blogpost-1.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-1.mdx
new file mode 100644
index 0000000..fa82eea
--- /dev/null
+++ b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-1.mdx
@@ -0,0 +1,16 @@
+---
+title: "MarkerPredict: predicting clinically relevant predictive biomarkers with machine learning"
+category: "content"
+date: "2026-02-14T12:19:16-05:00"
+tags: ["content", "reactome-research-spotlight"]
+---
+
+## [ MarkerPredict: predicting clinically relevant predictive biomarkers with machine learning ]()
+
+In their November 2025 NPJ Systems Biology and Applications paper “[MarkerPredict: predicting clinically relevant predictive biomarkers with machine learning]()”, Veres et al. constructed signaling networks using multiple curated interaction resources, with Reactome Functional Interaction (ReactomeFI) serving as a primary network due to its pathway-informed structure. Within these networks, they identified fully connected three-node motifs (“triangles”) containing known oncologic drug targets and intrinsically disordered proteins (IDPs). ReactomeFI showed the strongest enrichment of IDP–target triangles (enrichment ratio 11.91) relative to alternative networks, including CSN and SIGNOR, supporting its suitability for motif-based biomarker discovery.
+
+Features derived from network topology (e.g., motif participation and connectivity patterns) and protein disorder characteristics were used to train Random Forest and XGBoost classifiers. These models were evaluated for their ability to distinguish protein pairs associated with drug sensitivity from non-informative pairs. Model outputs were integrated into a Biomarker Probability Score (BPS), which ranks proteins by their likelihood of serving as predictive biomarkers.
+
+Applying MarkerPredict across targeted cancer therapies yielded 2,084 candidate predictive biomarkers. Among these, proteins such as LCK and ERK1 were highlighted as high-confidence candidates due to their network positioning and structural features, suggesting relevance for further experimental and clinical validation. The results indicate that proteins embedded in specific signaling motifs and exhibiting intrinsic disorder are more likely to function as effective predictors of therapeutic response.
+
+Overall, the study demonstrates that combining ReactomeFI-derived network topology with protein disorder information and supervised machine learning provides a scalable strategy for predictive biomarker discovery. MarkerPredict offers a complementary approach to existing biomarker identification methods and supports more informed selection of targeted therapies in oncology.
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-9.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-10.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-9.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-10.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-10.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-11.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-10.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-11.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-11.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-12.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-11.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-12.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-12.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-13.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-12.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-13.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-13.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-14.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-13.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-14.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-14.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-15.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-14.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-15.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-15.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-16.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-15.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-16.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-16.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-17.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-16.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-17.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-17.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-18.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-17.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-18.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-18.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-19.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-18.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-19.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-1.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-2.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-1.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-2.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-19.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-20.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-19.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-20.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-20.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-21.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-20.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-21.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-21.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-22.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-21.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-22.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-22.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-23.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-22.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-23.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-23.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-24.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-23.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-24.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-24.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-25.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-24.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-25.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-25.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-26.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-25.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-26.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-26.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-27.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-26.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-27.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-27.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-28.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-27.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-28.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-28.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-29.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-28.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-29.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-2.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-3.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-2.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-3.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-29.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-30.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-29.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-30.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-30.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-31.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-30.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-31.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-31.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-32.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-31.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-32.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-32.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-33.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-32.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-33.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-33.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-34.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-33.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-34.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-34.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-35.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-34.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-35.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-35.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-36.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-35.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-36.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-36.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-37.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-36.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-37.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-37.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-38.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-37.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-38.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-38.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-39.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-38.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-39.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-3.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-4.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-3.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-4.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-39.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-40.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-39.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-40.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-4.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-5.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-4.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-5.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-5.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-6.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-5.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-6.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-6.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-7.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-6.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-7.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-7.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-8.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-7.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-8.mdx
diff --git a/projects/website-angular/content/content/reactome-research-spotlight/article-8.mdx b/projects/website-angular/content/content/reactome-research-spotlight/blogpost-9.mdx
similarity index 100%
rename from projects/website-angular/content/content/reactome-research-spotlight/article-8.mdx
rename to projects/website-angular/content/content/reactome-research-spotlight/blogpost-9.mdx
diff --git a/projects/website-angular/content/documentation/cite.mdx b/projects/website-angular/content/documentation/cite.mdx
index d8732e7..fd91dd5 100644
--- a/projects/website-angular/content/documentation/cite.mdx
+++ b/projects/website-angular/content/documentation/cite.mdx
@@ -1,6 +1,6 @@
---
title: Referencing our Publications
-category: ""
+category: "documentation"
---
## Referencing our Publications
diff --git a/projects/website-angular/content/documentation/curator-guide.mdx b/projects/website-angular/content/documentation/curator-guide.mdx
index 8db3c63..2d68aec 100644
--- a/projects/website-angular/content/documentation/curator-guide.mdx
+++ b/projects/website-angular/content/documentation/curator-guide.mdx
@@ -1,6 +1,6 @@
---
title: Curator Guide
-category: ""
+category: "documentation"
---
## Curator Guide
diff --git a/projects/website-angular/content/documentation/data-model.mdx b/projects/website-angular/content/documentation/data-model.mdx
index cecd506..08af5be 100644
--- a/projects/website-angular/content/documentation/data-model.mdx
+++ b/projects/website-angular/content/documentation/data-model.mdx
@@ -1,6 +1,6 @@
---
title: Data Model
-category: ""
+category: "documentation"
---
## Data Model
diff --git a/projects/website-angular/content/documentation/dev.mdx b/projects/website-angular/content/documentation/dev.mdx
index 273ef4e..b5765db 100644
--- a/projects/website-angular/content/documentation/dev.mdx
+++ b/projects/website-angular/content/documentation/dev.mdx
@@ -1,6 +1,6 @@
---
title: "Developer's Zone"
-category: ""
+category: "documentation"
---
## Developer's Zone
diff --git a/projects/website-angular/content/documentation/dev/analysis.mdx b/projects/website-angular/content/documentation/dev/analysis.mdx
new file mode 100644
index 0000000..c09e6a5
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/analysis.mdx
@@ -0,0 +1,251 @@
+---
+title: Analysis Service
+category: "documentation"
+---
+
+## Analysis Service
+
+#### Explore our tools and web services and learn how to include them in your applications
+
+[ __ ](<#MoreInfo>)
+
+## [ More Info ](<#MoreInfo>)
+
+[ __ ](<#GetStarted>)
+
+## [ Get Started ](<#GetStarted>)
+
+[ __ ]()
+
+## [ API ]()
+
+[ __ ](<#Resources>)
+
+## [ Resources ](<#Resources>)
+
+The analysis tool suite contains an overrepresentation analysis, an expression data analysis and a species comparison tool. The tool suite is available via a [Web Service]( "Web Service") so all the available analysis tools can be easily integrated into third party software.
+
+### [__More Info](<#MoreInfo>)
+
+The Analysis Web-Service is token based, so for every analysis request a TOKEN is associated to the result. From this moment on, the results can be accessed via the API “token” method in order to retrieve more detailed information. Taking advantage of the token produced during every analysis, it is possible to link back to the PathwayBrowser and browse the results overlaid on either the pathways overview or a selected pathway.
+
+As mentioned afore, Reactome provides an overrepresentation analysis, an expression data analysis and a species comparison tool. For overrepresentation and expression analysis, the service can automatically detect which one to perform depending on the format of the submitted data (see [Table 1](<#AnalysisFormat>)). If the data contains expression values in a tab-separated-values (TSV) format, then the expression analysis will be executed, otherwise the overrepresentation analysis is the executed one.
+
+Table 1: Analysis input data formats
+
+
+Data must be submitted in a format that includes a first row of column headers. The header for column 1 must start with the # symbol. Column 1 must contain protein, compound or other suitable identifiers, such as OMIM IDs. Identifiers can only be placed in column 1. A one-column file is sufficient for pathway over-representation analysis. For expression analysis, columns 2 and onwards must contain numeric values, with no alphabetical characters. Decimals must use the full stop (period) symbol, not commas. Columns must be tab-delimited (this format is an option when saving from most spreadsheet programs such as Excel).
+
+The first line must start with “#” to indicate the name of the sample (first column) and the names of each expression column when expression data is submitted. Please note that while this line is optional for the overrepresentation format, it is mandatory for the expression data format, because the name of each column needs to be provided. It is recommended though to have this first line defined in the submitted data, so later on it is easier to identify what kind of experiment the data is related to.
+
+Every other line starts with the identifier of the gene/protein/chemical. For expression analysis, the rest of the columns should contain numbers corresponding to the expression values.
+
+#### [Data Submission](<#DataSubmission>)
+
+The first use case would be when the data to be analysed is submitted using the method:
+
+
+ https://reactome.org/AnalysisService/identifiers/
+
+In this case the data must be sent via POST (please refer to the API documentation for more details) so the analysis will be performed against the Reactome database content.
+
+A second use case is when a text flat file containing the data to be analysed (in the specified format mentioned afore) is sent. In this case the method to be used is:
+
+
+ https://reactome.org/AnalysisService/identifiers/form/
+
+A third use case would be when the data to be analysed is accessible through the Internet. In this case, there is a specific method that indicates to the analysis service where to get the data:
+
+
+ https://reactome.org/AnalysisService/identifiers/url/
+
+Please note all these methods have an option to project the identifiers to human and only show the result in this species (by adding the “projection/” suffix to any of them). Please refer to the API documentation for more details.
+
+#### [Analysis result handling](<#analysis-result-handling>)
+
+Once the analysis is finished, the result is sent back to the user in json format (please refer to the API documentation for more details). One of the fields in the analysis summary object is called “token”. The particular analysis result is associated to this token and it can be retrieved later on using the following method without having to send the same data again:
+
+
+ https://reactome.org/AnalysisService/token/
+
+Reactome ensures that the token will be available for the 7 next days after the analysis. After this period it goes into an LRU queue, so may be available for longer but this cannot be ensured as it depends on the frequency of its usage.
+
+The “pathways” field in the analysis result contains a list of the most significant pathways with the corresponding statistics results. By default, these pathways are sorted from the most to the least significant, so the first item would be the most statistically significant. However the user is advised to take into account the statistical values (pValue and FDR) and also the results coverage in the pathway, indicated by entities and reactions found.
+
+#### [Linking back to Reactome Pathway Browser](<#linking-back>)
+
+If linking back to Reactome to visualise the results is required, there are two methods. Providing the token is enough to link back to the PathwayBrowser and get the Fireworks results overlay view. To do so please replace {TOKEN} in the following URL by the one provided in the result.
+
+
+ https://reactome.org/PathwayBrowser/#DTAB=AN&ANALYSIS=**{TOKEN}**
+
+To build a link back pointing to a specific pathway to overlay the result on its diagram, please replace **{ST_ID}** by the pathway stable identifier (provided in the analysis result for each pathway) and **{TOKEN}** by the one provided in the result.
+
+
+ https://reactome.org/PathwayBrowser/#**{ST_ID}** &DTAB=AN&ANALYSIS=**{TOKEN}**
+
+### [__Get Started](<#GetStarted>)
+
+OK! You’ve reached this far :) so let’s see how to start using the analysis service.
+
+There are two ways to submit your sample of identifiers for analysis. The first method is to POST all the identifiers (or the file containing them). The second involves letting the analysis service know where the sample is by providing its URL. Many types of identifiers that can be submitted, including UniProt, chEBI, Ensembl, miRBase, GenBank/EMBL/DDBJ, RefPep, RefSeq, EntrezGene, OMIM, InterPro, Affymetrix, Agilent, Compound, Illumina, etc. Please get in touch with our This email address is being protected from spambots. You need JavaScript enabled to view it. if your sample identifiers are not supported.
+
+In the following examples we use [curl]() to query the analysis service from the command line. They show how to send some gene names (PIK3C2A, PTEN and UNC5B) to be analysed and also how to provide the location of your data via the **url** interface.
+
+The simplest approach is to send your gene names via POST to the ** **method.
+
+
+ curl -H "Content-Type: text/plain" -d "$(printf '**#Genes** \n**PIK3C2A** \n**PTEN** \n**UNC5B** ')" -X POST --url https://reactome.org/AnalysisService/identifiers/projection/\?pageSize\=1\&page\=1
+
+For long lists, the previous example might be impractical. A more convenient way is to POST the content of a file containing the sample to be analysed. Let’s assume you have a file called **genes.txt** that contains the following set of genes to be analysed:
+
+
+ #Genes
+ PIK3C2A
+ PTEN
+ UNC5B
+
+The command changes to indicate that the data is now taken from the file (but please note that the method of the analysis service remains the same):
+
+
+ curl -H "Content-Type: text/plain" --data-binary @genes.txt -X POST --url https://reactome.org/AnalysisService/identifiers/projection/\?pageSize\=1\&page\=1
+
+Both of the previous commands will produce the result shown in [Figure 1](<#AnalysisResult>)
+
+Figure 1: Analysis result example
+
+
+
+Let’s focus on the retrieved **pathways** list. It only contains one pathway because in the command we have specified **\?pageSize\=1\ &page\=1** and that forces the analysis to provide **ONLY** the most significant pathway for the submitted data. If you want the first 10 results, the way of doing it would be **\?pageSize\=10\ &page\=1**. To see the results from 11 to 20 use **\?pageSize\=10\ &page\=2**. The reason for this **paging** mechanism is to avoid overloading the client with the full set of results. Here we are querying from the command line and probably the memory usage isn’t an issue, but please consider web-clients. **Important note:** If **pageSize** and **page** are not specified, then the whole set of results is retrieved to the client.
+
+Other important fields in the result are **pathwaysFound** , **identifiersNotFound** , **summary.token** and **summary.type**. The first two are self-descriptive, so let’s focus on the **summary**. As already mentioned, the **summary.token** can be used to retrieve the results of a previously performed analysis without the need to submit the sample again. For example, the results of the previous analysis can be easily accessed by simply calling the **https://reactome.org/AnalysisService/token** method and providing the token.
+
+
+ curl https://reactome.org/AnalysisService/token/**MjAxNTEwMjAwNjU0MDBfMzMw** \?pageSize\=1\&page\=1
+
+As illustrated in [Figure 2](<#AnalysisTokenMethods>), the analysis service provides a thorough collection of token-based methods that can be used to access the results of a previously performed analysis.
+
+Figure 2: All the token-based methods provided by the analysis service
+
+
+
+The **summary.type** provides information about the type of the analysis performed and it can be OVERREPRESENTATION, EXPRESSION or SPECIES_COMPARISON.
+
+Every item in the **pathways** array, contains information about a specific pathway, i.e its name, stable identifier, the species it belongs to, the number of entities matching the submitted sample etc. **pValue** and **fdr** show the significance of the pathway as the result of the analysis.
+
+#### Pointing to your resource as data provider for the analysis
+
+Let’s suppose that you have some data to be analysed on your server and you want to point Reactome analysis to retrieve it directly. We use [PRIDE]() data for our examples. More specifically we analyse PRIDE data stored in . This can be done by using the **/identifiers/url** methods:
+
+
+ curl -H "Content-Type: text/plain" -d "https://www.ebi.ac.uk/pride/ws/archive/protein/list/assay/27929.acc" -X POST --url https://reactome.org/AnalysisService/identifiers/url/projection/\?pageSize\=1\&page\=1
+
+Just **POST** ing the **URL** , where the data is, to the **https://reactome.org/AnalysisService/identifiers/url/projection/** method is enough to perform the analysis. The rest of the parameters work exactly as explained above. It is also important to take into account that **URL** s sent to this method can either be **HTTP** or **HTTPS** , if your service uses [secure HTTP](), we can deal with it ;).
+
+#### [I have a JavaScript client. How do I query your service?](<#how-do-i-query>)
+
+First we will create a simple HTML page with one button and a place holder to show a table with the results of the query. Please note we have included the [jQuery library]().
+
+HTML: Base example
+
+
+
+
+ Connection to the Reactome Analysis Service
+ //We are using jQuery for this example
+
+
+
+
+
Connection to the Reactome Analysis Service
+
Please click the button to execute the analysis:
+
+
+
+
+
+
+
+
+
+As you can see in the [Figure 3](<#HTMLExample01Figure>), the resulting page is quite a simple one, but enough for our **get started** purposes.
+
+Figure 3: Example HTML
+
+
+
+The next step is to write the code that connects to the analysis service and presents the data back to the client. Please note that we will use the **/identifiers/url/projection** method in order to request the analysis of [data]() stored in the PRIDE repository.
+
+HTML: Example with JavaScript code included
+
+
+
+
+ Connection to the Reactome Analysis Service
+
+
+
+
+
Connection to the Reactome Analysis Service
+
Please click the button to execute the analysis:
+
+
+
+
+
+
+
+
+
+Quickly analysing the main parts of the [JavaScript code](<#HTMLExample02>) body, we see two main methods: [**$( document ).ready**]() and [**jQuery.ajax**](). The first one ensures the content of the associated callback function will be executed when all the HTML is fully loaded in the [DOM](). The second is responsible for performing the analysis request against Reactome’s analysis service.
+
+Focusing on the **.success** function in the ajax query, we see how the results for a “success” connection with a “success” response are handled (please note that a success connection differs from a success response, since there can be different reasons why a success connection can end up retrieving an error, i.e. data content is not in the right format). The results can be accessed within the **data** object passed to the **.success(function(data, textStatus){…})** method following the format shown in [Figure 1](<#AnalysisResult>).
+
+In the example, a table is created with the retrieved set of pathways ([Figure 4](<#HTMLExample02Figure>)). Each row of the table contains the pathway name, the pValue and the FDR. It’s also important to see how we’ve built the hyperlink from the name to the view of the pathway (with the analysis result overlay) directly to the [Pathway Browser](). Please note that it is also possible to link to the Pathway Browser in order to show the general result overlaid on the pathways overview. The link for this is:
+
+
+ https://reactome.org/PathwayBrowser/#/DTAB=AN&ANALYSIS=**{TOKEN}**
+
+Figure 4: Example HTML with results already loaded
+
+
+
+But this example doesn’t look pretty! Yes, we know but we are explaining how to access Reactome data. Please have a look to the [CSS]() documentation and start learning how to apply the styles to make the look fit with your expectations.
+
+### [__Resources](<#Resources>)
+
+[API Documentation]()
diff --git a/projects/website-angular/content/documentation/dev/content-service/index.mdx b/projects/website-angular/content/documentation/dev/content-service.mdx
similarity index 100%
rename from projects/website-angular/content/documentation/dev/content-service/index.mdx
rename to projects/website-angular/content/documentation/dev/content-service.mdx
diff --git a/projects/website-angular/content/documentation/dev/content-service/diagram-exporter.mdx b/projects/website-angular/content/documentation/dev/content-service/diagram-exporter.mdx
new file mode 100644
index 0000000..988c8a0
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/content-service/diagram-exporter.mdx
@@ -0,0 +1,134 @@
+---
+title: Diagram exporter
+category: "documentation"
+---
+
+## Diagram exporter
+
+### Introduction
+
+In Reactome, pathway diagrams are stored using a custom format. Aimming to allow researchers to include images of their favourite pathway diagrams into their publications, posters or presentations, we have developed the diagram exporter. This tool makes it easy to export pathway diagrams in bitmap format, allowing users to specify the output format, the quality and the decorators. The diagram exporter has been deployed as part of our [Content Service](), which constitutes an easy API that provides access to the Reactome knowledgebase. To learn more about the Content Service and how you can use it, have a look [here]().
+
+This tutorial will walk you through the simple steps of using this tool to generate images of your favourite pathway diagram.
+
+### Get started
+
+Generating the image of a diagram using our API is as easy as just calling the service with the identifier of the diagram and the desired extension (for example ".png", ".jpg" etc.)
+
+
+ [/ContentService/exporter/diagram/R-HSA-169911.png](). Higher quality value results in higher image quality.
+
+
+ [/ContentService/exporter/diagram/R-HSA-169911.png?quality=7]()
+
+
+
+In case the requested pathway has an interactive illustration (EHLD) associated with it, the diagram exporter will export the illustration as an image. In the following example, the EHLD of Hemostasis is requested in png format. Follow this [link]() if you are interested to learn more about our EHLDs.
+
+
+ [/ContentService/exporter/diagram/R-HSA-109582.png]()
+
+
+
+We can also use the _flg_ argument to flag diagram entities (genes, molecules, etc.)
+
+
+ [/ContentService/exporter/diagram/R-HSA-428359.png?flg=Q9NZI8]()
+
+
+
+The selection and flagging arguments work in exactly the same way for EHLDs.
+
+
+ [/ContentService/exporter/diagram/R-HSA-109582.png?sel=R-HSA-983231&flg=THBD]()
+
+
+
+### Analysis overlay
+
+Reactome offers a pathway analysis service that supports enrichment and expression analysis. The diagram exporter allows you to overlay the results of the analysis on top of the exported diagrams. To do so, use the _token_ argument to specify the unique token assosiated with the performed analysis. To learn more about our Analysis Service and how to use it have a look to this [page]().
+
+In the next example, we use the token acquired from an overrepresentation analysis, to overlay the results on top of a diagram and export it in high quality (7) jpeg format.
+
+
+ [/ContentService/exporter/diagram/R-HSA-8937144.jpeg?quality=7&token=]()
+
+
+
+In the same way, we can overlay the results of an expression analysis on top of a diagram and export it in our prefered format and quality. In case the submitted sample (for this type of analysis) includes more than one column, we can specify it by using the _column_ argument, like in the following example.
+
+Please keep in mind that in case a column is not specified (null), the first one is selected by default. Also in case a column is not specified and the requested format is gif, then an animated image with all the columns is generated.
+
+
+ [/ContentService/exporter/diagram/R-HSA-432047.jpg?quality=7&column=1&token=]()
+
+
+
+Analysis results can also be overlaid on top of our interactive illustrations.
+
+
+ [/ContentService/exporter/diagram/R-HSA-69278.png?token=]()
+
+
+
+### Animated GIFs
+
+When you run an expression analysis and want to export an animated GIF with a frame per analysis column, set extension to gif and don’t specify column:
+
+
+ [/ContentService/exporter/diagram/R-HSA-432047.gif?quality=7&sel=R-ALL-879865&token=]()
+
+
+
+In the same way, you can export animaged gifs of EHLDs
+
+
+ [/ContentService/exporter/diagram/R-HSA-69278.gif?quality=7&sel=R-HSA-69242&token=]()
+
+
+
+### Color profiles
+
+Reactome provides several color profiles for diagram and analysis. You can use _diagramProfile_ and _analysisProfile_ arguments to modify the default ones.
+
+
+ [/ContentService/exporter/diagram/R-HSA-879415.png?quality=7&sel=R-ALL-879865&diagramProfile=standard&analysisProfile=strosobar&token=]()
+
+
+
+### Arguments
+
+The folowing table presents a full list of the supported arguments alond with their default values.
+
+Name| Description| Values| Default
+---|---|---|---
+_quality_ | Quality of the image | 1 - 10 | 5
+_format_ | Format of the image | png, jpg, jpeg, gif | png
+_flags_ | List of elements to be flagged | stId, dbId, identifer, geneName | null
+_selection_ | List of elements to be selected | stId, dbId, identifer, geneName | null
+_diagramProfile_ | Color profile for diagram | modern, standard | modern
+_analysisProfile_ | Color profile for analysis | standard, strosobar, copper plus | standard
+_token_ | Analysis token | String | null
+_column_ | The specific expression analysis results column to be overlaid If column is not specified (null), the first one is selected. If column is not specified (null) and format is gif, then an animated gif is generated with all the columns. | Integers | null
+
+###
diff --git a/projects/website-angular/content/documentation/dev/diagram/index.mdx b/projects/website-angular/content/documentation/dev/diagram.mdx
similarity index 100%
rename from projects/website-angular/content/documentation/dev/diagram/index.mdx
rename to projects/website-angular/content/documentation/dev/diagram.mdx
diff --git a/projects/website-angular/content/documentation/dev/diagram/gwt.mdx b/projects/website-angular/content/documentation/dev/diagram/gwt.mdx
new file mode 100644
index 0000000..534f2e3
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/diagram/gwt.mdx
@@ -0,0 +1,195 @@
+---
+title: Diagram GWT
+category: "documentation"
+---
+
+## Diagram GWT
+
+Reactome’s Diagram GWT widget is our original implementation of the diagram viewer in [GWT](). It is meant to be reused by third party resources in order to display Reactome Pathway Diagrams directly in their web pages and enable users to interact with them.
+
+[ __ ](<#MoreInfo>)
+
+## [ More Info ](<#MoreInfo>)
+
+[ __ ](<#GetStarted>)
+
+## [ Get Started ](<#GetStarted>)
+
+[ __ ](<#API>)
+
+## [ API ](<#API>)
+
+[ __ ](<#Resources>)
+
+## [ Resources ](<#Resources>)
+
+### [__More Info](<#MoreInfo>)
+
+[GWT]() is a development toolkit for building and optimizing complex browser-based applications. Its goal is to enable productive development of high-performance web applications without the developer having to be an expert in browser quirks, XMLHttpRequest, and JavaScript. GWT is used by many products at Google, including AdWords, AdSense, Flights, Hotel Finder, Offers, Wallet, Blogger. It’s open source, completely free, and used by thousands of developers around the world.
+
+### [__Get Started](<#GetStarted>)
+
+We assume that you already have your [GWT project]() up and running configured with the [ MAVEN archetype](). So the first step is to add the EBI Nexus repository in your pom.xml file:
+
+
+
+ ...
+
+
+ nexus-ebi-repo
+ The EBI internal repository
+ http://www.ebi.ac.uk/Tools/maven/repos/content/groups/ebi-repo/
+
+ true
+
+
+ false
+
+
+
+
+ nexus-ebi-snapshot-repo
+ The EBI internal snapshot repository
+ http://www.ebi.ac.uk/Tools/maven/repos/content/groups/ebi-snapshots/
+
+ false
+
+
+ true
+
+
+
+
+
+Once the repository is in place, the next thing to do is adding the Reactome's pathways overview dependency:
+
+
+
+ ...
+
+ org.reactome.web
+ diagram
+ 3.2.1
+
+
+
+
+Before writing the code to use the pathways overview, add the dependency in your **gwt.xml** file, so the compiler will find the pathways overview:
+
+
+
+
+
+And finally, assuming the widget is now going to be displayed by its own, the code in your **EntryPoint** class would look like:
+
+
+ /**
+ * For test purposes we assume the diagram loads as a single item in the web page.
+ *
+ * Entry point classes define onModuleLoad().
+ */
+ public class Main implements EntryPoint {
+
+ private final DiagramViewer diagram;
+
+ public Main() {
+ //We create the diagram. Please note that different pathways diagrams can be loaded in the same
+ //instance (the diagram manages the cache of previously loaded diagrams using an LRU)
+ diagram = DiagramFactory.createDiagramViewer();
+ }
+
+ @Override
+ public void onModuleLoad() {
+ Scheduler.get().scheduleDeferred(new Scheduler.ScheduledCommand() {
+ @Override
+ public void execute() {
+ DiagramFactory.CONSOLE_VERBOSE = true; //This is optional (for dev purposes)
+ DiagramFactory.EVENT_BUS_VERBOSE = true; //This is optional (for dev purposes)
+
+ //For this use case we add the diagram as the main and only object of the webpage
+ RootLayoutPanel.get().add(diagram);
+
+ //This will run the loadDiagram method asynchronously, ensuring the DOM has been updated
+ Scheduler.get().scheduleDeferred(new Scheduler.ScheduledCommand() {
+ @Override
+ public void execute() {
+ diagram.loadDiagram("R-HSA-111804");
+ }
+ });
+ }
+ });
+ }
+ }
+
+### [__API](<#API>)
+
+The available methods are defined in the **DiagramViewer** interface in the **org.reactome.web.diagram.client** package. These are:
+
+
+ //subscribe to this event to be notified when the user closes the analysis overlay
+ HandlerRegistration addAnalysisResetHandler(AnalysisResetHandler handler);
+
+ //Subscribe to this event to be notified if the browser does not support canvas, so diagrams can not be displayed
+ HandlerRegistration addCanvasNotSupportedEventHandler(CanvasNotSupportedHandler handler);
+
+ //Subscribe to this event to be notified when the user selects an object in the diagram
+ HandlerRegistration addDatabaseObjectSelectedHandler(GraphObjectSelectedHandler handler);
+
+ //Subscribe to this event to be notified when the user hovers over an object in the diagram
+ HandlerRegistration addDatabaseObjectHoveredHandler(GraphObjectHoveredHandler handler);
+
+ //Subscribe to this event to be notified when a new diagram is displayed in the viewport
+ HandlerRegistration addDiagramLoadedHandler(DiagramLoadedHandler handler);
+
+ //Subscribe to this event to be notified when elements in the diagram are flagged
+ HandlerRegistration addDiagramObjectsFlaggedHandler(DiagramObjectsFlaggedHandler handler);
+
+ //Subscribe to this event to be notified when flagged elements in the diagram are not longer visible
+ HandlerRegistration addDiagramObjectsFlagResetHandler(DiagramObjectsFlagResetHandler handler);
+
+ //Subscribe to this event to be notified when the user selects the "Go to overview" button
+ HandlerRegistration addFireworksOpenedHandler(FireworksOpenedHandler handler);
+
+ //Flags the elements in the diagram which identifiers match with "identifier"
+ void flagItems(String identifier);
+
+ //Highlights the element(s) in the diagram with the "stableIdentifier"
+ void highlightItem(String stableIdentifier);
+
+ //Highlights the element(s) in the diagram with the "dbIdentifier"
+ void highlightItem(Long dbIdentifier);
+
+ //Loads the diagram with the "stableIdentifier"
+ void loadDiagram(String stId);
+
+ //Loads the diagram with the "dbId"
+ void loadDiagram(Long dbId);
+
+ //Resets the analysis overlay
+ void resetAnalysis();
+
+ //Resets the flagged items
+ void resetFlaggedItems();
+
+ //Resets the highlighted items
+ void resetHighlight();
+
+ //Resets the selected items
+ void resetSelection();
+
+ //Selects the element(s) in the diagram with the "stableIdentifier"
+ void selectItem(String stableIdentifier);
+
+ //Selects the element(s) in the diagram with the "stableIdentifier"
+ void selectItem(Long dbIdentifier);
+
+ //Sets the analysis overlay for a given "token" and "resource" (this will query the Analysis Service)
+ void setAnalysisToken(String token, String resource);
+
+ //Set the widget visibility
+ void setVisible(boolean visible);
+
+### [__Resources](<#Resources>)
+
+ * [__GWT Diagram code]()
+ * [__Reactome Pathways Portal code]()
diff --git a/projects/website-angular/content/documentation/dev/diagram/js.mdx b/projects/website-angular/content/documentation/dev/diagram/js.mdx
new file mode 100644
index 0000000..94ed54d
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/diagram/js.mdx
@@ -0,0 +1,117 @@
+---
+title: Diagram JS
+category: "documentation"
+---
+
+## Diagram JS
+
+Reactome's DiagramJs widget is our diagram viewer in an ordinary JavaScript API. It is meant to be reused by third party resources in order to display Reactome Pathway Diagrams directly in their web pages and enable users to interact with them.
+
+### [__Reusing Reactome's Diagram Widget?](<#reuse-diagram-widget>)
+
+To reuse our viewer you need to follow the following steps
+
+1\. Include the diagram javascript dependency in you HTML header
+
+
+
+
+2\. Add a place holder in the body of your web page
+
+
+
+
+3\. Create and initialise the diagram viewer from your javascript code
+
+
+ //Creating the Reactome Diagram widget
+ //Take into account a proxy needs to be set up in your server side pointing to www.reactome.org
+ function onReactomeDiagramReady(){ //This function is automatically called when the widget code is ready to be used
+ var diagram = Reactome.Diagram.create({
+ "placeHolder" : "diagramHolder",
+ "width" : 950, // minimum recommended width
+ "height" : 500
+ });
+
+ //Initialising it to the "Hemostasis" pathway
+ diagram.loadDiagram("R-HSA-109582");
+
+ //Adding different listeners
+
+ diagram.onDiagramLoaded(function (loaded) {
+ console.info("Loaded ", loaded);
+ diagram.flagItems("FYN");
+ if (loaded == "R-HSA-109582") diagram.selectItem("R-HSA-109582");
+ });
+
+ diagram.onObjectHovered(function (hovered){
+ console.info("Hovered ", hovered);
+ });
+
+ diagram.onObjectSelected(function (selected){
+ console.info("Selected ", selected);
+ });
+ }
+
+### [__DiagramJs API](<#API>)
+
+The current implementation supports the following listeners and methods:
+
+Method| Params| Description
+---|---|---
+**create** :: Constructor
+Reactome.Diagram.create(params); | **param** :: json object
+{
+'proxyPrefix' : string,
+'placeHolder' : string,
+'width' : int (optional),
+'height' : int (optional)
+} | Creates and returns a new Reactome.Diagram object
+**loadDiagram(stId)** :: void | Pathway stable identifier
+**stId** : string | Loads the specified pathway
+**flagItems(term)** :: void | Entity identifier (gene name)
+**term** : string | Flags all entities matching with the "term"
+**highlightItem(stId)** :: void | Item stable identifier
+**stId** : string | Highlights the specified item if it exists in the diagram
+**resetAnalysis()** :: void | | Resets the analysis overlay
+**resetFlaggedItems()** :: void | | Resets the flagged items
+**resetHighlight()** :: void | | Clears the highlights in the diagram
+**resetSelection()** :: void | | Clears the selection in the diagram
+**resize(width, height)** :: void | **widht** : int
+**height** : int | Resizes the viewport to the specified width and height
+**selectItem(stId)** :: void | Item stable identifier
+**stId** : string | Selects the specified item if it exists in the diagram
+**setAnalysisToken(token, resultFilter)** :: void | Analysis token
+**token** : string
+Resource
+**resultFilter** : **obj** **obj** resultFilter: {
+'resource' : string,
+'pValue' : double,(optional)
+'includeDisease' : boolean,(optional)
+'min' : Integer, (optional) 'max':Integer, (optioinal) 'speciesList': [List]
+} | Overlays the analysis result corresponding to the specified (token, resultFilter)
+**onObjectSelected(function(obj))** :: void | **obj** is selected item:
+{
+'stId' : string,
+'displayName' : string,
+'schemaClass' : string,
+'identifier' : string (optional),
+'geneNames' : Array (optional)
+} | The function is called when an object in the diagram is selected by the user action
+**onObjectHovered(function(obj))** :: void | **obj** is hovered item:
+{
+'stId' : string,
+'displayName' : string,
+'schemaClass' : string,
+'identifier' : string (optional),
+'geneNames' : Array (optional)
+} | The function is called when an object in the diagram is hovered by the user action
+**onDiagramLoaded(function(stId))** :: void | The stable identifier of the loaded diagram
+**stId** : string | The function is called when a diagram is loaded in the viewer
+**onFlagsReset(function())** :: void | The function receives no parameters | The function is called when the flagged items have been reset by the user action
+**onAnalysisReset(function())** :: void | The function receives no parameters | The function is called when the users resets the analysis overlay
+
+diagram.onAnalysisReset(function(){ /* your code here */ });
+**onCanvasNotSupported(function())** :: void | The function receives no parameters. | The function is called when the browser doesn't support HTML5 Canvas so the viewer cannot be instantiated
+
+diagram.onCanvasNotSupported(function(){ /* your code here */ });
diff --git a/projects/website-angular/content/documentation/dev/diagram/pathway-diagram-specs.mdx b/projects/website-angular/content/documentation/dev/diagram/pathway-diagram-specs.mdx
new file mode 100644
index 0000000..bc2ded9
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/diagram/pathway-diagram-specs.mdx
@@ -0,0 +1,966 @@
+---
+title: Pathway Diagrams Specifications
+category: "documentation"
+---
+
+## Pathway Diagrams Specifications
+
+## Introduction
+
+This document provides a set of guidelines for developers on how to properly render Reactome pathway diagrams. Pathway Diagrams represent pathways as a series of connected molecular events, known in Reactome as ‘reactions’, which can be considered as steps in the pathway. For a more detailed guide about diagrams refer to [our user guide]().
+
+Currently, there are two projects that follow these guidelines:
+
+ * diagram: [ https://github.com/reactome-pwp/diagram ]()
+ * diagram-exporter: [ https://github.com/reactome-pwp/diagram-exporter ]()
+
+Regulation of Apoptosis diagram
+
+## Diagram layout and graph files
+
+All the class diagrams can be found as UML in xmi (Umbrello) format. [Download]()
+
+The layout information of every diagram is stored in a JSON file, named after its stable identifier, i.e. _R-HSA-12345.json_ . This file contains all the information regarding what to draw and how to draw it, including coordinates and sizes. In addition to the layout file, Reactome provides another JSON file per diagram containing a small graph view of the diagram content, i.e. _R-HSA-12345.graph.json_. We discuss this diagram graph file in more detail further on.
+
+_The general structure of a diagram _layout_ file: _
+
+
+ {
+ "dbId":169911,
+ "stableId":"R-HSA-169911",
+ "displayName":"Diagram of Regulation of Apoptosis",
+ "disease":false,
+ "minX":61,
+ "maxX":1126,
+ "minY":202,
+ "maxY":656,
+ "nodes":[],
+ "edges":[],
+ "compartments":[],
+ "notes":[],
+ "links":[],
+ "shadows":[]
+ }
+
+
+_To get a more detailed description of the diagram _layout_ , expand the following tree:_
+
+ * Diagram
+ * nodes : **Node[]**
+ * nodeAttachments : **NodeAttachment[]**
+ * reactomeId : **Long**
+ * label : **String**
+ * description : **String**
+ * shape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * interactorsSummary : **SummaryItem**
+ * pressed : **Boolean**
+ * shape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * hit : **Boolean**
+ * type : **String**
+ * number : **Integer**
+ * trivial : **Boolean**
+ * connectors : **Connector[]**
+ * edgeId : **Long**
+ * segments : **Segment[]**
+ * to : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * from : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * endShape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * stoichiometry : **Stoichiometry**
+ * shape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * value : **Integer**
+ * isDisease : **Boolean**
+ * isFadeOut : **Boolean**
+ * type : **String**
+ * [_NodeCommon_] prop : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] innerProp : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] identifier : **Identifier**
+ * resource : **String**
+ * id : **String**
+ * [_NodeCommon_] textPosition : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_NodeCommon_] insets : **Bound**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] bgColor : **Color**
+ * r : **Integer**
+ * g : **Integer**
+ * b : **Integer**
+ * [_NodeCommon_] fgColor : **Color**
+ * r : **Integer**
+ * g : **Integer**
+ * b : **Integer**
+ * [_NodeCommon_] isCrossed : **Boolean**
+ * [_NodeCommon_] needDashedBorder : **Boolean**
+ * [_DiagramObject_] reactomeId : **Long**
+ * [_DiagramObject_] schemaClass : **String**
+ * [_DiagramObject_] renderableClass : **String**
+ * [_DiagramObject_] position : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_DiagramObject_] isDisease : **Boolean**
+ * [_DiagramObject_] isFadeOut : **Boolean**
+ * [_DiagramObject_] minX : **Double**
+ * [_DiagramObject_] minY : **Double**
+ * [_DiagramObject_] maxX : **Double**
+ * [_DiagramObject_] maxY : **Double**
+ * [_DiagramObject_] id : **Long**
+ * [_DiagramObject_] displayName : **String**
+ * isDisease : **Boolean**
+ * minX : **Integer**
+ * minY : **Integer**
+ * maxX : **Integer**
+ * maxY : **Integer**
+ * notes : **Note[]**
+ * [_NodeCommon_] prop : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] innerProp : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] identifier : **Identifier**
+ * resource : **String**
+ * id : **String**
+ * [_NodeCommon_] textPosition : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_NodeCommon_] insets : **Bound**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] bgColor : **Color**
+ * r : **Integer**
+ * g : **Integer**
+ * b : **Integer**
+ * [_NodeCommon_] fgColor : **Color**
+ * r : **Integer**
+ * g : **Integer**
+ * b : **Integer**
+ * [_NodeCommon_] isCrossed : **Boolean**
+ * [_NodeCommon_] needDashedBorder : **Boolean**
+ * [_DiagramObject_] reactomeId : **Long**
+ * [_DiagramObject_] schemaClass : **String**
+ * [_DiagramObject_] renderableClass : **String**
+ * [_DiagramObject_] position : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_DiagramObject_] isDisease : **Boolean**
+ * [_DiagramObject_] isFadeOut : **Boolean**
+ * [_DiagramObject_] minX : **Double**
+ * [_DiagramObject_] minY : **Double**
+ * [_DiagramObject_] maxX : **Double**
+ * [_DiagramObject_] maxY : **Double**
+ * [_DiagramObject_] id : **Long**
+ * [_DiagramObject_] displayName : **String**
+ * edges : **Edge[]**
+ * [_EdgeCommon_] segments : **Segment[]**
+ * to : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * from : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_EdgeCommon_] endShape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * [_EdgeCommon_] catalysts : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] inhibitors : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] activators : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] precedingEvents : **Long[]**
+ * [_EdgeCommon_] reactionType : **String**
+ * [_EdgeCommon_] followingEvents : **Long[]**
+ * [_EdgeCommon_] interactionType : **String**
+ * [_EdgeCommon_] reactionShape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * [_EdgeCommon_] inputs : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] outputs : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_DiagramObject_] reactomeId : **Long**
+ * [_DiagramObject_] schemaClass : **String**
+ * [_DiagramObject_] renderableClass : **String**
+ * [_DiagramObject_] position : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_DiagramObject_] isDisease : **Boolean**
+ * [_DiagramObject_] isFadeOut : **Boolean**
+ * [_DiagramObject_] minX : **Double**
+ * [_DiagramObject_] minY : **Double**
+ * [_DiagramObject_] maxX : **Double**
+ * [_DiagramObject_] maxY : **Double**
+ * [_DiagramObject_] id : **Long**
+ * [_DiagramObject_] displayName : **String**
+ * links : **Link[]**
+ * [_EdgeCommon_] segments : **Segment[]**
+ * to : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * from : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_EdgeCommon_] endShape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * [_EdgeCommon_] catalysts : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] inhibitors : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] activators : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] precedingEvents : **Long[]**
+ * [_EdgeCommon_] reactionType : **String**
+ * [_EdgeCommon_] followingEvents : **Long[]**
+ * [_EdgeCommon_] interactionType : **String**
+ * [_EdgeCommon_] reactionShape : **Shape**
+ * r : **Double**
+ * b : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * a : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * c : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * s : **String**
+ * r1 : **Double**
+ * empty : **Boolean**
+ * type : **String**
+ * [_EdgeCommon_] inputs : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_EdgeCommon_] outputs : **ReactionPart[]**
+ * stoichiometry : **Integer**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * id : **Long**
+ * [_DiagramObject_] reactomeId : **Long**
+ * [_DiagramObject_] schemaClass : **String**
+ * [_DiagramObject_] renderableClass : **String**
+ * [_DiagramObject_] position : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_DiagramObject_] isDisease : **Boolean**
+ * [_DiagramObject_] isFadeOut : **Boolean**
+ * [_DiagramObject_] minX : **Double**
+ * [_DiagramObject_] minY : **Double**
+ * [_DiagramObject_] maxX : **Double**
+ * [_DiagramObject_] maxY : **Double**
+ * [_DiagramObject_] id : **Long**
+ * [_DiagramObject_] displayName : **String**
+ * compartments : **Compartment[]**
+ * componentIds : **Long[]**
+ * [_NodeCommon_] prop : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] innerProp : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] identifier : **Identifier**
+ * resource : **String**
+ * id : **String**
+ * [_NodeCommon_] textPosition : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_NodeCommon_] insets : **Bound**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * [_NodeCommon_] bgColor : **Color**
+ * r : **Integer**
+ * g : **Integer**
+ * b : **Integer**
+ * [_NodeCommon_] fgColor : **Color**
+ * r : **Integer**
+ * g : **Integer**
+ * b : **Integer**
+ * [_NodeCommon_] isCrossed : **Boolean**
+ * [_NodeCommon_] needDashedBorder : **Boolean**
+ * [_DiagramObject_] reactomeId : **Long**
+ * [_DiagramObject_] schemaClass : **String**
+ * [_DiagramObject_] renderableClass : **String**
+ * [_DiagramObject_] position : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_DiagramObject_] isDisease : **Boolean**
+ * [_DiagramObject_] isFadeOut : **Boolean**
+ * [_DiagramObject_] minX : **Double**
+ * [_DiagramObject_] minY : **Double**
+ * [_DiagramObject_] maxX : **Double**
+ * [_DiagramObject_] maxY : **Double**
+ * [_DiagramObject_] id : **Long**
+ * [_DiagramObject_] displayName : **String**
+ * shadows : **Shadow[]**
+ * prop : **NodeProperties**
+ * x : **Double**
+ * y : **Double**
+ * width : **Double**
+ * height : **Double**
+ * points : **Coordinate[]**
+ * x : **Double**
+ * y : **Double**
+ * colour : **String**
+ * [_DiagramObject_] reactomeId : **Long**
+ * [_DiagramObject_] schemaClass : **String**
+ * [_DiagramObject_] renderableClass : **String**
+ * [_DiagramObject_] position : **Coordinate**
+ * x : **Double**
+ * y : **Double**
+ * [_DiagramObject_] isDisease : **Boolean**
+ * [_DiagramObject_] isFadeOut : **Boolean**
+ * [_DiagramObject_] minX : **Double**
+ * [_DiagramObject_] minY : **Double**
+ * [_DiagramObject_] maxX : **Double**
+ * [_DiagramObject_] maxY : **Double**
+ * [_DiagramObject_] id : **Long**
+ * [_DiagramObject_] displayName : **String**
+ * dbId : **Long**
+ * stableId : **String**
+ * cPicture : **String**
+ * forNormalDraw : **Boolean**
+ * displayName : **String**
+
+## Diagram objects
+
+Diagram layout is made up of six types of elements:
+
+ * **Compartments**
+ * **Nodes**
+ * **Edges**
+ * **Links**
+ * **Notes**
+ * **Shadows**
+
+These elements follow this hierarchy.
+
+
+
+### Compartments
+
+
+
+Compartments are rendered in the background and represent where reactions happen. They are rendered as rounded rectangles using _NodeCommon.prop_. Compartments might have an additional layer when surrounded by a membrane, which must be specified using _NodeCommon.innerProp. Text (_DiagramObject.displayName_) is displayed in one line, using _NodeCommon.textPosition._ Color is taken from the diagram profile sheet (_Profile.compartment_). Compartments cannot be rendered on top of any other node._
+
+### Nodes
+
+Nodes represent the participants of reactions, i.e. inputs, outputs, catalysts, regulators and other pathways. The type of each node is coded into _renderableClass._ The renderable class can be one of **Chemical, ChemicalDrug, Complex, Entity, EntitySet, Gene, ProcessNode, EncapsulatedNode, Protein** and **RNA.**
+
+Nodes are laid in 2 layers, background and foreground. The background is rendered using a different shape for each class with dimensions defined in _NodeCommon.prop_. Some nodes may have a foreground (**ProcessNode** , **EncapsulatedNode** and **EntitySet**). The foreground shape and colour are class specific. The text is centred to the node with a padding of 5 points to the _NodeCommon.prop_. In case it is too large, then it is wrapped in several lines and, if needed, the size of the font is reduced. When a foreground is present, the text is padded to foreground limits.
+
+**Class**| **Background**| **Foreground**| **Example**
+---|---|---|---
+_Chemical_ | ellipse | | 
+_ChemicalDrug_ | ellipse | | 
+_Complex_ | edged rectangle (octagon) | | 
+_Entity_ | rectangle | | 
+_EntitySet_ | rounded rectangle | rounded rectangle (padding = 4) | 
+_* Gene_ | recatangle with 2 rounded corners | | 
+_ProcessNode_ | rectangle | rectangle (padding = 10) | 
+_EncapsulatedNode_ | hexagon | hexagon (padding = 10) | 
+_Protein_ | rounded rectangle | | 
+___ RNA _ | bone | | 
+
+*Genes are made up using a different approach. They have 2 shapes, a rectangle with 2 rounded corners (y=NodeCommon.prop.y + 25) and a triangle. The rectangle is filled but not stroked. Both shapes are joint using 3 perpendicular lines.
+
+
+ shape = SemiRoundedRectangle(prop.x, prop.y + 25, prop.width. prop.height)
+ arrow = Path()
+ arrow.moveTo(prop.maxX, prop.getY + 8)
+ arrow.lineTo(prop.maxX, prop.getY - 8)
+ arrow.lineTo(prop.maxX + 8, prop.getY)
+ arrow.closePath()
+ path = Path()
+ path.moveTo(prop.x, prop.y + 25)
+ path.lineTo(prop.maxX, prop.y + 25)
+ path.moveTo(prop.maxX - 4, prop.y + 25)
+ path.lineTo(prop.maxX - 4, prop.y)
+ path.lineTo(prop.maxX, prop.y) # already closed
+ fill(shape, arrow)
+ stroke(path, arrow)
+
+
+_RNA shape looks after a bone_ :
+
+
+ x1 = rna.x + 16
+ x2 = rna.maxX - 16
+ y1 = rna.y + 8
+ y2 = rna.maxY - 8
+ path = Path()
+ path.moveTo(x1, y1)
+ path.lineTo(x2, y1)
+ path.quadTo(rna.maxX, rna.y, rna.maxX, rna.centerY)
+ path.quadTo(rna.maxX, rna.maxY, x2, y2)
+ path.lineTo(x1, y2)
+ path.quadTo(rna.x, rna.maxY, rna.x, rna.centerY)
+ path.quadTo(rna.x, rna.y, x1, y1)
+ fill(path)
+ stroke(path)
+
+
+Some nodes contain attachments, such as post-translational modifications (PTMs) . Attachments are styled after their owner. The border of nodes is rendered after the background and foreground shapes, except for genes, which have a custom border shape. When a node has _NodeCommon.needDashedBorder_ then the border must be dashed.
+
+Drugs use the same shape as chemicals, but with different colors. They also have a small reactangle (14x7) in the bottom right corner with the text Rx (recipe, latin word for prescription).
+
+### Edges
+
+Edges represent reactions in the diagram. They are layout as lines with shapes. They can have 2 shapes: a _reaction shape_ , to indicate the type of reaction, and an _end shape_. Both can be omitted. Lines are taken from _EdgeCommon.segments_. Shape is created using several properties inside _EdgeCommon_.
+
+**Reaction type**| **shape.type**| **shape.empty**| **shape.text**| **Example**
+---|---|---|---|---
+Association | circle | fill | | 
+Dissociation | double circle | empty | | 
+Omitted process | box | empty | \\\ | 
+Transition | box | empty | | 
+Uncertain | box | empty | ? | 
+All (end shape) | arrow | fill | | 
+
+Shapes are specified in the _Shape_ element inside _Edge.reactionType_ and _Edge.endShape_.
+
+**_Shape.type_ **| **Shape**
+---|---
+ARROW | triangle with points: a, b, c
+BOX | rectangle from a (top left) to b (bottom right)
+CIRCLE | circle with center c and radius r
+STOP | line from a to b
+DOUBLE CIRCLE | 2 circles with center c and radii r and r1
+
+When _Shape.s_ is present, the value of _Shape.s_ is written in the centre of the shape. When _Shape.emtpy_ is true, the Shape is filled with _Profile.reaction.fill_ color (usually white), otherwise, it is filled with _Profile.reaction.stroke_ (usually black).
+
+### Connectors
+
+As each reaction can have several participants, a connector is created in the participant with _Connector.EdgeId_ = _DiagramObject.Id_. Connectors are styled after the reaction they belong. Connectors have segments and 2 shapes: _end shape_ and _stoichiometry shape_. Stoichiometries are always empty boxes with the stoichiometry value as text only if stoichiometry value is greater than 1.
+
+**endShape.type**| **Shape**| **Empty**| **Example**
+---|---|---|---
+Inhibitor | line | fill | 
+Catalyst | circle | empty | 
+Output | arrow | fill | 
+Activator | arrow | empty | 
+Input | _no shape_ | |
+
+### Links
+
+Links are used for linking elements which are related, normally subpathways or distant nodes.
+
+**Renderable class**| **Dashed**| **Link type**| **Shape**| **Empty**| **Example**
+---|---|---|---|---|---
+EntitySetAndMemberLink | dashed | | | | 
+EntitySetAndEntitySetLink | dashed | | | | 
+FlowLine | false | | ARROW | fill | 
+Interaction | false | Activate | ARROW | empty | 
+Interaction | false | Inhibit | ARROW | empty | 
+
+### Notes
+
+Notes are just texts. The text is displayed in one line beginning at _NodeCommon.textPosition._ Color is taken from _Profile.note.text_.
+
+### Shadows
+
+Shadows are used when there are several subpathways in the same diagram. In particular, these coloured rectangles highlight the specific diagram reactions belonging to each subpathway, allowing further zooming in to the areas of interest within a given pathway diagram. Rendering them is optional, but it is recommended that they are drawn on top of every element, as rectangles, using _Shadow.points_ and _Shadow.color_ for text and filling. It is also adviced to use transparency for the filling so that all subpathway participants are visible. The text contains the name of the subpathway and should be rendered using a bigger font size.
+
+## Text
+
+The default font for texts is Arial black with size 9. For shadows, font size is 24.
+
+## Colouring
+
+Element colors are taken from a JSON stylesheet. Currently, we have 2 styles: [ standard ]() and [ modern]().
+
+_To get a more detailed description of the diagram colour profile, expand the following tree:_
+
+ * DiagramProfile
+ * stoichiometry : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * entity : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * complex : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * note : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * interactor : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * attachment : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * flowline : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * gene : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * processnode : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * encapsulatednode : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * protein : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * link : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * otherentity : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * chemical : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * chemicaldrug : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * entityset : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * compartment : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * reaction : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * rna : **DiagramProfileNode**
+ * stroke : **String**
+ * lineWidth : **String**
+ * fadeOutFill : **String**
+ * fadeOutStroke : **String**
+ * lighterFill : **String**
+ * lighterStroke : **String**
+ * text : **String**
+ * fadeOutText : **String**
+ * lighterText : **String**
+ * fill : **String**
+ * thumbnail : **DiagramProfileThumbnail**
+ * hovering : **String**
+ * highlight : **String**
+ * selection : **String**
+ * edge : **String**
+ * node : **String**
+ * name : **String**
+ * properties : **DiagramProfileProperties**
+ * hovering : **String**
+ * highlight : **String**
+ * selection : **String**
+ * halo : **String**
+ * flag : **String**
+ * trigger : **String**
+ * button : **String**
+ * text : **String**
+ * disease : **String**
+
+The **DiagramProfileNode** defines 3 colors per each node: _fill, stroke_ and _text_ . If an object is fadeout (_DiagramObject.isFadeOut),_ then we must choose _fadeOutFill, fadeOutStroke_ and _fadeOutText_. If an analysis is being run, then we use _lighterFill, lighterStroke_ and _lighterText_. And by default _fill, stroke_ and _text_.
+
+There is 1 exception: reactions and stoichiometries’ texts; text color is ignored and stroke is used instead. When _DiagramObject.isDisease_ is true, the color of node border, edge border, edge text and segments is changed to _Profile.properties.disease_ (usually red).
+
+### Decorators
+
+Some elements can be decorated in the diagram. This information is not in the diagram and has to be inserted from the outside. Reactome supports 2 types of decoration: **selection** and **flagging**.
+
+When a node is selected, the border is changed in 2 ways: the color is changed to _Profile.properties.selection_ and the width is increased. The node, the reactions in which it participates and the nodes that participate in these reactions are haloed. Haloing is made by drawing the border behind the node with a larger width and with _Profile.properties.halo_ color.
+
+When a reaction is selected, its segments, texts and borders change color to _Profile.properties.selection_ and segments increase its width. The reaction and its participants are haloed.
+
+When a node is flagged, the border is drawn in the background with a larger width using _Profile.properties.flag._ At this moment, edges are not being flagged.
+
+ Representation of layers for a node. From bottom to top: flag, halo, background, analysis, foreground, border and text.
+
+ Edge layers. From bottom to top: halo, segments, filling, border and text.
+
+## Analysis
+
+When an analysis is overlaid, we must modify the color of elements to _lighter_. Then, those nodes hit by the analysis must render a new layer between the background and the foreground. At first, we must calculate how much a node is hit by the analysis. We can do it by using the graph JSON file and the analysis results (or the analysis token). We must traverse the graph from each node through its children (_EntityNode.children)_.
+
+ C1 has 1/2 children hit. S1 has 3/4 leaves hit. P1, P3 and P4 have 1/1 leaves hit.
+
+_To get a more detailed description of the diagram graph, expand the following tree:_
+
+ * Graph
+ * nodes : **EntityNode[]**
+ * identifier : **String**
+ * parents : **Long[]**
+ * children : **Long[]**
+ * geneNames : **String[]**
+ * diagramIds : **Long[]**
+ * [_GraphNode_] schemaClass : **String**
+ * [_GraphNode_] dbId : **Long**
+ * [_GraphNode_] stId : **String**
+ * [_GraphNode_] speciesID : **Long**
+ * [_GraphNode_] displayName : **String**
+ * edges : **EventNode[]**
+ * catalysts : **Long[]**
+ * inhibitors : **Long[]**
+ * activators : **Long[]**
+ * inputs : **Long[]**
+ * outputs : **Long[]**
+ * diagramIds : **Long[]**
+ * preceding : **Long[]**
+ * following : **Long[]**
+ * requirements : **Long[]**
+ * [_GraphNode_] schemaClass : **String**
+ * [_GraphNode_] dbId : **Long**
+ * [_GraphNode_] stId : **String**
+ * [_GraphNode_] speciesID : **Long**
+ * [_GraphNode_] displayName : **String**
+ * dbId : **Long**
+ * stId : **String**
+ * subpathways : **SubpathwayNode[]**
+ * dbId : **Long**
+ * stId : **String**
+ * events : **Long[]**
+ * displayName : **String**
+
+ProcessNodes percentages are taken from the _SubpathwaySummary_ : entity.found / entity.total. For the analysis, there is another stylesheet. We draw enrichment using the _AnalysisSheet.enrichment.gradient.max_ color. There are 3 analysis stylesheets: [ Standard](), [ Copper Plus]() and [ Strosobar]().
+
+_To get a more detailed description of the analysis colour profile, expand the following tree:_
+
+ * AnalysisProfile
+ * enrichment : **OverlayNode**
+ * text : **String**
+ * gradient : **ProfileGradient**
+ * stop : **String**
+ * min : **String**
+ * max : **String**
+ * legend : **OverlayLegend**
+ * hover : **String**
+ * median : **String**
+ * expression : **OverlayNode**
+ * text : **String**
+ * gradient : **ProfileGradient**
+ * stop : **String**
+ * min : **String**
+ * max : **String**
+ * legend : **OverlayLegend**
+ * hover : **String**
+ * median : **String**
+ * ribbon : **String**
+ * name : **String**
+
+If the analysis is an expression analysis, we must take the individual expression values from the leaves. We can show only 1 column at a time. If we want to show them all we must create an animation (play button in the pathway browser and GIFs in the diagram exporter). Once we have the expression values for an element, we divide it into regions and use _AnalysisSheet.expression.gradient_ to calculate the color of each region, using _AnalysisResult.expression.max_ and _AnalysisResult.expression.min_ as limit values. Leaves are sorted using the identifier from the analysis (_FoundEntity.id_).
+
+ Subsection of a diagram with an analysis overlaid  Example of a diagram with an expression analysis
+
+## Elements order
+
+From bottom to top, elements must be rendered in the following order:
+
+ 1. Compartments
+ 2. Fade out reactions
+ 3. Fade out nodes
+ 4. Flags
+ 5. Halos
+ 6. Reactions
+ 7. Nodes
+ 8. Notes
+ 9. Shadows
+
+## More Information
+
+To learn more about our pathway diagrams and the techniques we use to render them efficiently please go through our relevant publications:
+
+ 1. [Reactome diagram viewer: data structures and strategies to boost performance]()
+ 2. [Reactome enhanced pathway visualization]()
diff --git a/projects/website-angular/content/documentation/dev/graph-database/index.mdx b/projects/website-angular/content/documentation/dev/graph-database.mdx
similarity index 100%
rename from projects/website-angular/content/documentation/dev/graph-database/index.mdx
rename to projects/website-angular/content/documentation/dev/graph-database.mdx
diff --git a/projects/website-angular/content/documentation/dev/graph-database/extract-participating-molecules.mdx b/projects/website-angular/content/documentation/dev/graph-database/extract-participating-molecules.mdx
new file mode 100644
index 0000000..109a988
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/graph-database/extract-participating-molecules.mdx
@@ -0,0 +1,504 @@
+---
+title: "Tutorial: Extracting participating molecules using the Graph Database"
+category: "documentation"
+---
+
+## Tutorial: Extracting participating molecules using the Graph Database
+
+ * [The participating molecules use case](<#participating-molecules-use-case>)
+ * [Retrieving objects based on their identifier](<#retrieving-objects>)
+
+ * [Identifiers for proteins or chemicals](<#identifiers-proteins-or-chemicals>)
+
+ * [Breaking down complexes and sets to get their participants](<#complexes-sets-participants>)
+
+ * [Retrieving pathways,subpathways and superpathways](<#retrieving-pathways>)
+
+ * [Retrieving the reactions for a given pathway](<#retrieving-reactions>)
+
+ * [Retrieving the participants of a Reaction](<#retrieving-participants>)
+
+ * [Joining the pieces: Participating molecules for a pathway](<#joining-pieces>)
+
+The Reactome Graph Database, also called a graph-oriented database, is a type of NoSQL database that uses graph theory to store, map and query relationships relating to our data content. Each node represents an entity (such as a pathway, reaction or proteins) and each edge represents a relationship between two nodes. Every node in our graph database is defined by a unique identifier, a set of outgoing edges and/or incoming edges and a set of properties expressed as key/value pairs. Each edge is defined by a unique identifier, a starting-place and/or ending-place node and a set of properties.
+
+[Cypher]() is Neo4j’s open graph query language. Cypher’s syntax provides a familiar way to match patterns of nodes and relationships in a graph. If you want to learn more about what is Cypher, please visit the following [link]().
+
+Graph databases are well-suited for analyzing interconnections, which is why there has been a lot of interest in using graph databases to mine data from biological pathways and reactions. The Reactome Graph database has many advantages, but one is its responsiveness in managing data. Furthermore, even though data queries increase exponentially, the performance of a graph database does not drop, compared to what happens with relational databases. When software developers work with data, they are looking for flexibility and scalability. Our Graph Database contributes a lot in this regard because when needs increase, the possibilities of adding more nodes and relationships to an existing graph are huge.
+
+### [The participating molecules use case](<#participating-molecules-use-case>)
+
+This tutorial explains how to query Reactome using Cypher. It's assumed that the reader has a basic knowledge of Cypher as well as an understanding of our [data model]() and how data is stored in our [schema]().
+
+Even though it is not possible to cover all possible queries of the Reactome Graph Database in a single tutorial, the sections in this document build up a query, which will retrieve the resource and identifier of each participating molecule of a given Pathway. There are several intermediate stages to be explained before reaching that point:
+
+ 1. How to retrieve objects like proteins, reactions, pathways, etc.
+ 2. How to get the identifier of proteins or chemicals
+ 3. How to deconstruct complexes or sets to get their participants
+ 4. How to retrieve the subpathways for a given pathway
+ 5. How to retrieve the reactions of a pathway
+ 6. How to retrieve the participants of a reaction
+
+The enumeration above represents the **basic bricks** from which to construct the final query that retrieves the **participating molecules** for a given **pathway**.
+
+### [Retrieving objects based on their identifier](<#retrieving-objects>)
+
+To retrieve the Pathway "**Antigen processing-Cross presentation** " with identifier **R-HSA-1236975** , the query is as follows:
+
+
+ //Selecting an Pathway by its stable identifier
+ MATCH (pathway:Pathway{stId:"R-HSA-1236975"})
+ RETURN pathway
+
+
+The result of the query is:
+
+
+ ╒═════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
+ │pathway │
+ ╞═════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
+ │{speciesName: Homo sapiens, oldStId: REACT_111119, isInDisease: false, releaseDate: 2011-09-20, displayName: Antigen │
+ │ processing-Cross presentation, stIdVersion: R-HSA-1236975.1, dbId: 1236975, releaseStatus: UPDATED, name: [Antigen │
+ │ processing-Cross presentation], stId: R-HSA-1236975, hasDiagram: false, isInferred: false} │
+ └─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┘
+
+
+In the same way, let's focus on an EntityWithAccessionedSequence (EWAS) which corresponds to a protein in Reactome. For this example we use one form of **PTEN** in the **cytosol** with identifier **R-HSA-199420**
+
+
+ //Selecting an EWAS by its stable identifier
+ MATCH (ewas:EntityWithAccessionedSequence{stId:"R-HSA-199420"})
+ RETURN ewas
+
+
+The result is one node of the database:
+
+
+ ╒════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
+ │ewas │
+ ╞════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
+ │{speciesName: Homo sapiens, startCoordinate: 1, isInDisease: false, displayName: PTEN [cytosol], dbId: 199420, name: [PTEN, │
+ │ Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase PTEN, PTEN_HUMAN, MMAC1, TEP1], referenceType: ReferenceGeneProduct,│
+ │ endCoordinate: 403, stId: R-HSA-199420} │
+ └────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┘
+
+
+### [Identifiers for proteins or chemicals](<#identifiers-proteins-or-chemicals>)
+
+Continuing with the same form of PTEN, another use case could be to retrieve only a couple of fields for the target node. The following query retrieves the EWAS **display name** and its **identifier** (from the reference entity):
+
+
+ //Following the reference entity link in order to get the identifier
+ MATCH (ewas:EntityWithAccessionedSequence{stId:"R-HSA-199420"}),
+ (ewas)-[:referenceEntity]->(re:ReferenceEntity)
+ RETURN ewas.displayName AS EWAS, re.identifier AS Identifier
+
+
+Please note that the **identifier** is not directly stored in the node for the EWAS but is a property of another node pointed from the EWAS which is a **ReferenceEntity**. The previous query accesses that node from the EWAS following the **referenceEntity** edge. The result of this query is as shown below:
+
+
+ ╒══════════════╤══════════╕
+ │EWAS │Identifier│
+ ╞══════════════╪══════════╡
+ │PTEN [cytosol]│P60484 │
+ └──────────────┴──────────┘
+
+
+TIP: The _MATCH_ part of the previous query could be written in one line as follows:
+
+
+ //Equivalent to the previous one
+ MATCH (ewas:EntityWithAccessionedSequence{stId:"R-HSA-199420"})-[:referenceEntity]->(re:ReferenceEntity)
+ RETURN ewas.displayName AS EWAS, re.identifier AS Identifier
+
+
+Continuing on, it is possible to construct a query to retrieve the **reference database** on top of the previously retrieved fields. Please note that in this case the **reference database** is a **node** pointed from **ReferenceEntity** by an edge called **referenceDatabase** :
+
+
+ //Following the reference entity and database links in order to get the identifier and the database of reference
+ MATCH (ewas:EntityWithAccessionedSequence{stId:"R-HSA-199420"}),
+ (ewas)-[:referenceEntity]->(re:ReferenceEntity)-[:referenceDatabase]->(rd:ReferenceDatabase)
+ RETURN ewas.displayName AS EWAS, re.identifier AS Identifier, rd.displayName AS Database
+
+
+
+ ╒══════════════╤══════════╤════════╕
+ │EWAS │Identifier│Database│
+ ╞══════════════╪══════════╪════════╡
+ │PTEN [cytosol]│P60484 │UniProt │
+ └──────────────┴──────────┴────────┘
+
+
+### [Breaking down complexes and sets to get their participants](<#complexes-sets-participants>)
+
+The components of a complex, which are also physical entities, are stored in the "hasComponent" slot. Let's use the complex "**Ag-substrate:E3:E2:Ub** " with identifier **R-HSA-983126** as example in this case:
+
+
+ //First level components for the complex with stable identifier R-HSA-983126
+ MATCH (Complex{stId:"R-HSA-983126"})-[:hasComponent]->(pe:PhysicalEntity)
+ RETURN pe.stId AS component_stId, pe.displayName AS component
+
+
+The result of the query is
+
+
+ ╒══════════════╤═══════════════════════════════════════════════╕
+ │component_stId│component │
+ ╞══════════════╪═══════════════════════════════════════════════╡
+ │R-NUL-983035 │antigenic substrate [cytosol] │
+ ├──────────────┼───────────────────────────────────────────────┤
+ │R-HSA-976075 │E3 ligases in proteasomal degradation [cytosol]│
+ ├──────────────┼───────────────────────────────────────────────┤
+ │R-HSA-976165 │Ubiquitin:E2 conjugating enzymes [cytosol] │
+ └──────────────┴───────────────────────────────────────────────┘
+
+
+In this example, the "**E3 ligases in proteasomal degradation** " is a Set and "**E3 ligases in proteasomal degradation** " is a Complex. To further deconstruct the initial complex there are some minor changes which should be applied to the Cypher query. Sets can either be DefineSets, OpenSets or CandidateSets. The way to find out which other physical entities are part of them is "traversing" through the "**hasMember** " or "**hasCandidate** " slots. The following query will break down the initial complex into ALL its participants:
+
+
+ //All distinct components for the complex with stable identifier R-HSA-983126
+ MATCH (Complex{stId:"R-HSA-983126"})-[:hasComponent|hasMember|hasCandidate*]->(pe:PhysicalEntity)
+ RETURN DISTINCT pe.stId AS component_stId, pe.displayName AS component
+
+
+This query returns 284 entities for v63:
+
+
+ ╒════════════════╤════════════════════════╕
+ │ component_stId │ component │
+ ╞════════════════╪════════════════════════╡
+ │ R-HSA-141412 │ CDC20 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174242 │ ANAPC7 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174211 │ ANAPC5 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174052 │ CDC26 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174244 │ UBE2C [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174126 │ ANAPC11 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174156 │ CDC16 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174189 │ ANAPC1 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174100 │ UBE2E1 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174229 │ ANAPC2 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174168 │ ANAPC4 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174073 │ CDC27 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174137 │ CDC23 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174142 │ ANAPC10 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-174236 │ UBE2D1 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-976009 │ CBLB [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-976042 │ MKRN1 [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-939214 │ UBB(1-76) [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-939213 │ UBB(77-152) [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │ R-HSA-939239 │ UBC(533-608) [cytosol] │
+ ├────────────────┼────────────────────────┤
+ │... │... │
+ └────────────────┴────────────────────────┘
+
+
+### [Retrieving pathways, subpathways and superpathways](<#retrieving-pathways>)
+
+In this example, we focus on the pathway "**Class I MHC mediated antigen processing & presentation**" with identifier **R-HSA-983169**. To find out its subpathways, the slot to query is "**hasEvent** ":
+
+
+ //Direct subpathways for the pathway with stable identifier R-HSA-198933
+ MATCH (p:Pathway{stId:"R-HSA-983169"})-[:hasEvent]->(sp:Pathway)
+ RETURN p.stId AS Pathway, sp.stId AS SubPathway, sp.displayName as DisplayName
+
+
+The result for v63 returns 3 subpathways:
+
+
+ ╒════════════╤═════════════╤══════════════════════════════════════════════════════════════════════════╕
+ │Pathway │SubPathway │DisplayName │
+ ╞════════════╪═════════════╪══════════════════════════════════════════════════════════════════════════╡
+ │R-HSA-983169│R-HSA-983168 │Antigen processing: Ubiquitination & Proteasome degradation │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-1236975│Antigen processing-Cross presentation │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-983170 │Antigen Presentation: Folding, assembly and peptide loading of class I MHC│
+ └────────────┴─────────────┴──────────────────────────────────────────────────────────────────────────┘
+
+
+It is important to note that subpathways might contain other subpathways, so to get ALL the supathways of R-HSA-198933, the query is as follows:
+
+
+ //ALL subpathways for the pathway with stable identifier R-HSA-198933
+ MATCH (p:Pathway{stId:"R-HSA-983169"})-[:hasEvent*]->(sp:Pathway)
+ RETURN p.stId AS Pathway, sp.stId AS SubPathway, sp.displayName as DisplayName
+
+
+In this case the number of subpathways is increased to 7:
+
+
+ ╒════════════╤═════════════╤══════════════════════════════════════════════════════════════════════════╕
+ │Pathway │SubPathway │DisplayName │
+ ╞════════════╪═════════════╪══════════════════════════════════════════════════════════════════════════╡
+ │R-HSA-983169│R-HSA-983170 │Antigen Presentation: Folding, assembly and peptide loading of class I MHC│
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-1236975│Antigen processing-Cross presentation │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-1236978│Cross-presentation of soluble exogenous antigens (endosomes) │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-1236977│Endosomal/Vacuolar pathway │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-1236974│ER-Phagosome pathway │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-1236973│Cross-presentation of particulate exogenous antigens (phagosomes) │
+ ├────────────┼─────────────┼──────────────────────────────────────────────────────────────────────────┤
+ │R-HSA-983169│R-HSA-983168 │Antigen processing: Ubiquitination & Proteasome degradation │
+ └────────────┴─────────────┴──────────────────────────────────────────────────────────────────────────┘
+
+
+Following the same approach, retrieving the superpathway is as easy as changing the direction of the edge in the query:
+
+
+ //Direct superpathway for the pathway with stable identifier R-HSA-198933
+ MATCH (p:Pathway{stId:"R-HSA-983169"})<-[:hasEvent]-(sp:Pathway)
+ RETURN p.stId AS Pathway, sp.stId AS SuperPathway, sp.displayName as DisplayName
+
+
+It will then retrieve the only one pathway containing R-HSA-198933:
+
+
+ ╒════════════╤═════════════╤══════════════════════╕
+ │Pathway │SuperPathway │DisplayName │
+ ╞════════════╪═════════════╪══════════════════════╡
+ │R-HSA-983169│R-HSA-1280218│Adaptive Immune System│
+ └────────────┴─────────────┴──────────────────────┘
+
+
+As subpathways, the superpathways might have other superpathways, so following the "**hasEvent** " slot recursively will show ALL the superpathways up to the root:
+
+
+ //ALL superpathways for the pathway with stable identifier R-HSA-198933
+ MATCH (p:Pathway{stId:"R-HSA-983169"})<-[:hasEvent*]-(sp:Pathway)
+ RETURN p.stId AS Pathway, sp.stId AS SuperPathway, sp.displayName as DisplayName
+
+
+There are 2 superpathways for R-HSA-198933 in v63:
+
+
+ ╒════════════╤═════════════╤══════════════════════╕
+ │Pathway │SuperPathway │DisplayName │
+ ╞════════════╪═════════════╪══════════════════════╡
+ │R-HSA-983169│R-HSA-1280218│Adaptive Immune System│
+ ├────────────┼─────────────┼──────────────────────┤
+ │R-HSA-983169│R-HSA-168256 │Immune System │
+ └────────────┴─────────────┴──────────────────────┘
+
+
+### [Retrieving the reactions for a given pathway](<#retrieving-reactions>)
+
+Continuing with the pathway "**Class I MHC mediated antigen processing & presentation**" with identifier **R-HSA-983169** , to get ALL the reactions contained either directly in it or as part of any of its subpathways, the query has to recursively traverse the "**hasEvent** " slot:
+
+
+ //All reactions for the pathway with stable identifier R-HSA-198933
+ MATCH (p:Pathway{stId:"R-HSA-983169"})-[:hasEvent*]->(rle:ReactionLikeEvent)
+ RETURN p.stId AS Pathway, rle.stId AS Reaction, rle.displayName AS ReactionName
+
+
+As shown in the table below, this pathway contains 51 reactions for v63:
+
+
+ ╒══════════════╤═══════════════╤═══════════════════════════════════════════════════════════════════════════════════╕
+ │ Pathway │ Reaction │ ReactionName │
+ ╞══════════════╪═══════════════╪═══════════════════════════════════════════════════════════════════════════════════╡
+ │ R-HSA-983169 │ R-HSA-983148 │ Interaction of Erp57 with MHC class I HC │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-8951499 │ Loading of antigenic peptides on to class I MHC │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983146 │ Interaction of beta-2-microglobulin (B2M) chain with class I HC │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983145 │ Binding of newly synthesized MHC class I heavy chain (HC) with calnexin │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983144 │ Transport of Antigen peptide in to ER │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-203979 │ Coat Assembly │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983142 │ Formation of peptide loading complex (PLC) │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983427 │ Expression of peptide bound class I MHC on cell surface │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983138 │ Transport of MHC heterotrimer to ER exit site │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983426 │ Capturing cargo and formation of prebudding complex │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983425 │ Recruitment of Sec31p:Sec13p to prebudding complex and formation of COPII vesicle │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983424 │ Budding of COPII coated vesicle │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983422 │ Disassembly of COPII coated vesicle │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983421 │ Journey of cargo through Golgi complex │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-983169 │ R-HSA-983161 │ Dissociation of the Antigenic peptide:MHC:B2M peptide loading complex │
+ ├──────────────┼───────────────┼───────────────────────────────────────────────────────────────────────────────────┤
+ │... │... │... │
+ └──────────────┴───────────────┴───────────────────────────────────────────────────────────────────────────────────┘
+
+
+### [Retrieving the participants of a Reaction](<#retrieving-participants>)
+
+In this case let's use the reaction "**IKKB phosphorylates SNAP23** " with identifier **R-HSA-8863895**. Reactions have inputs, outputs, catalysts and regulations, so to know the participants of a reaction, all these slots have to be taken into account. Please note that the physical entity acting as catalyst is stored in the "physicalEntity" slot of the class "CatalystActivity" and the one belonging to the regulation is stored in the "regulator" slot of the "Regulation" class. So the query is as follows:
+
+
+ //First level paticipating molecules for reaction R-HSA-8863895
+ MATCH (r:ReactionLikeEvent{stId:"R-HSA-8863895"})-[:input|output|catalystActivity|physicalEntity|regulatedBy|regulator*]->(pe:PhysicalEntity)
+ RETURN DISTINCT r.stId AS Reaction, pe.stId as Participant, pe.displayName AS DisplayName
+
+
+The result of it is 6 physical entities, where two of them are complexes:
+
+
+ ╒═════════════╤═════════════╤═══════════════════════════════════════════════════╕
+ │Reaction │Participant │DisplayName │
+ ╞═════════════╪═════════════╪═══════════════════════════════════════════════════╡
+ │R-HSA-8863895│R-HSA-168113 │CHUK:IKBKB:IKBKG [cytosol] │
+ ├─────────────┼─────────────┼───────────────────────────────────────────────────┤
+ │R-HSA-8863895│R-ALL-113592 │ATP [cytosol] │
+ ├─────────────┼─────────────┼───────────────────────────────────────────────────┤
+ │R-HSA-8863895│R-HSA-8863966│SNAP23 [phagocytic vesicle membrane] │
+ ├─────────────┼─────────────┼───────────────────────────────────────────────────┤
+ │R-HSA-8863895│R-HSA-8863923│p-S95-SNAP23 [phagocytic vesicle membrane] │
+ ├─────────────┼─────────────┼───────────────────────────────────────────────────┤
+ │R-HSA-8863895│R-ALL-29370 │ADP [cytosol] │
+ ├─────────────┼─────────────┼───────────────────────────────────────────────────┤
+ │R-HSA-8863895│R-HSA-937033 │oligo-MyD88:Mal:BTK:activated TLR [plasma membrane]│
+ └─────────────┴─────────────┴───────────────────────────────────────────────────┘
+
+
+As shown above, to break down complexes and sets into their participants, the "hasComponent", "hasMember" and "hasCandidate" slots have to be taken into account. Adding them into the previous query will retrieve ALL the participants of the reaction:
+
+
+ //ALL paticipating molecules for reaction R-HSA-8863895
+ MATCH (r:ReactionLikeEvent{stId:"R-HSA-8863895"})-[:input|output|catalystActivity|physicalEntity|regulatedBy|regulator|hasComponent|hasMember|hasCandidate*]->(pe:PhysicalEntity)
+ RETURN DISTINCT r.stId AS Reaction, pe.stId as Participant, pe.displayName AS DisplayName
+
+
+With the modifications, the result of the query goes up to 43 physical entities in v63:
+
+
+ ╒═══════════════╤═══════════════╤══════════════════════════════════════════════════════════════════════════════╕
+ │ Reaction │ Participant │ DisplayName │
+ ╞═══════════════╪═══════════════╪══════════════════════════════════════════════════════════════════════════════╡
+ │ R-HSA-8863895 │ R-HSA-168113 │ CHUK:IKBKB:IKBKG [cytosol] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-168114 │ IKBKB [cytosol] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-168104 │ CHUK [cytosol] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-168108 │ IKBKG [cytosol] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-ALL-113592 │ ATP [cytosol] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-8863966 │ SNAP23 [phagocytic vesicle membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-8863923 │ p-S95-SNAP23 [phagocytic vesicle membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-ALL-29370 │ ADP [cytosol] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-937033 │ oligo-MyD88:Mal:BTK:activated TLR [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-937013 │ MyD88 oligomer [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-937017 │ MYD88 [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-2201325 │ activated TLR2/4:p-4Y-MAL:PI(4,5)P2:BTK [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-5365824 │ p-4Y-TIRAP:PI(4,5)P2 [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-ALL-179856 │ PI(4,5)P2 [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-2201321 │ p-4Y-TIRAP [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-181230 │ Activated TLR1:2 or TLR 2:6 heterodimers or TLR4 homodimer [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-181410 │ TLR6:TLR2:ligand:CD14:CD36 [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-2559461 │ TLR6/2 ligand:CD14:CD36 [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │ R-HSA-8863895 │ R-HSA-166033 │ GPIN-CD14(20-345) [plasma membrane] │
+ ├───────────────┼───────────────┼──────────────────────────────────────────────────────────────────────────────┤
+ │... │... │... │
+ └───────────────┴───────────────┴──────────────────────────────────────────────────────────────────────────────┘
+
+
+### [Joining the pieces: Participating molecules for a pathway](<#joining-pieces>)
+
+The aim of this tutorial was to describe a number of different queries to the Reactome Graph Database that joined together would retrieve the resource and identifier of each participating molecule for a given Pathway. This final example will demonstrate how to concatenate all the individual query described previously into a single query.
+
+Starting from the pathway "**Class I MHC mediated antigen processing & presentation**" with identifier **R-HSA-983169** , first we need to find out all the reactions contained in it. For each reaction we want to find their participants and for the cases of complexes and sets, we want to break them down into single physical entities. Finally, for each physical entity we are interested in their identifier and resource:
+
+
+ //ALL paticipating molecules for pathway R-HSA-983169
+ MATCH (p:Pathway{stId:"R-HSA-983169"})-[:hasEvent*]->(rle:ReactionLikeEvent),
+ (rle)-[:input|output|catalystActivity|physicalEntity|regulatedBy|regulator|hasComponent|hasMember|hasCandidate*]->(pe:PhysicalEntity),
+ (pe)-[:referenceEntity]->(re:ReferenceEntity)-[:referenceDatabase]->(rd:ReferenceDatabase)
+ RETURN DISTINCT re.identifier AS Identifier, rd.displayName AS Database
+
+
+For version 63, the pathway has 463 participating molecules as shown below:
+
+
+ ╒════════════╤══════════╕
+ │ Identifier │ Database │
+ ╞════════════╪══════════╡
+ │ P11021 │ UniProt │
+ ├────────────┼──────────┤
+ │ P27824 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30501 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30486 │ UniProt │
+ ├────────────┼──────────┤
+ │ P01893 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30447 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30685 │ UniProt │
+ ├────────────┼──────────┤
+ │ P18465 │ UniProt │
+ ├────────────┼──────────┤
+ │ P18464 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30460 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30490 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30495 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30493 │ UniProt │
+ ├────────────┼──────────┤
+ │ P13747 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30488 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30511 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30483 │ UniProt │
+ ├────────────┼──────────┤
+ │ P30462 │ UniProt │
+ ├────────────┼──────────┤
+ │ P04439 │ UniProt │
+ ├────────────┼──────────┤
+ │ ... │ ... │
+ └────────────┴──────────┘
+
+
+This concludes the introductory tutorial on how to build a Cypher query to retrieve all the participating molecules for a given pathway. For questions or suggestions, please get in touch our This email address is being protected from spambots. You need JavaScript enabled to view it.
diff --git a/projects/website-angular/content/documentation/dev/graph-database/neo4j-desktop.mdx b/projects/website-angular/content/documentation/dev/graph-database/neo4j-desktop.mdx
new file mode 100644
index 0000000..1ecc619
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/graph-database/neo4j-desktop.mdx
@@ -0,0 +1,76 @@
+---
+title: Untitled
+category: "documentation"
+---
+
+### Introduction
+
+This article is going to describe step by step how to import our great Graph Database dump file into Neo4j Desktop
+
+### Download
+
+ * [Neo4j Desktop]()
+ * [Graph Database Dump file]()
+
+### Neo4j Desktop
+
+Install Neo4j Desktop following the [official documentation website]().
+
+### Importing Graph Database
+
+ 1. Create new Project
+
+
+
+ 2. Add reactome.graphdb.dump
+
+
+
+ 3. Click on the Open button next to the file you have added and select "Create new DBMS from Dump".
+
+
+
+ 4. Choose a name, password and version (preferably >4.x.x)
+
+
+
+ 5. Not mandatory, open Neo4j Settings, edit "dbms.allow_upgrade=true
+ 1. This step is only needed if you are using Reactome Graph Database which is not compatible with v4.X.X.
+
+
+
+
+
+ 6. Now Click on `Start` and you are ready to explore the Neo4j Browser or Neo4j Bloom.
+
+
+
+### Adding a different version of the Reactome database
+
+* Only Neo4j >4.x supports multiple DMBS, make sure you installed Neo4j 4.x.x
+
+ 1. Add reactome.previous.graphdb.dump
+
+
+
+ 2. Click on the Open button next to the file you have added and select "Import dump into existing DBMS".
+ 3. Select Reactome and create database "reactome"
+
+
+
+ 1. Open Neo4j Settings
+
+
+
+ 2. Not mandatory, open Neo4j Settings, edit "dbms.allow_upgrade=true
+
+ 1. This step is only needed if you are using Reactome Graph Database which is not compatible with v4.X.X.
+
+
+
+ 3. If you want this database to be your default, then use its name in the settings file.
+
+
+
+ 4. Now Click on `Start` and you are ready to explore the Neo4j Browser or Neo4j Bloom.
+ 1.
diff --git a/projects/website-angular/content/documentation/dev/index.mdx b/projects/website-angular/content/documentation/dev/index.mdx
deleted file mode 100644
index b5765db..0000000
--- a/projects/website-angular/content/documentation/dev/index.mdx
+++ /dev/null
@@ -1,44 +0,0 @@
----
-title: "Developer's Zone"
-category: "documentation"
----
-
-## Developer's Zone
-
-#### Explore our tools and web services and learn how to include them in your applications
-
-[ __ ]()
-
-## [ Analysis Service ]()
-
-Use the Analysis Service to analyse your data against Reactome’s content
-
-[ __ ]()
-
-## [ Content Service ]()
-
-Use the Content Service to access all our knowledgebase content from your client
-
-[ __ ]()
-
-## [ Graph Database ]()
-
-Access to the Reactome knowledgebase content as an interconnected graph database
-
-[ __ ]()
-
-## [ Pathways Overview ]()
-
-Use this widget to include our pathways overview in your web application
-
-[ __ ]()
-
-## [ Pathway Diagrams ]()
-
-Use this widget to include our pathway diagrams in your web application
-
-[ __ ]()
-
-## [ Reactome Partners ]()
-
-Check out who is currently using Reactome web services and widgets
diff --git a/projects/website-angular/content/documentation/dev/pathways-overview/index.mdx b/projects/website-angular/content/documentation/dev/pathways-overview.mdx
similarity index 100%
rename from projects/website-angular/content/documentation/dev/pathways-overview/index.mdx
rename to projects/website-angular/content/documentation/dev/pathways-overview.mdx
diff --git a/projects/website-angular/content/documentation/dev/pathways-overview/gwt.mdx b/projects/website-angular/content/documentation/dev/pathways-overview/gwt.mdx
new file mode 100644
index 0000000..06a6d2f
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/pathways-overview/gwt.mdx
@@ -0,0 +1,190 @@
+---
+title: Pathways Overview GWT
+category: "documentation"
+---
+
+## Pathways Overview GWT
+
+Reactome’s Fireworks GWT widget is our original implementation of the pathways overview in GWT. It is meant to be reused by third party resources in order to display Reactome pathways overview directly in their web pages and enable users to interact with them.
+
+[ __ ](<#MoreInfo>)
+
+## [ More Info ](<#MoreInfo>)
+
+[ __ ](<#GetStarted>)
+
+## [ Get Started ](<#GetStarted>)
+
+[ __ ](<#API>)
+
+## [ API ](<#API>)
+
+[ __ ](<#Resources>)
+
+## [ Resources ](<#Resources>)
+
+### [__More Info](<#MoreInfo>)
+
+[GWT]() is a development toolkit for building and optimizing complex browser-based applications. Its goal is to enable productive development of high-performance web applications without the developer having to be an expert in browser quirks, XMLHttpRequest, and JavaScript. GWT is used by many products at Google, including AdWords, AdSense, Flights, Hotel Finder, Offers, Wallet, Blogger. It’s open source, completely free, and used by thousands of developers around the world.
+
+### [__Get Started](<#GetStarted>)
+
+We assume that you already have your [GWT project]() up and running configured with the [ MAVEN archetype](). So the first step is to add the EBI Nexus repository in your pom.xml file:
+
+
+
+ ...
+
+
+ nexus-ebi-repo
+ The EBI internal repository
+ http://www.ebi.ac.uk/Tools/maven/repos/content/groups/ebi-repo/
+
+ true
+
+
+ false
+
+
+
+
+ nexus-ebi-snapshot-repo
+ The EBI internal snapshot repository
+ http://www.ebi.ac.uk/Tools/maven/repos/content/groups/ebi-snapshots/
+
+ false
+
+
+ true
+
+
+
+
+
+Once the repository is in place, the next thing to do is adding the Reactome's pathways overview dependency:
+
+
+
+ ...
+
+ org.reactome.web
+ fireworks
+ 1.0.0
+
+
+
+
+Before writing the code to use the pathways overview, add the dependency in your **gwt.xml** file, so the compiler will find the pathways overview:
+
+
+
+
+
+And finally, assuming the widget is now going to be displayed by its own, the code in your **EntryPoint** class would look like:
+
+
+ /**
+ * For test purposes we assume the pathway overview loads as a single item in the web page.
+ *
+ * Entry point classes define onModuleLoad().
+ */
+ public class Main implements EntryPoint {
+
+ private final FirewowksViewer fireworks;
+
+ public Main() {
+ //We create the fireworks. Please note that different pathways overview cannot be loaded in the same instance
+ fireworks = FireowksFactory.createFireworksViewer();
+ }
+
+ @Override
+ public void onModuleLoad() {
+ Scheduler.get().scheduleDeferred(new Scheduler.ScheduledCommand() {
+ @Override
+ public void execute() {
+ FireworksFactory.CONSOLE_VERBOSE = true; //This is optional (for dev purposes)
+ FireworksFactory.EVENT_BUS_VERBOSE = true; //This is optional (for dev purposes)
+
+ //For this use case we add the firewoks as the main and only object of the webpage
+ RootLayoutPanel.get().add(fireworks);
+ }
+ });
+ }
+ }
+
+
+### [__API](<#API>)
+
+The available methods are defined in the **DiagramViewer** interface in the **org.reactome.web.diagram.client** package. These are:
+
+
+ //subscribe to this event to be notified when the user closes the analysis overlay
+ HandlerRegistration addAnalysisResetHandler(AnalysisResetHandler handler);
+
+ //Subscribe to this event to be notified if the browser does not support canvas, so diagrams can not be displayed
+ HandlerRegistration addCanvasNotSupportedHandler(CanvasNotSupportedHandler handler);
+
+ //Subscribe to this event to be notified when the overview is displayed in the viewport
+ HandlerRegistration addFireworksLoaded(FireworksLoadedHandler handler);
+
+ //Subscribe to this event to be notified when
+ HandlerRegistration addExpressionColumnChangedHandler(ExpressionColumnChangedHandler handler);
+
+ //Subscribe to this event to be notified when the user hovers over an node in the overview
+ HandlerRegistration addNodeHoverHandler(NodeHoverHandler handler);
+
+ //Subscribe to this event to be notified when the user leave the hovered node in the overview
+ HandlerRegistration addNodeHoverResetHandler(NodeHoverResetHandler handler);
+
+ //Subscribe to this event to be notified when the user opens a node in the overview
+ HandlerRegistration addNodeOpenedHandler(NodeOpenedHandler handler);
+
+ //Subscribe to this event to be notified when the user selects a node in the overview
+ HandlerRegistration addNodeSelectedHandler(NodeSelectedHandler handler);
+
+ //Subscribe to this event to be notified when the user unselects a node in the overview
+ HandlerRegistration addNodeSelectedResetHandler(NodeSelectedResetHandler handler);
+
+ //Subscribe to this event to be notified when the user changes the colour profile
+ HandlerRegistration addProfileChangedHandler(ProfileChangedHandler handler);
+
+ //Returns the selected node
+ Node getSelected();
+
+ //Highlights the node with the provided stable identifier
+ void highlightNode(String stableIdentifier);
+
+ //Highlights the node with the provided stable identifier
+ void highlightNode(Long dbIdentifier);
+
+ //Zooms-in to the node with the provided stable identifier
+ void openPathway(String stableIdentifier);
+
+ //Zooms-in to the node with the provided database identifier
+ void openPathway(Long dbIdentifier);
+
+ //Resets the analysis overlay
+ void resetAnalysis();
+
+ //Resets the highlighted nodes
+ void resetHighlight();
+
+ //Resets the selected nodes
+ void resetSelection();
+
+ //Selects the node with the provided stable identifier
+ void selectNode(String stableIdentifier);
+
+ //Selects the node with the provided database identifier
+ void selectNode(Long dbIdentifier);
+
+ //Sets the analysis overlay for a given "token" and "resource" (this will query the Analysis Service)
+ void setAnalysisToken(String token, String resource);
+
+ //Show all the nodes in the viewport
+ void showAll();
+
+### [__Resources](<#Resources>)
+
+ * [__GWT Fireworks code]()
+ * [__Reactome Pathways Portal code]()
diff --git a/projects/website-angular/content/documentation/dev/pathways-overview/js.mdx b/projects/website-angular/content/documentation/dev/pathways-overview/js.mdx
new file mode 100644
index 0000000..84afb68
--- /dev/null
+++ b/projects/website-angular/content/documentation/dev/pathways-overview/js.mdx
@@ -0,0 +1,105 @@
+---
+title: Pathways Overview JS
+category: "documentation"
+---
+
+## Pathways Overview JS
+
+Reactome's Fireworks JS widget is our pathways overview viewer in an ordinary JavaScript API. It is meant to be reused by third party resources in order to display Reactome pathways overview directly in their web pages and enable users to interact with them.
+
+### [__Reusing Reactome's Pathways Overview Widget?](<#reuse-pathways-widget>)
+
+To reuse our viewer you need to follow the following steps
+
+1\. Include the fireworks javascript dependency in you HTML header
+
+
+
+
+
+2\. Add a place holder in the body of your web page
+
+
+
+
+
+3\. Set a proxy in your server under "/reactome" pointing to "https://reactome.org"
+
+4\. Create and initialise the pathways overview from your javascript code
+
+
+ //Creating the Reactome pathways overview widget
+ function onReactomeFireworksReady(){ //This function is automatically called when the widget code is ready to be used
+ var fireworks = Reactome.Fireworks.create({
+ "placeHolder" : "fireworksHolder",
+ "width" : 930,
+ "height" : 500
+ });
+
+ //Adding different listeners
+
+ fireworks.onFireworksLoaded(function (loaded) {
+ console.info("Loaded ", loaded);
+ });
+
+ fireworks.onNodeHovered(function (hovered){
+ console.info("Hovered ", hovered);
+ });
+
+ fireworks.onNodeSelected(function (selected){
+ console.info("Selected ", selected);
+ });
+ }
+
+
+### [__FireworksJs API](<#API>)
+
+The current implementation supports the following listeners and methods:
+
+Method| Params| Description
+---|---|---
+**create** :: Constructor
+Reactome.Fireworks.create(params); | **param** :: json object
+{
+'proxyPrefix' : string,
+'placeHolder' : string,
+'width' : int (optional),
+'height' : int (optional)
+} | Creates and returns a new Reactome.Fireworks object
+**flagItems(String identifier)** :: void | Item identifier
+**identifier** : string | Flags pathways where the identifier is found. It accepts main identifiers but also cross references, gene names and physical entity stable identifiers
+**highlightNode(stId)** :: void | Item stable identifier
+**stId** : string | Highlights the specified item if it exists in the fireworks
+**resetAnalysis()** :: void | | Resets the analysis overlay
+**resetHighlight()** :: void | | Clears the highlights in the fireworks
+**resetSelection()** :: void | | Clears the selection in the fireworks
+**resize(width, height)** :: void | **width** : int
+**height** : int | Resizes the viewport to the specified with and height
+**selectNode(stId)** :: void | Item stable identifier
+**stId** : string | Selects the specified item if it exists in the fireworks
+**setAnalysisToken(token, resource)** :: void | Analysis token
+**token** : string
+Resource
+**resource** : string | Overlays the analysis result corresponding to the specified (token, resource)
+**showAll()** :: void | | Shows all the pathways in the viewport
+**onNodesFlaggedReset(function())** :: void | The function receives no parameters | The function is called when the user resets the flagged items
+**onNodeSelected(function(obj))** :: void | **obj** is selected item:
+{
+'stId' : string,
+'displayName' : string,
+'schemaClass' : string,
+} | The function is called when an object in the fireworks is selected by the user action
+**onNodeHovered(function(obj))** :: void | **obj** is hovered item:
+{
+'stId' : string,
+'displayName' : string,
+'schemaClass' : string,
+} | The function is called when an object in the fireworks is hovered by the user action
+**onFireworksLoaded(function(id))** :: void | The db identifier of the loaded species in fireworks
+**id** : string | The function is called when a fireworks is loaded in the viewer
+**onAnalysisReset(function())** :: void | The function receives no parameters | The function is called when the users resets the analysis overlay
+
+fireworks.onAnalysisReset(function(){ /* your code here */ });
+**onCanvasNotSupported(function())** :: void | The function receives no parameters. | The function is called when the browser doesn't support HTML5 Canvas so the viewer cannot be instantiated
+
+fireworks.onCanvasNotSupported(function(){ /* your code here */ });
diff --git a/projects/website-angular/content/documentation/faq/analysis/api-and-r/215-pathway-analysis-api.mdx b/projects/website-angular/content/documentation/faq/analysis/api-and-r/215-pathway-analysis-api.mdx
new file mode 100644
index 0000000..e6a597c
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/api-and-r/215-pathway-analysis-api.mdx
@@ -0,0 +1,8 @@
+---
+title: "Is there an API for Reactome? Can I do pathway analysis through the API?"
+category: "documentation"
+---
+
+## Is there an API for Reactome? Can I do pathway analysis through the API?
+
+Pathway analysis can be performed through our API, documented [here]().
diff --git a/projects/website-angular/content/documentation/faq/analysis/api-and-r/216-gene-symbol.mdx b/projects/website-angular/content/documentation/faq/analysis/api-and-r/216-gene-symbol.mdx
new file mode 100644
index 0000000..641b0d5
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/api-and-r/216-gene-symbol.mdx
@@ -0,0 +1,8 @@
+---
+title: "Should I use the gene symbol or the NCBI identifiers to make requests to the identifiers endpoint of the Analysis Service?"
+category: "documentation"
+---
+
+## Should I use the gene symbol or the NCBI identifiers to make requests to the identifiers endpoint of the Analysis Service?
+
+We suggest using the HGVS gene name or the Uniprot identifier in your analysis. Different results may be obtained using other identifiers (like NCBIs) if those identifiers also map to other resources, bringing in more pathways.
diff --git a/projects/website-angular/content/documentation/faq/analysis/api-and-r/217-pathway-analysis-r.mdx b/projects/website-angular/content/documentation/faq/analysis/api-and-r/217-pathway-analysis-r.mdx
new file mode 100644
index 0000000..e6a74ce
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/api-and-r/217-pathway-analysis-r.mdx
@@ -0,0 +1,8 @@
+---
+title: "Can I do pathway analysis using R?"
+category: "documentation"
+---
+
+## Can I do pathway analysis using R?
+
+Although we don't provide a specific R package suitable for visualizing expression values on Reactome pathways, we do provide a full BioConductor package for quantitative pathway analysis based on expression data. Please see [here]() for documentation, and [here]() for the relevant publication.
diff --git a/projects/website-angular/content/documentation/faq/analysis/fiviz/218-non-human-fiviz.mdx b/projects/website-angular/content/documentation/faq/analysis/fiviz/218-non-human-fiviz.mdx
new file mode 100644
index 0000000..97fdb9f
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/fiviz/218-non-human-fiviz.mdx
@@ -0,0 +1,8 @@
+---
+title: "I need to do a network analysis for (species) from RNA seq data. Can you please guide me on how to generate an FI network?"
+category: "documentation"
+---
+
+## I need to do a network analysis for (species) from RNA seq data. Can you please guide me on how to generate an FI network?
+
+Unfortunately our ReactomeFIViz tool currently supports networks for human and mouse only. For bacterial species, you can try [StringDB](), which provides software tools to build networks for your genes.
diff --git a/projects/website-angular/content/documentation/faq/analysis/fiviz/219-fiviz-differential-expression.mdx b/projects/website-angular/content/documentation/faq/analysis/fiviz/219-fiviz-differential-expression.mdx
new file mode 100644
index 0000000..19fa120
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/fiviz/219-fiviz-differential-expression.mdx
@@ -0,0 +1,8 @@
+---
+title: "Can I submit a list of differentially expressed genes, as log2 fold change values, coming from two different conditions to have a quantitative representation of pathways involvement?"
+category: "documentation"
+---
+
+## Can I submit a list of differentially expressed genes, as log2 fold change values, coming from two different conditions to have a quantitative representation of pathways involvement?
+
+If you want to use your log fold change data directly, you may use the Reactome Cytoscape app, described [here]() (search for “Perform GSEA analysis”). This feature will sort your genes based on log fold change and then calculate a score for each pathway using GSEA’s ranked gene list input.
diff --git a/projects/website-angular/content/documentation/faq/analysis/general/197-convert-to-human.mdx b/projects/website-angular/content/documentation/faq/analysis/general/197-convert-to-human.mdx
new file mode 100644
index 0000000..5c55981
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/general/197-convert-to-human.mdx
@@ -0,0 +1,14 @@
+---
+title: "I used the “Analyse gene expression” tool with my mouse proteomics data set. I notice that the database is based on human pathways – does the software automatically convert my mouse gene names to the human orthologs? Or can’t I use the software for mouse?"
+category: "documentation"
+---
+
+## I used the “Analyse gene expression” tool with my mouse proteomics data set. I notice that the database is based on human pathways – does the software automatically convert my mouse gene names to the human orthologs? Or can’t I use the software for mouse?
+
+Reactome’s manual curation covers human proteins, but we do support analysis of non-human data sets.
+
+There are two methods to analyze data in Reactome.
+
+The “Analyse gene list” tool, accessed after “Analysis” is selected from the home page, allows users to upload human or non-human data sets for overrepresentation analysis. After uploading the data with this tool, the user is given the option of ‘projecting to human’ (which is selected by default). If this toggle is selected, non-human identifiers in the data set are converted to their human equivalents using orthology information from Panther.
+
+Data sets can also be analyzed with the Reactome Gene Set Analysis (Reactome GSA) tool, accessed through the “Analyze Gene Expression” button after “Analysis” is selected from the home page. Reactome GSA performs quantitative pathway analyses, increasing the statistical power of the differential gene expression analysis. The analysis software automatically converts the mouse proteins from your proteomics set to their human orthologs.__
diff --git a/projects/website-angular/content/documentation/faq/analysis/general/207-statistical-analysis.mdx b/projects/website-angular/content/documentation/faq/analysis/general/207-statistical-analysis.mdx
new file mode 100644
index 0000000..70fb626
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/general/207-statistical-analysis.mdx
@@ -0,0 +1,13 @@
+---
+title: "The analysis report includes lists of significantly up- and down-regulated single genes. Which statistical analysis does the software use for these values?"
+category: "documentation"
+---
+
+## The analysis report includes lists of significantly up- and down-regulated single genes. Which statistical analysis does the software use for these values?
+
+As described in our [on-line documentation](), the analysis returns:
+
+ * Entities p-value: the result of the statistical binomial test for over-representation for molecules of the results type selected.
+ * Entities FDR: False discovery rate. Corrected over-representation probability.
+
+Note that if performing an analysis with non-human data, the statistics may be skewed due to changes in the sizes of gene families between human and the species associated with the submitted data.
diff --git a/projects/website-angular/content/documentation/faq/analysis/general/208-visualizing-gene-expression.mdx b/projects/website-angular/content/documentation/faq/analysis/general/208-visualizing-gene-expression.mdx
new file mode 100644
index 0000000..17b9c29
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/general/208-visualizing-gene-expression.mdx
@@ -0,0 +1,22 @@
+---
+title: "Can I visualize gene expression data (color nodes) according to gene expression level with Reactome?"
+category: "documentation"
+---
+
+## Can I visualize gene expression data (color nodes) according to gene expression level with Reactome?
+
+From the [home page](), click the “Analysis Tools” button, and then select “Analyse gene list” (preselected on the analysis page).
+
+Submit your list of differentially expressed genes, or try one of the sample data sets.
+
+The leftmost column of the uploaded data set must have gene names or other gene/protein identifiers. The remainder of the data set consists of as many columns with numerical values as necessary. Columns may have headers, or not.
+
+Click “Continue” and then “Analyse”, maintaining the options in the second window (‘Project to human’, ‘Include interactors’) at their default values.
+
+Reactome will perform a standard gene set enrichment analysis, based only on the gene list. In the results overview, pathways will be greyed out if they are not significantly enriched. If a pathway is enriched, its colour will be determined as follows:
+
+The top end of the colour map is the highest value of all submitted numerical values. The bottom end of the colour map is the lowest of all submitted numerical values, across all columns.
+
+The colour of a pathway is based on the average expression value for all genes which are in the submitted dataset and in the pathway. In the initial view, this average is based on the first column. If the data set has more than one column, users can cycle through the subsequent columns with the “Play” button at the bottom of the pathway window.
+
+In the pathways overview (either the "Fireworks" or the "Reacfoam" view, selectable in the top left area of the main window), the user can double click (long click in Reacfoam to select a pathway and zoom in to the detailed molecular map view. In this view, proteins are coloured according to the user-provided values, and again the user can cycle through multiple columns with the "Play" controls.
diff --git a/projects/website-angular/content/documentation/faq/analysis/general/209-expanded-event-hierarchy.mdx b/projects/website-angular/content/documentation/faq/analysis/general/209-expanded-event-hierarchy.mdx
new file mode 100644
index 0000000..8e571c1
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/general/209-expanded-event-hierarchy.mdx
@@ -0,0 +1,12 @@
+---
+title: "After submitting a list of genes for Reactome Analysis, how do I download the fully expanded 'Event Hierarchy' list on the left side of the screen?"
+category: "documentation"
+---
+
+## After submitting a list of genes for Reactome Analysis, how do I download the fully expanded 'Event Hierarchy' list on the left side of the screen?
+
+We provide an [endpoint]() to retrieve the full event hierarchy for a given species. For example, you can get human data with a token like the one below. Please replace “your_analysis_token” in the URL below with your own token.
+
+__
+
+[__]()_https://reactome.org/ContentService/data/eventsHierarchy/9606?pathwaysOnly=false &token=__your_analysis_token_ _& resource=TOTAL&interactors=false_
diff --git a/projects/website-angular/content/documentation/faq/analysis/general/210-query-genes-per-pathway.mdx b/projects/website-angular/content/documentation/faq/analysis/general/210-query-genes-per-pathway.mdx
new file mode 100644
index 0000000..dcebdb9
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/general/210-query-genes-per-pathway.mdx
@@ -0,0 +1,15 @@
+---
+title: "Is it possible to know how many of my uploaded genes are found within each specific hierarchical pathway category identified in the Reacfoam view? For instance, how many of my genes belong to pathways that would be included in the \"Immune\" category, etc."
+category: "documentation"
+---
+
+## Is it possible to know how many of my uploaded genes are found within each specific hierarchical pathway category identified in the Reacfoam view? For instance, how many of my genes belong to pathways that would be included in the "Immune" category, etc.
+
+At the moment we don’t have a way to filter the results by top level pathway, or diagram-level pathway or other criteria. This feature is on our radar to implement.
+
+In the meantime, you can get a sense of this as follows:
+
+ * Click on the ‘Analysis’ tab in the Details panel (below the pathway diagram window- the Analysis tab should be open by default after performing analysis).
+ * To the left of the Details panel, click on the ‘Download’ button.
+ * Click on the “Pathway analysis results”. This gives the analysis results for all pathways, and for higher-order pathways the columns "Entities found" and "Entities total" contain the aggregated results from the sub-pathways.
+ * Import the results into a spreadsheet. Although we don't have a column "Pathway level" or similar, which would allow you to filter for only top level pathways, if you sort the pathways by decreasing "Entities found", you get a good view of the top level, and thus typically (but not always) largest pathways.
diff --git a/projects/website-angular/content/documentation/faq/analysis/general/211-permanent-analysis-results.mdx b/projects/website-angular/content/documentation/faq/analysis/general/211-permanent-analysis-results.mdx
new file mode 100644
index 0000000..fddad83
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/general/211-permanent-analysis-results.mdx
@@ -0,0 +1,8 @@
+---
+title: "Is there a permanent link available for my analysis results?"
+category: "documentation"
+---
+
+## Is there a permanent link available for my analysis results?
+
+Unfortunately, there is no permanent link for analysis results. Analysis data is available through the token for 7 days after your last usage. Analysis results are deleted when Reactome releases new data, regardless of when you performed your analysis or last accessed your data.
diff --git a/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/212-gsa-training-material.mdx b/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/212-gsa-training-material.mdx
new file mode 100644
index 0000000..b99dfa8
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/212-gsa-training-material.mdx
@@ -0,0 +1,12 @@
+---
+title: "I have a proteomics data set that I would like to use for quantitative pathway analysis. Is there training material available to help me?"
+category: "documentation"
+---
+
+## I have a proteomics data set that I would like to use for quantitative pathway analysis. Is there training material available to help me?
+
+To process quantitative proteomics data, the best, but also most complex analysis option is ReactomeGSA. A description of the tool, its use, and documentation are all part of [this publication]()
+
+There is also a comprehensive training video [here]().
+
+If after going through these resources you have additional questions, please reach out to the help desk at This email address is being protected from spambots. You need JavaScript enabled to view it.
diff --git a/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/213-differentially-expressed-genes.mdx b/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/213-differentially-expressed-genes.mdx
new file mode 100644
index 0000000..b67f1d4
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/213-differentially-expressed-genes.mdx
@@ -0,0 +1,10 @@
+---
+title: "Can I submit a list of differentially expressed genes, as log2 fold change values, coming from two different conditions to have a quantitative representation of pathways involvement? If yes, how can I prepare and submit my file?"
+category: "documentation"
+---
+
+## Can I submit a list of differentially expressed genes, as log2 fold change values, coming from two different conditions to have a quantitative representation of pathways involvement? If yes, how can I prepare and submit my file?
+
+For this, please use the Reactome Gene Set Analysis tool, accessed through the “Analyze Gene Expression” button after “Analysis” is selected from the home page. The detailed analysis approaches underlying this feature are described [here]() _._
+
+The Reactome GSA tool is an enhancement of [GSEA]() and can take your original expression data directly, perform differential expression analysis and then highlight pathways according scores.
diff --git a/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/214-truncated-gsa-analysis-output.mdx b/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/214-truncated-gsa-analysis-output.mdx
new file mode 100644
index 0000000..3ca561c
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/analysis/reactome-gsa/214-truncated-gsa-analysis-output.mdx
@@ -0,0 +1,8 @@
+---
+title: "I submitted a list for expression analysis using the PADOG or CAMERA tools and the output Excel file with the statistics is missing a lot of significant genes. What is the explanation for this?"
+category: "documentation"
+---
+
+## I submitted a list for expression analysis using the PADOG or CAMERA tools and the output Excel file with the statistics is missing a lot of significant genes. What is the explanation for this?
+
+Although there is no limit on the number of genes that can be returned, the analysis is restricted to those genes that are present in Reactome. Since Reactome is a manually curated resource, it does not cover all human genes.
diff --git a/projects/website-angular/content/documentation/faq/general-website/195-no-results.mdx b/projects/website-angular/content/documentation/faq/general-website/195-no-results.mdx
new file mode 100644
index 0000000..7a768ba
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/195-no-results.mdx
@@ -0,0 +1,25 @@
+---
+title: I searched for (...) and didn’t get any results.
+category: "documentation"
+---
+
+## I searched for (...) and didn’t get any results.
+
+Your search may be correct as Reactome does not yet have complete coverage of human biology. But before concluding that there is no content relevant to your query, please check that your search is formatted appropriately:
+
+The search bar on Reactome’s homepage takes a variety of inputs including but not limited to: HGNC gene names, protein names, identifiers from Uniprot, ChEBI or other resources, and simple word or phrase searches (“glucose”; “signaling by ERBB2”; “Li Fraumeni syndrome”).
+
+Multiple search terms separated by a space may be entered in a single query; the results will be the total hits generated by each of the terms searched independently. Use of other punctuation (slashes, commas) may not yield full results and are better avoided.
+
+To search for an exact match, enclose your search term(s) in quotation marks.
+
+[Boolean operators]() (AND, OR, NOT) may also be used in formatting a search.
+
+Wild cards may be used to expand your search:
+
+ * ? represents one character e.g. A1?? Matches both A1CF and A1BG
+ * * represents n characters e.g. *A1* matches A1CF, A1BG and A1A4E9; also VWA1, ATP1A1
+
+When entering a database identifier, more accurate results will be generated if the search term is formatted using the syntax database:id (for instance, uniprot:P60484). For complex database names like Guide to Pharmacology, replace spaces with dashes (Guide-to-Pharmacology:7382)
+
+Hits of a successful search indicate that the search term is included somewhere in the identified record. This may be identification of a physical entity or event that directly involves the search term, but the search term may also be used in an event summary or as part of an associated literature reference, for example.
diff --git a/projects/website-angular/content/documentation/faq/general-website/199-non-human-species.mdx b/projects/website-angular/content/documentation/faq/general-website/199-non-human-species.mdx
new file mode 100644
index 0000000..ce4f827
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/199-non-human-species.mdx
@@ -0,0 +1,12 @@
+---
+title: "Does Reactome contain pathway information from non-human species?"
+category: "documentation"
+---
+
+## Does Reactome contain pathway information from non-human species?
+
+Reactome is centered on the molecular functions of human proteins. When possible, we annotate these functions with published evidence from work with human systems. When such evidence isn’t available, but the function is known to be well conserved across species and experimental evidence exists for a non-human species, we annotate the reaction for the protein in that species and manually infer the reaction involving the homologous human protein.
+
+We also computationally infer reactions for a small group of model organisms from our manually annotated human data. This group of organisms is centered on organisms of interest to the Alliance for Genome Resources. The process for these computationally inferred events is [here]().
+
+In a collaborative project, the Reactome data model has been adopted by [Plant Reactome]() to capture pathway information for diverse species of plants, including crop plants.
diff --git a/projects/website-angular/content/documentation/faq/general-website/201-identifiers.mdx b/projects/website-angular/content/documentation/faq/general-website/201-identifiers.mdx
new file mode 100644
index 0000000..d22a9d5
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/201-identifiers.mdx
@@ -0,0 +1,14 @@
+---
+title: Explain the identifiers associated with entities and events.
+category: "documentation"
+---
+
+## Explain the identifiers associated with entities and events.
+
+Each Reactome instance of any type gets a unique internal database identifier (DB_ID) when it is created that persists unchanged for the life of that instance and is not reused if that instance is deleted.
+
+The stable IDs visible on our website and in our downloads are generated only for instances that are physical entities (chemicals, genome-encoded entities, complexes and sets of these) and events (reactions and pathways). Each stable ID takes the following form: R (Reactome) - three-letter code for the species of the instance- DB_ID.version (to indicate its version if the instance has been revised since its creation). So human entities and events have stable ids with R-HSA (for Homo sapiens)-########.##, stable ids for Caenorhabditis elegans are R-CEL-########.##, etc. For simple chemicals, ALL is used as the species code (R-ALL-########.##). Like DB_IDs, stable identifiers persist for the lifetime of the instance, unchanged except for versioning, and are not reused if the instance is deleted.
+
+For manually annotated events, the species code is that of the species in which the event occurs. Barring curation errors, that species code always corresponds to the species listed in the details panel of the web page for the entity or event.
+
+For computationally inferred events, the species code is that of the model organism species and the DB identifier is the one assigned to the human event that is the basis of the inference. This is to reflect the fact that these inferences are intrinsically UNstable (because as the model organism genome build changes, the inferences based on sequence similarity to that model organism will also change).
diff --git a/projects/website-angular/content/documentation/faq/general-website/202-earlier-versions.mdx b/projects/website-angular/content/documentation/faq/general-website/202-earlier-versions.mdx
new file mode 100644
index 0000000..7e175f1
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/202-earlier-versions.mdx
@@ -0,0 +1,14 @@
+---
+title: "Where can I find earlier versions of the Reactome database?"
+category: "documentation"
+---
+
+## Where can I find earlier versions of the Reactome database?
+
+We are working at making earlier versions of the Reactome database publicly available. In the meantime, MySQL copies of earlier database releases are available through the general URL structure:
+
+https://download.reactome.org/XX/databases/gk_current.sql.gz
+
+where XX is the version code for the release (ie 80 for Version 80, released 3/2022). See [here]() for a list of Reactome version release dates. Note that version 60, 65 and 70 onward are available for download.
+
+Note that we are currently transitioning from a MySQL to Neo4J database. The current release of the database is available in both formats [here]().
diff --git a/projects/website-angular/content/documentation/faq/general-website/203-pathways-per-organ.mdx b/projects/website-angular/content/documentation/faq/general-website/203-pathways-per-organ.mdx
new file mode 100644
index 0000000..c2bd469
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/203-pathways-per-organ.mdx
@@ -0,0 +1,14 @@
+---
+title: "How can I find all pathways reported for one organ? For example, I want to extract all reported pathways in (tissue X) for (species Y)."
+category: "documentation"
+---
+
+## How can I find all pathways reported for one organ? For example, I want to extract all reported pathways in (tissue X) for (species Y).
+
+Reactome does not provide organ-specific annotation, we aim to annotate a generic (human) cell, and users can then use expression data overlay for their organ (cell, tissue) of interest through our [analysis tool]() (select "Microarray data" for an example).
+
+You can also use the "Tissue Distribution" tool to see how data from the [Human Protein Atlas]() maps to Reactome pathway space.
+
+Documentation is at []()[https://reactome.org/userguide/analysis]()
+
+Please keep in mind that Reactome curation focuses on human, and while you can switch to one of the other species supported by our inference protocol through the drop down in the upper left area of the pathway browser, this data is inferred from human by orthology.
diff --git a/projects/website-angular/content/documentation/faq/general-website/204-kegg-to-reactome.mdx b/projects/website-angular/content/documentation/faq/general-website/204-kegg-to-reactome.mdx
new file mode 100644
index 0000000..5b24bba
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/204-kegg-to-reactome.mdx
@@ -0,0 +1,12 @@
+---
+title: "Is there a mapping file between KEGG and Reactome pathways?"
+category: "documentation"
+---
+
+## Is there a mapping file between KEGG and Reactome pathways?
+
+Reactome does not have a mapping file to KEGG. The best reference for such a comparison is [ComPath]() (reference [here]()), however this resource may not be up to date.
+
+As an alternate approach, users can try the method used to establish links from models in the BioModels database to Reactome. For each model, we took its gene set, ran a gene set enrichment analysis against Reactome using the Reactome API documented [here](), and then added links to the Reactome pathways below a certain p value cutoff.
+
+You could use that method to identify Reactome pathways that are related to a given KEGG pathway, and consider pathways with no match below your p value cutoff as not matched. The Reactome analysis interface is fast, each query should return in a few seconds, so this is perfectly feasible for all of KEGG. We'd be happy to communicate further on this.
diff --git a/projects/website-angular/content/documentation/faq/general-website/205-inferred-pathways-download.mdx b/projects/website-angular/content/documentation/faq/general-website/205-inferred-pathways-download.mdx
new file mode 100644
index 0000000..7b8740e
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/general-website/205-inferred-pathways-download.mdx
@@ -0,0 +1,8 @@
+---
+title: "Is it possible to download the inferred pathways for mouse (or another species), similar to the GMT file for human pathways on the downloads page?"
+category: "documentation"
+---
+
+## Is it possible to download the inferred pathways for mouse (or another species), similar to the GMT file for human pathways on the downloads page?
+
+Unfortunately, this is not possible at this time.
diff --git a/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/198-install-neo4j.mdx b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/198-install-neo4j.mdx
new file mode 100644
index 0000000..d819043
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/198-install-neo4j.mdx
@@ -0,0 +1,14 @@
+---
+title: I am having trouble installing a local Reactome Neo4J server.
+category: "documentation"
+---
+
+## I am having trouble installing a local Reactome Neo4J server.
+
+Unfortunately, installation of a local Neo4J server can be complicated due to variations in individual setup (computer, operating system, version of Neo4J and so forth).
+
+Please follow this [guide]() to install the graph database locally, there are different ways to install the graph database, and we also provide graph tar file and dump file to meet different requirements.
+
+As a long-term solution to this issue, we are working on providing the Neo4J databases in Docker containers. When these are ready for deployment, they will be added to the Reactome Download page.
+
+In the meantime, if you are getting an error message that the database is unavailable, please feel free to contact our developers through the help email ([]()This email address is being protected from spambots. You need JavaScript enabled to view it.) and we will try to assist you.
diff --git a/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/221-cypher-gene-list-to-pathways.mdx b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/221-cypher-gene-list-to-pathways.mdx
new file mode 100644
index 0000000..3b65904
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/221-cypher-gene-list-to-pathways.mdx
@@ -0,0 +1,26 @@
+---
+title: "I have a list of input genes. I want to retrieve all the pathways in which the genes are involved. How can I do this using a Neo4j Cypher query?"
+category: "documentation"
+---
+
+## I have a list of input genes. I want to retrieve all the pathways in which the genes are involved. How can I do this using a Neo4j Cypher query?
+
+There is a [tutorial]() on using neo4j Cypher query here.
+
+To retrieve all human pathways for a single gene (for example: [P36897](), the Uniprot identifier for TGFR1)
+
+
+
+ MATCH (n)-[:referenceDatabase]->(rd:ReferenceDatabase)
+ WHERE toLower(rd.displayName) = toLower("UniProt") AND (n.identifier = "P36897" OR n.variantIdentifier ="P36897" OR "P36897" IN n.geneName OR "P36897" IN n.name)
+ WITH DISTINCT n
+ MATCH (pe:PhysicalEntity)-[:referenceEntity|referenceSequence|crossReference|referenceGene*]->(n)
+ WITH DISTINCT pe
+ MATCH (rle:ReactionLikeEvent)-[:input|output|catalystActivity|physicalEntity|entityFunctionalStatus|diseaseEntity|regulatedBy|regulator|hasComponent|hasMember|hasCandidate|repeatedUnit*]->(pe)
+ WITH DISTINCT rle
+ MATCH (:Species{taxId:"9606"})<-[:species]-(p:Pathway)-[:hasEvent]->(rle)
+ RETURN DISTINCT p
+ ORDER BY p.stId
+
+
+If you have a list of genes, we suggest using [UNWIND]() neo4j cypher to flatten your list back to individual items and then execute the query.
diff --git a/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/222-disease-to-pathways-api-or-neo4j.mdx b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/222-disease-to-pathways-api-or-neo4j.mdx
new file mode 100644
index 0000000..811e8e2
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/222-disease-to-pathways-api-or-neo4j.mdx
@@ -0,0 +1,57 @@
+---
+title: "I have a list of Reactome pathway IDs. I want to find whether any of them are disease pathways and also retrieve associated disease names. Is it possible to retrieve the information using API or neo4j graph database query?"
+category: "documentation"
+---
+
+## I have a list of Reactome pathway IDs. I want to find whether any of them are disease pathways and also retrieve associated disease names. Is it possible to retrieve the information using API or neo4j graph database query?
+
+To do this, you can do a POST query with your pathway identifiers to the endpoint described [here](<#/query/findByIds>):
+
+To know if a Pathway is a disease, look at the “isInDisease” property. The disease name is contained in the displayName of the diseases in the “disease” array of the Pathway models.
+
+One query example would be
+
+
+
+ curl -X 'POST' \
+ 'https://reactome.org/ContentService/data/query/ids' \
+ -H 'accept: */*' \
+ -H 'Content-Type: text/plain' \
+ -d 'R-HSA-9679506,R-HSA-8876384' > pathwaysDescription.json
+
+
+To do this for multiple ids using cypher, you can use this query:
+
+
+
+ MATCH (p:Pathway)
+ WHERE p.stId in ["R-HSA-9679506","R-HSA-8876384"]
+ OPTIONAL MATCH (p)-[:disease]->(d:Disease)
+ RETURN p,d
+
+
+Using the optional match allows you to get both pathways associated with a disease and those which are not.
+
+If you want to filter for only disease pathways, use the following:
+
+
+
+ MATCH (p:Pathway)-[:disease]->(d:Disease)
+ WHERE p.stId in ["R-HSA-9679506","R-HSA-8876384"]
+ RETURN p,d
+
+
+You just need to provide your list of stId (reactome identifiers) in the “where” list.
+
+Finally, if you want to export a table instead of viewing it in the browser, use the following:
+
+
+
+ MATCH (p:Pathway)
+ WHERE p.stId in ["R-HSA-9679506","R-HSA-8876384"]
+ OPTIONAL MATCH (p)-[:disease]->(d:Disease)
+ WITH p, collect(d) as diseases
+ RETURN p.stId, p.displayName, p.isInDisease, [d in diseases | d.displayName] as diseaseNames, [d in diseases | d.databaseName + ":" + d.identifier] as diseaseIdentifiers
+
+
+Note that some pathways, like R-HSA-9608290 are associated with several diseases.
diff --git a/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/223-neo4j-all-genes-for-a-pathway.mdx b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/223-neo4j-all-genes-for-a-pathway.mdx
new file mode 100644
index 0000000..2d66681
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/223-neo4j-all-genes-for-a-pathway.mdx
@@ -0,0 +1,16 @@
+---
+title: "How can I use Neo4J to identify all the genes for a given pathway?"
+category: "documentation"
+---
+
+## How can I use Neo4J to identify all the genes for a given pathway?
+
+The query for collecting all genes for a given pathway is
+
+
+
+ MATCH (n:DatabaseObject{stId:"R-HSA-3371599"})-[:hasEvent|input|output|catalystActivity|physicalEntity|entityFunctionalStatus|diseaseEntity|regulatedBy|regulator|hasComponent|hasMember|hasCandidate|repeatedUnit|referenceEntity*]->(m:ReferenceGeneProduct)
+ RETURN DISTINCT m
+
+
+For guidance on formatting other queries, please take a look at our [data schema](). Our graph core projects also include most of the queries used in Reactome; you can find lots of examples [here]().
diff --git a/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/224-ppi-to-pathways-graph.mdx b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/224-ppi-to-pathways-graph.mdx
new file mode 100644
index 0000000..db8a5af
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/graph-database-and-cypher-query/224-ppi-to-pathways-graph.mdx
@@ -0,0 +1,16 @@
+---
+title: "Is it possible to directly connect my protein protein interaction network with the Reactome database or do I have to extract the required data and separately build the graph?"
+category: "documentation"
+---
+
+## Is it possible to directly connect my protein protein interaction network with the Reactome database or do I have to extract the required data and separately build the graph?
+
+Yes, you can directly query in our graph database, for example:
+
+
+
+ MATCH (ref1:ReferenceEntity)<-[:referenceEntity]-(p1:EntityWithAccessionedSequence)<-[:input|output|catalystActivity|physicalEntity|entityFunctionalStatus|diseaseEntity|regulatedBy|regulator|hasComponent|hasMember|hasCandidate|repeatedUnit*]-(reaction:ReactionLikeEvent)-[:input|output|catalystActivity|physicalEntity|entityFunctionalStatus|diseaseEntity|regulatedBy|regulator|hasComponent|hasMember|hasCandidate|repeatedUnit*]->(p2:EntityWithAccessionedSequence)-[:referenceEntity]->(ref2:ReferenceEntity)
+ WHERE ref1.databaseName = 'UniProt' AND ref2.databaseName = 'UniProt' AND ref1.identifier = 'P60484’ AND ref2.identifier = 'P42685'
+ MATCH eventPath=(reaction)<-[:hasEvent*]-(pathway)
+ UNWIND (nodes(eventPath)) as p
+ RETURN DISTINCT p.stId
diff --git a/projects/website-angular/content/documentation/faq/illustrations-figures/38-illustrations-figures.mdx b/projects/website-angular/content/documentation/faq/illustrations-figures/38-illustrations-figures.mdx
new file mode 100644
index 0000000..521a98c
--- /dev/null
+++ b/projects/website-angular/content/documentation/faq/illustrations-figures/38-illustrations-figures.mdx
@@ -0,0 +1,10 @@
+---
+title: "Is there a way to download high level images which can be zoomed in locally (after download)?"
+category: "documentation"
+---
+
+## Is there a way to download high level images which can be zoomed in locally (after download)?
+
+If you would like to download icons and Enhanced High Level Diagrams(EHLD), yes, we provide SVG format for both. Please visit our icon library [here](), you can download any EHLDs in pathway browser by clicking the download button at the top right corner to export diagram to different formats including SVG.
+
+However, if you would like to download a high level image of Reacfoam, unfortunately, there is no way to export it for now, basically, it takes a screenshot of the current window when you download a Reacfoam image.
diff --git a/projects/website-angular/content/documentation/icon-info.mdx b/projects/website-angular/content/documentation/icon-info.mdx
index bc70e9b..92e9483 100644
--- a/projects/website-angular/content/documentation/icon-info.mdx
+++ b/projects/website-angular/content/documentation/icon-info.mdx
@@ -1,6 +1,6 @@
---
title: "EHLD Specs & Guidelines"
-category: ""
+category: "documentation"
---
[ __ ]()
diff --git a/projects/website-angular/content/documentation/icon-info/ehld-specs-guideline.mdx b/projects/website-angular/content/documentation/icon-info/ehld-specs-guideline.mdx
new file mode 100644
index 0000000..910adc7
--- /dev/null
+++ b/projects/website-angular/content/documentation/icon-info/ehld-specs-guideline.mdx
@@ -0,0 +1,88 @@
+---
+title: "EHLD Specs & Guidelines"
+category: "documentation"
+---
+
+## EHLD Specs & Guidelines
+
+The Enhanced High Level Diagrams (EHLD) project aims to improve the graphical representation of higher-level pathways in the Reactome events hierarchy, e.g., “signal transduction”, “apoptosis”, or “metabolism” whose pathway diagrams consist of green boxes labeled with the names of sub-events, optionally located in cellular compartments and connected by arrows. These green-box diagrams feature limited navigation: clicking on a green box takes the user to that sub-event. In some cases, an illustrator has also created a graphic representation of the event. However, these illustrator diagrams offer no navigation functionality.
+
+There was a general agreement that green-box diagrams are not that appealing, and put off users accustomed to textbook-quality illustrations of biological processes with striking, intuitively clear iconography. Meanwhile, while our illustrations often are up to textbook standards, they are static images. The project included generation of scalable vector graphic (SVG) versions of illustrations, and the development of software that will make these SVG images navigable in the way that the green-box figures are now, and also to provide user interactivity by progressive zoom in/out, actions when hovering over the items or selecting them and showing the associated content in the details panel for the selected items.
+
+This project has also driven the development of a consistent iconography, a controlled visual vocabulary for biological processes like the controlled word vocabularies we already have for names of physical entities and reactions.
+
+This document includes two main sections. The first section describes in detail the process that should be followed to create an EHLD. The second section of this document provides guide through the process of creating and including new graphic elements into the Reactome EHLD icons library.
+
+### Enhanced High Level Diagrams guidelines
+
+This section aims to guide you through the process of creating an EHLD. The following paragraphs aim to summarise the set of requirements and guidelines the illustrator should follow in order to produce EHLDs that are compatible with Reactome Pathway Browser v3.4.
+
+#### Generic guidelines
+
+In Reactome, an EHLD is an interactive graphical representation of a higher level pathway diagram, typically containing two or more subpathways as active regions. An active region refers to a group of shapes representing a single subpathway. In particular, users can interact with active regions by hovering or selecting them, as well as use them to navigate to the subpathways they represent.
+
+
+
+_EHLD representing the Hemostasis branch of the Reactome event hierarchy. (a) Pathway hierarchy view for Hemostasis. (b) Hemostasis as an EHLD representation. (c) Hemostasis EHLD overlaid with pathway enrichment analysis results._
+
+### Generic guidelines
+
+To make all these possible, there is a set of initial requirements to be taken into account during the generation of the EHLDs.
+
+ * EHLDs must be in SVG format. Illustrators are free to use the design tool of their preference but at the end their diagram must be exported to SVG.
+ * The use of raster graphics is strongly discouraged as it results in larger file sizes and it has a negative impact on software performance. Also, by including bitmap images we cancel out resolution-independent zooming, which is one of SVG’s main advantage.
+ * All exported SVG files should include the styles in an internal CSS stylesheet. Please make use of this option when exporting from Adobe Illustrator: _Export As - > SVG -> Styling: Internal CSS_. If you are using another application to create the EHLDs, you will need to configure it appropriately.
+ * The name of each EHLD file has to be the Reactome Identifier of the corresponding pathway diagram, followed by “.svg”. For example, the _Apoptosis_ EHLD should be named as _R-HSA-109581.svg_. Please keep in mind that only one EHLD should be created per high level pathway diagram.
+ * Active regions are annotated using the id attribute of their group element (inside the SVG file) and can be classified in the following two groups:
+ * Regions containing a group of irregular shapes (including text) that can be selected (and hovered) in order to navigate to the respective pathway diagram. These regions should be annotated by setting their id attribute to “ _REGION-_ ” _\+ Reactome Identifier_ of their represented subpathway, e.g. “ _REGION-R-HSA-109581_ ”. This type of active regions may include arrows pointing to or from them and they are not mandatory.
+ * Regions containing the label of the represented subpathway in a box-like shape that can be not only selected but also overlaid with the results of the analysis. These regions should be annotated by setting their id attribute to “ _OVERLAY-_ ” _\+ Reactome Identifier_ of their represented subpathway, e.g. “ _OVERLAY-R-HSA-109581_ ”. Regions of this category must not include arrows pointing to or from them, and they are mandatory.
+ * Please keep in mind that if we need to annotate both types of active regions for a given subpathway, the region (“REGION-R-HSA-109581”) has to include the overlay component (“OVERLAY-R-HSA-109581”). In other words, the group of the selectable shapes has to include the group of shapes that can be overlaid.
+ * Any group of shapes, text or graphical elements that are not annotated as active regions are considered decorators and users cannot interact with them. They have pure esthetic value and they cannot be selected, hovered or overlaid.
+ * If possible, no background covering the whole image should be put in place. Please note that compartments or other necessary backgrounds can be kept, but we suggest to avoid “unnecessary” backgrounds. In case a background is required, but it is too big to fit in the illustration, it is suggested to draw only a small part of it with clear and well defined boundaries (See the blood vessel in the EHLD of Hemostasis above).
+ * Text elements should not be converted to groups of shapes. Instead, use the normal SVG element.
+
+
+
+Example illustrating the two types of active regions. Selectable regions (annotated with “REGION-” + Reactome Identifier) are highlighted in red, while regions that can be overlaid are highlighted in brown. Shapes and text that are not annotated as active regions are considered decorators (presented in blue) and users cannot interact with them.
+
+Taking the aforementioned requirements into consideration, the EHLD of the figure above should have the following structure:
+
+ * REGION-Pathway A _# The whole pathway will be selectable #_
+ * OVERLAY-Pathway A _# Only the label will be overlaid #_
+ * REGION-Pathway B _# The whole pathway will be selectable including the arrow #_
+ * OVERLAY-Pathway B _# Only the label will be overlaid #_
+ * REGION-Pathway C _# The whole pathway will be selectable including the arrow #_
+ * OVERLAY-Pathway C _# Only the label will be overlaid #_
+ * OVERLAY-Pathway D _# The whole pathway will be selectable and overlaid #_
+ * DECORATORS _# None of the shapes are selectable or can be overlaid #_
+
+### Creating an EHLD step by step
+
+We suggest starting a new EHLD by creating an Adobe Illustrator document with **1366px** in width and **768px** in height and using the **RGB colour mode**.
+
+Pathway Labels should have **centered,** **white** , **uppercased** , text using **Arial Bold** font at **12pt** inside a **rounded rectangle** with **8pt** **radius** , minimum **width of 170px** and **height** of**30px** for single lines or **43px** for double lines. The colour of the rounded rectangle should be **#0F82BC** (R:15 G:130 B:188). We suggest writing the name of the subpathway it represents as it appears in the Reactome hierarchy. In case of a discrepancy, or any kind of oddity, please contact the author or the curator related to the pathway.
+
+Please use the same specifications for any other written element except for:
+
+ * **Descriptions** of a process should always be black and in lowercase letters.
+ * Any **written element** that belongs to a compound, protein or cell (generically inside of any coloured shape) should be white and in uppercase letters (when possible).
+ * Any **written element** that belongs to a receptor and/or describes a cell element or a process (generically outside of any shape) should be black and in lowercase letters (when possible, usually receptors are named by acronyms and capital letters should be used in these cases).
+ * Greek or other Unicode characters must be avoided in text elements. Thus, names with greek characters should include the whole word (e.g. alpha instead of α) written in lowercase.
+ * Reactome **logo** should always have **50%** **opacity**.
+
+Every pathway label should have one **Analysis Information Label**. The latter is used to display additional details about the hit elements and the false discovery rate (FDR). All analysis information labels should be annotated as “ANALINFO” in the Layer Hierarchy Panel. Place “XXX/YYY” as a text placeholder inside it. Text should be centered and uppercased in **Arial Bold** **9pt** , and in white colour. The box should be a rounded rectangle with **8px radius** , minimum **width of 170px** and **20px height**. Its fill colour should be **#C6C6C6** (R:198 G:198 B:198). Also, set the opacity of the group to 0%, but make sure you leave the opacity of the shapes and text inside the group to 100%. Each analysis information label should be placed beside its pathway label and in the opposite side of the content for that pathway. For instance, if the pathway label is above its content, the analysis information label should be placed above the pathway label and vice versa.
+
+In order to better shape the active region (or the clickable area) of a pathway we can add a white shape with **0% opacity** under any element of the Pathway in the Layer Hierarchy Panel. In this way we can extent the clickable area and improve the overall user experience while navigating the EHLDs. In particular, we should try to surround the elements of the subpathway covering the gaps within the individual graphic elements, and thus, avoiding the inconvenient situation where the mouse cursor changes from the usual arrow shape to the hand constantly. Additionally, in EHLDs where elements of a particular subpathway are too thin and users find it difficult to select them, this technique can provide a bigger and more convenient clickable area around them. However, we should use this with caution, as creating too large clickable areas can mislead users.
+
+It is recommended to place all arrows and neutral text elements in a group at the top of the layer hierarchy. As well, all decorators and elements in the background that we do not want to highlight or be clickable, we recommend to place them in a group at the bottom of the layer hierarchy.
+
+Reactome displays EHLDs in a blank zoomable space. For a better user experience we suggest to have all shapes and elements portrayed in full. However, we understand that some pathways are large and contain too many and too big elements to fit in one image. In cases where the illustrator needs to cut an element and it is difficult to make a clean representation of the cut, we suggest the addition of a gradient so that elements vanish keeping the limits of the canvas clear.
+
+In order to export an EHLD we should use the “Export as” option and save the file as an SVG with the following options:
+
+ * Styling: Internal CSS.
+ * Font: SVG.
+ * Images: Preserve.
+ * Object IDs: Layer Names.
+ * Decimal Points: 3.
+ * Minify and Responsive, both ticked.
diff --git a/projects/website-angular/content/documentation/icon-info/icons-guidelines.mdx b/projects/website-angular/content/documentation/icon-info/icons-guidelines.mdx
new file mode 100644
index 0000000..88b1ac1
--- /dev/null
+++ b/projects/website-angular/content/documentation/icon-info/icons-guidelines.mdx
@@ -0,0 +1,156 @@
+---
+title: Icons Library Guidelines
+category: "documentation"
+---
+
+## Icons Library Guidelines
+
+This section will guide you through the process of creating and including your own graphic elements into the [Reactome Icon Library]() and become a part of this growing community.
+
+Since Reactome is a free and open-source project, the Reactome Library is under a Creative Commons license. As a result, if your creations are included in our library, they will follow the same policy. Feel free to use any tool or application that allows you to create **vector graphics** and, please, make sure that the following specifications are implemented in the files you will be creating.
+
+#### Generic guidelines
+
+ * Keep simplicity in mind. The main goal for this Icon Library is to provide easy to read graphic components for our users and simplicity is key to this matter.
+ * Consistency is important for Reactome and the Icon Library. Please, before submitting a proposal, read the guidelines we have set for the different types of components we use.
+ * Each icon should represent a single element only. After receiving feedback we open this rule to receptors when binding other receptors in order to make it easy for our users.
+ * Every element should be placed into a separate file, **200** by **200px** in size, with **RGB colour mode** and **White background**.
+ * Use only **Arial Bold 12pt**. Text colour should be chosen depending on the text’s position. In case the text is **inside** a coloured shape, its colour should be **White** , while in case the text is **outside** of a coloured shape, its colour should be **Black**.
+ * All text should be **Uppercased** except for words describing **greek** characters (e.g alpha, beta, gamma, etc.). In that case those words should be written in **Lowercase**. It should be noted that Greek or other Unicode characters must be avoided, as they are not supported for the moment.
+ * Your elements should only comprise vector graphics. Please make sure that you have not included any bitmap image in it. Including raster images often results in larger file sizes and thus negatively impacts software performance.
+ * You can place your new element in any of the seven categories of the Reactome Library; Cell elements, Cell types, Compounds, Human tissue, Ion channels, Proteins and Receptors.
+
+#### Cell elements
+
+Keeping simplicity is one of our targets so we recommend the use of **simple figures** , such as circles, squares and triangles as a base. For colouring we suggest the use of shades of **Primary Colours** (Red, Blue and Yellow) and **Secondary Colours** (Purple, Green and Orange).
+
+Vector graphic applications allow you to use **Gradient tools** to blend two or more colours for the same shape. We recommend you to use these tools so you can create more appealing graphics. See the Mitochondrion below:
+
+
+
+_(a) Without the use of gradients, the Mitochondrion element looks plain while in_
+
+_(b) through the correct use of gradients the element looks more 3D-like,_
+
+_resulting in a more attractive and playful image._
+
+#### Cell types
+
+In order to ensure a sense of unity and a consistent look and feel, we suggest applying the same principles; simple shapes and colours.
+
+For representing types of cells, please keep in mind that we are not using scale to portray the different organelles and elements inside the cell cytoplasm. Additionally, to maintain simplicity, try to avoid overloading the the overall design with too much detail. The following table features 3 examples:
+
+ | _This**Microbe** element is formed by a circle with a radial gradient of two colours. To represent the membrane, we have added an outline with a different gradient. To convey the idea of danger we use this squares as spikes and a red shade in their gradients._
+---|---
+ | _We have drawn this**Macrophage** element with shaky tentacles and we have used a not uniform outline (membrane) to give it a soft and flexible feeling._
+ | _For this**Pathogen** element we used a large light gradient to represent a shiny and hard capsule._ _To show its death we draw holes on its surface and play with a darkened, sad colour._
+
+#### Compounds
+
+Compounds are represented by very geometrical elements, following more specific guidelines.
+
+Simple chemical elements or compounds are usually portrayed in the Library with their chemical element symbol (e.g. Ca for Calcium) or the name of the compound (e.g. IFN-gamma) using **Arial bold 12pt** in **White** colour.
+
+Simple chemical elements are represented by an **octagon** of **28px** height and two plain colours, one for the fill and a different one for the stroke (**2pt**). In case the text of the compound is too long to fit in the octagon, we suggest stretching the shape until it fits the name, respecting its lateral edges, like follows:
+
+
+---
+_An example of a simple chemical element._
+
+__Complex compounds can be represented with a variety of shapes, always simple and easy to differentiate one from each other.
+
+ | _We can use circles with or without outline._
+---|---
+ | _We can use hexagons, in the same way as for the simple compounds, just adding a gradient on their stroke._
+
+#### Human tissue
+
+Human tissue elements are mainly used as backgrounds in our diagrams and the way of representing these organs depends on the illustrator’s skills and taste. To ensure unity in the Reactome Library we suggest to keep these illustrations as simple as possible and visualise them as toys; so they convey a nice, soft feeling, with a plastic-like, very friendly and approachable look.
+
+ | 
+---|---
+_Examples of human tissue elements; Liver and Blood vessel._
+
+#### Ion channels
+
+These representations are quite simplified and, thus, elements are designed to look like small funnels that cross through the membrane, allowing elements to move from inside of the cell to outside and vice versa. There are hundreds of different ion channels and we suggest differentiating them with the use of colours in various combinations.
+
+ | 
+---|---
+_Examples of ion channels._
+
+#### Proteins
+
+There are tens of thousands of different proteins, some of them with specific shapes. Proteins in the Reactome Library follow the standard representation as rounded rectangle shapes. As a result, we suggest using the **Rounded Rectangle Tool** to draw a shape with **10px** radius for the corners and **20px** height for single line of text, or **30px** for a double line. The stroke should be **3pt** and we suggest choosing an irregular profile (in Adobe Illustrator, use the default **Width Profile 2**). For the name of the proteins we suggest to follow the same guides as before, **White Uppercase Arial Bold 12pt**.
+
+The shape will contain the simplified name of the protein following the general instructions for Reactome Library. It should be filled with a simple gradient of two colours. One of the colours of this gradient should also be used as the stroke colour.
+
+ | 
+---|---
+_Examples of protein elements in their rounded rectangle representation._
+
+In case a protein has a specific shape, we suggest keeping the standard representation as long as it is not too complicated. Otherwise, we suggest opting for a more simplified version.
+
+ | 
+---|---
+_Examples of protein elements represented in their standard representation._
+
+#### Receptors
+
+Like proteins, there is a huge number of receptors. In cases where there is a standard, but quite complex, way to represent them, we suggest adopting a more simplified version. If this is too difficult we suggest finding a more simple representation.
+
+ | 
+---|---
+_Examples of receptor elements_
+
+#### Metadata
+
+You are encouraged to accompany your element with more information including a short description and extra details about its designer and curator. Upon inclusion of your element into the Reactome Library, all metadata information will be available to the public through our landing page.
+
+In order to do so, you need to create an additional metadata file using your favorite text editor. The file should have the following structure:
+
+
+
+
+ **{CATEGORY}**
+
+ **{CURATOR_NAME}**
+ **{DESIGNER_NAME}**
+ **{ICON_NAME}**
+ **{DESCRIPTION}**
+
+
+ **{REFERENCE_NAME}**
+ **{REFERENCE_ID}**
+
+
+
+ **{SYNONYM}**
+
+
+
+
+Please follow these steps:
+
+ 1. Create a new file using your favorite text editor (TextEdit, NotePad, etc.).
+ 2. Copy and Paste the text above into your file.
+ 3. Edit the file and:
+ 1. Fill {CATEGORY} with one or more of the suggested categories: cell_element, cell_type, compound, human_tissue, protein, receptor or transporter. We accept one or more different categories as we think they are just a guide that helps our users to find the element they need.
+ 2. Fill {ORCID} and/or {URL} for the curator and designer roles. This is [how to find ORCID](). In case both URL and ORCID are provided, the link for that person will point to the URL.
+ 3. Fill {CURATOR_NAME} and {DESIGNER_NAME} as the curator and designer name, respectively. If the name is the same, please fill both with same information.
+ 4. Fill {DESCRIPTION} with any information relevant to describe the component. Please be brief.
+ 5. Fill {REFERENCE_NAME} for the name of the resource you have used to find this component (e.g.:CHEBI, UNIPROT, GO … ). In case of not finding a suitable reference, please delete this line.
+ 6. Fill {REFERENCE_ID} as it appears in the resource’s URL (e.g.: CHEBI:1234, Q1234, GO:1234 … ). In case of not finding a suitable reference, please delete this line.
+ 7. Fill {SYNONYM} as any synonyms or alternative names your component might be known of. In case of not finding a suitable synonym, please delete this line.
+ 4. Save the file using the same name as the element but make sure to use .xml as its extension. For example if your graphic element is stored with the name “Element.svg” then your metadata file should be named as “Element.xml”.
+ 5. Make sure to submit your metadata file along with the graphics file.
+
+### Export Settings
+
+Currently, every element in the Reactome Library is available in 3 different formats, SVG, EMF and PNG. Therefore, you are kindly requested to export your work into these formats and include them in your submission.
+
+For PNG files we suggest to export with 300 DPI and transparent background.
+
+### How to submit your work
+
+To include your graphic elements in the Reactome Library, you need to contact our helpdesk (This email address is being protected from spambots. You need JavaScript enabled to view it.). We will then review your submitted files and, depending on the outcome, we will include them in our Library.
diff --git a/projects/website-angular/content/documentation/index.mdx b/projects/website-angular/content/documentation/index.mdx
index 5c036c9..c3e8a4c 100644
--- a/projects/website-angular/content/documentation/index.mdx
+++ b/projects/website-angular/content/documentation/index.mdx
@@ -1,21 +1,21 @@
---
-title: Documentation
+title: "Documentation"
category: "documentation"
---
-[  ]()
+[  ]()
Take the most out of our tools and data analysis. All you need is our User Guide.
#### __[ For Users ]()
-[  ]()
+[  ]()
Are you interested in programatically querying our data or integrating our Widgets ?
#### __[ For Developers ]()
-[  ]()
+[  ]()
Have our data been useful in your research or experiment ? Please, remember to cite us!
diff --git a/projects/website-angular/content/documentation/inferred-events.mdx b/projects/website-angular/content/documentation/inferred-events.mdx
index 8ef08e0..45b820f 100644
--- a/projects/website-angular/content/documentation/inferred-events.mdx
+++ b/projects/website-angular/content/documentation/inferred-events.mdx
@@ -1,6 +1,6 @@
---
title: Computationally Inferred Events
-category: ""
+category: "documentation"
---
## Computationally Inferred Events
@@ -21,4 +21,4 @@ These electronically inferred reactions are predictions based on a number of ass
A modified version of the orthoinference process was used to create a first draft of the [SARS-CoV-2 infection pathway](). Events in the SARS-Cov-2 pathway corresponding to each event in the [SARS-CoV-1 pathway]() were created and populated with SARS-CoV-2 protein-containing physical entities based on orthology to SARS-CoV-1 proteins. We continue to replace predicted SARS-Cov-2 events with experimentally validated events when the relevant experimental evidence becomes available.
-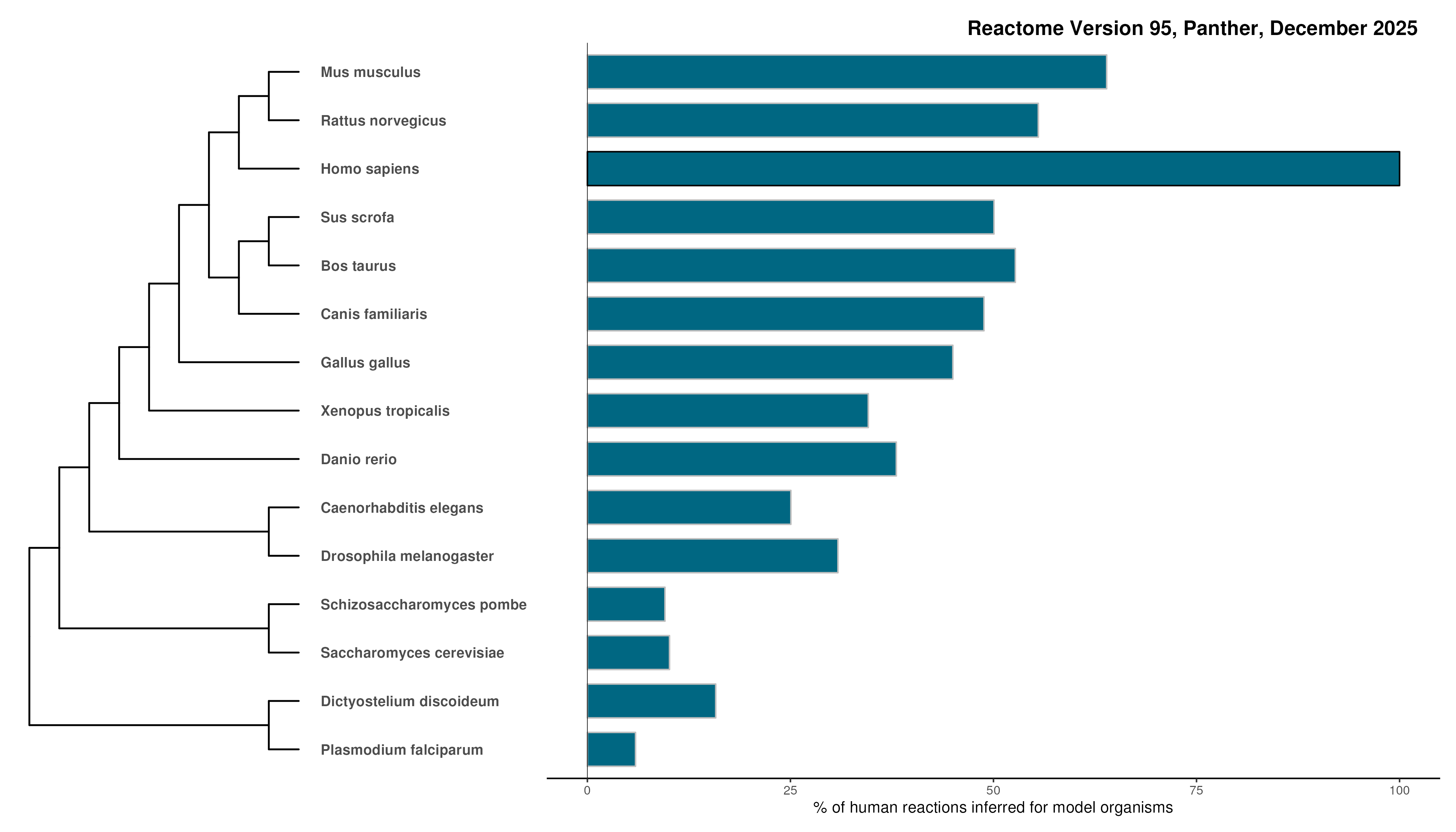
+
diff --git a/projects/website-angular/content/documentation/linking-to-us/identifiers.mdx b/projects/website-angular/content/documentation/linking-to-us/identifiers.mdx
new file mode 100644
index 0000000..b78e6c8
--- /dev/null
+++ b/projects/website-angular/content/documentation/linking-to-us/identifiers.mdx
@@ -0,0 +1,53 @@
+---
+title: External Identifiers
+category: "documentation"
+---
+
+## External Identifiers
+
+Listed below are the external bioinformatics databases that Reactome provides link outs to from our website.
+
+ * [AraCyc]()
+ * [BioGPS]()
+ * [BioModels]()
+ * [CAS]()
+ * [COMPOUND]()
+ * [COSMIC]()
+ * [CTD Gene]()
+ * [ChEBI]()
+ * [ClinGen]()
+ * [dbSNP Gene]()
+ * [dictyBase]()
+ * [DOCK Blaster]()
+ * [DOID]()
+ * [EC]()
+ * [ENSEMBL]()
+ * [Flybase]()
+ * [GO]()
+ * [GeneCards]()
+ * [Guide to Pharmacology]()
+ * [HGNC]()
+ * [IntEnz]()
+ * [KEGG Gene]()
+ * [miRBase]()
+ * [MOD]()
+ * [NCBI Gene]()
+ * [NCBI Nucleotide]()
+ * [NCBI_Protein]()
+ * [OMIM]()
+ * [ORCID]()
+ * [Orphanet]()
+ * [PRF]()
+ * [PlasmoDB]()
+ * [Protein Data Bank]()
+ * [PubChem Compound]()
+ * [PubChem Substance]()
+ * [RefSeq]()
+ * [Rhea]()
+ * [SGD]()
+ * [SO]()
+ * [TAIR]()
+ * [UCSC human]()
+ * [UniProt]()
+ * [Wormbase]()
+ * [ZINC]()
diff --git a/projects/website-angular/content/documentation/release-documentation.mdx b/projects/website-angular/content/documentation/release-documentation.mdx
new file mode 100644
index 0000000..d540010
--- /dev/null
+++ b/projects/website-angular/content/documentation/release-documentation.mdx
@@ -0,0 +1,8 @@
+---
+title: Release Documentation
+category: "documentation"
+---
+
+## Release Documentation
+
+The current Release SOP with associated appendices (last updated V95) is available for download [here]().
diff --git a/projects/website-angular/content/documentation/userguide.mdx b/projects/website-angular/content/documentation/userguide.mdx
index 6caa160..ec3ae5c 100644
--- a/projects/website-angular/content/documentation/userguide.mdx
+++ b/projects/website-angular/content/documentation/userguide.mdx
@@ -1,6 +1,6 @@
---
title: Pathway Browser
-category: ""
+category: "documentation"
---
### What is Reactome?
@@ -9,7 +9,7 @@ Reactome is a curated database of pathways and reactions in human biology. React
### What is this guide For?
-This guide introduces features of the Reactome website using a combination of short explanations and exercises. You will learn how to search, interpret the views, use the tools and if necessary find documentation or [contact us]() for help.
+This guide introduces features of the Reactome website using a combination of short explanations and exercises. You will learn how to search, interpret the views, use the tools and if necessary find documentation or This email address is being protected from spambots. You need JavaScript enabled to view it. for help.
[ __ ]()
@@ -43,19 +43,19 @@ This guide introduces features of the Reactome website using a combination of sh
Available via the EBI Train Online system
-[  ]()
+[  ]()
This quick tour provides a brief introduction to Reactome knowledgebase and its various features.
#### __[ Quick tour ]()
-[  ]()
+[  ]()
This course provides a more detailed view of the Reactome pathway database web interface and the database content.
#### __[ Exploring Pathways ]()
-[  ]()
+[  ]()
This course introduces the analysis tools available via the Reactome pathway database web interface.
diff --git a/projects/website-angular/content/documentation/userguide/analysis/analysis.mdx b/projects/website-angular/content/documentation/userguide/analysis.mdx
similarity index 100%
rename from projects/website-angular/content/documentation/userguide/analysis/analysis.mdx
rename to projects/website-angular/content/documentation/userguide/analysis.mdx
diff --git a/projects/website-angular/content/documentation/userguide/analysis/gsa.mdx b/projects/website-angular/content/documentation/userguide/analysis/gsa.mdx
new file mode 100644
index 0000000..150e58c
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/analysis/gsa.mdx
@@ -0,0 +1,115 @@
+---
+title: ReactomeGSA
+category: "documentation"
+---
+
+## ReactomeGSA
+
+####
+
+#### Quantitative, multi-dataset Pathway Analysis (ReactomeGSA)
+
+ReactomeGSA is a new pathway analysis tool integrated into the Reactome ecosystem. Its main feature is that it performs quantitative pathway analyses (so-called gene set analyses). This increases the statistical power of the differential expression analysis, which is directly performed on the pathway level.
+
+ReactomeGSA can analyse multiple datasets simultaneously resulting in a comparative pathway analysis. Thereby, it is possible to quickly assess whether the same effect was observed in independent experiments or studies.
+
+ReactomeGSA currently supports quantitative proteomics, transcriptomics, and microarray data. Datasets from all of these methods can be combined in a single analysis. Thereby, ReactomeGSA can perform multi-omics pathway analyses.
+
+#### Using ReactomeGSA
+
+We currently offer three ways to access ReactomeGSA:
+
+ 1. Reactome’s web-based pathway browser (see below)
+ 2. From R using our ReactomeGSA Bioconductor R package (see [here]())
+ 3. Programmatically, using the ReactomeGSA API at []()
+
+> #### **Analyze Gene Expression using ReactomeGSA**
+
+You can visit ReactomeGSA [here](). When you click on the “Analyse gene expression” tab of the Analysis Tools, you will be redirected to ReactomeGSA, where you can perform analyses.
+
+The following tutorial will show you how to: navigate to ReactomeGSA, submit data for analysis, annotate your data, and perform your first analysis. This following tutorial shows a simplified analysis workflow.
+
+
+
+#### ReactomeGSA Analysis Algorithms
+
+In this first screen, you need to select the algorithm to use for the differential pathway analysis. At the time of writing, ReactomeGSA offers three algorithms. PADOG and Camera perform a differential expression analysis between two groups of samples. ssGSEA is a so-called gene set variation approach that returns pathway-level quantitative data for each sample.
+
+The detailed parameters for each algorithm can be adapted by clicking the blue icon on the left of the algorithm’s box.
+
+#### Adding datasets
+
+After clicking “Next”, you are presented with the now empty list of datasets. Click the “+ Add dataset” button to add a new dataset.
+
+First, you have to select the type of dataset you want to load.
+
+To upload your own data, select one of the options under “Select a file from a local folder”. The file must be a tab-delimited text file (CSV or TSV file) where the first column contains the gene or protein identifiers and all subsequent columns the respective samples. The first row contains the sample names and all subsequent rows the genes / proteins.
+
+To test the tool, it is possible to quickly load example data. Currently, ReactomeGSA provides three example datasets, two (matched) datasets on melanoma associated B cells (proteomics and transcriptomics measurements) and one scRNA-seq dataset on B cells.
+
+#### Annotating experimental metadata
+
+Once a dataset is added, you need to annotate the experimental metadata. This is necessary in order to define the groups for the differential expression analysis in the next step.
+
+You can adapt the dataset’s name in the “Dataset name” box at the top. This name will be used for all results. To increase the readability, we suggest to use as short names as possible.
+
+In case you load data from ExpressionAtlas or choose one of the example datasets (as shown in the screenshot) the sample annotation table will already be pre-filled with certain metadata. In case you uploaded your own dataset, the table will only show the orange sample labels on the left.
+
+To add an annotation, click the “plus” symbol on the right. This will add a new empty column to the table. First, add a heading to define the name of the property (for example, “treatment”). Next, add the values for every sample that you want to include in your comparisons. Samples without any values will simply be ignored.
+
+#### Defining the experimental design
+
+In the final step of adding a dataset, you have to define which groups to compare. The “comparison factor” drop-down menu contains all parameters that were annotated in the sample table before (if they contain at least two different values). “1st group” and “2nd group” define which groups of samples to compare against each other. The 1st group is the control or baseline.
+
+Depending on which “comparison factor” you select, the available values for the “1st group” and “2nd group” will change automatically.
+
+Additionally, some gene set analysis methods allow you to define so-called “covariates”. These are parameters that might cause a bias in your result (ie. the sequencing facility used) that you would like to correct for. Simply select the relevant ones for your experiment.
+
+Once you click “Continue” you will be returned to the list of datasets where you will now see your annotated dataset in the list. If you want, you can add any number of datasets to a single request.
+
+#### Starting the analysis
+
+“Create REACTOME visualizations” is always selected. If it is de-selected, the result cannot be visualised in Reactome’s pathway browser. This option is generally only relevant to users of the [ReactomeGSA R package]().
+
+If you select “Create reports” ReactomeGSA will automatically create a Microsoft Excel and PDF report of your results. Additionally, it will create a short R script that allows you to load your data directly into an R session.
+
+In case you provide your email address, you are automatically notified as soon as the analysis is complete. The mail will contain direct links to the generated reports (if you chose to create them) and a link to the visualisation in the PathwayBrowser.
+
+Launch the analysis by clicking the “GO” button.
+
+If you provided an email address, you can also close your browser. The analysis will still continue on our servers and you will be notified as soon as it is done. Some analysis with many large datasets (for example comparing five TCGA datasets in a single analysis) may require up to an hour to complete.
+
+> #### Loading Public Datasets
+
+The following tutorial will show you how to: load public data for analysis from EBI Expression Atlas, EBI Single Cell Expression Atlas, GREIN and GEO data.
+
+
+
+Finally, it is possible to directly load datasets from several sources.
+
+ 1. ExpressionAtlas. To do so, first navigate to ExpressionAtlas (in a separate tab or window) at [](). Once you have identified a dataset of interest, you can get the dataset’s id by opening the dataset and copying the portion after “[www.ebi.ac.uk/]()” and before the next “/”. For example, if you opened []() the identifier of this dataset would be “E-MTAB-6592”.
+
+
+ 2. Single Cell Expression Atlas. For single-cell experiments you additionally have to define the parameter “k” in order to define which clusters should be used. The effect of “k” on the results can be visualised on the first page of the respective single cell experiment in ExpressionAtlas (see []()). In this case, the dataset identifier would be “E-MTAB-7008" and a possible value for “k” would be 10.
+
+
+ 3. GREIN data. The accession ID is provided in the URL.
+ 4. GEO Microarray data. After choosing the Pathway algorithm the user can choose public datasets in the "Public datasets" panel and the "GEO query" section to fetch public microarray data via the GEO accession id. e.g. GSE1563. In the process the data is fetched directly from the GEO database and can be annotated, downloaded and analysed within the ReactomeGSA workflow.
+
+> #### Citing ReactomeGSA
+
+#### **If you use ReactomeGSA in your research, please cite the following publication:**
+
+ReactomeGSA - Efficient Multi-Omics Comparative Pathway Analysis
+
+Johannes Griss, Guilherme Viteri, Konstantinos Sidiropoulos, Vy Nguyen, Antonio Fabregat, Henning Hermjakob
+
+_Mol Cell Proteomics._ 2020 Dec; 19(12): 2115–2124; [Link]()
+
+#### **If you use ReactomeGSA's data integration, please cite the following publication:**
+
+ReactomeGSA: new features to simplify public data reuse
+
+Alexander Grentner, Eliot Ragueneau, Chuqiao Gong, Adrian Prinz, Sabina Gansberger, Inigo Oyarzun, Henning Hermjakob, Johannes Griss
+
+_Bioinformatics_ , Volume 40, Issue 6, June 2024; [Link]()
diff --git a/projects/website-angular/content/documentation/userguide/claim-your-work.mdx b/projects/website-angular/content/documentation/userguide/claim-your-work.mdx
new file mode 100644
index 0000000..1c61097
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/claim-your-work.mdx
@@ -0,0 +1,36 @@
+---
+title: "Tutorial: How to claim your works"
+category: "documentation"
+---
+
+## Tutorial: How to claim your works
+
+ * [Find yourself in Reactome](<#guide>)
+ * [I have an ORCID ID but it is not in Reactome. What should I do ?](<#missing_orcid>)
+
+### Find yourself in Reactome
+
+ 1. The simple text search tool is located at top right of the Home page. To search type your name or ORCID. The search has an auto-complete function; if the text you wanted to use appears in the drop-down list, select it and results will be displayed. For other text click Search.
+
+ 2. Locate yourself in the Result List and Click your name.
+
+ 3. You'll land in the Contributor's page. You may see up to four categorized tables showing a maximum of 15 rows each.
+
+ 4. In order to claim your work you may need to Authenticate yourself in ORCID, in a similar way like others websites offer 'Sign in with Google', 'Sign in with Facebook'. Simply click "Are you John Doe ?". A pop-up like to the one below will open. Please input your ORCID credentials, then Click Sign-in.
+
+ 5. Reactome.org will ask your permission to add works into your ORCID records. Authorize if you wish to grant access, otherwise we can't have access to
+
+ 6. After authorizing, we will automatically proceed with the authorization process and the pop up will be closed and current page will be refreshed. A button "Claim all Work" may appear next to your ORCID number if we are able to match ORCID IDs. Click it to make sure all your contributions are uploaded.
+
+ 7. Depending on the number of entries you are uploading it may take a few minutes. You may see a Claiming Summary when the entire process is finished.
+
+ 8. Visit your ORCID page and Navigate to the Works Section.
+ 9. All your work now is uploaded.
+
+### I have an ORCID ID but it is not in Reactome. What should I do ?
+
+ 1. Follow intructions 1 to 5 in the previous section.
+ 2. You may see a Link "Let us know your ORCID". Please Click.
+
+ 3. After clicking, use the following form to send your ORCID details to our curators. Our help team will validate and add your ORCID into our database and will be available to you on our next data release. Follow us on Twitter for any updates.
+
diff --git a/projects/website-angular/content/documentation/userguide/cytomics.mdx b/projects/website-angular/content/documentation/userguide/cytomics.mdx
new file mode 100644
index 0000000..2ab1f92
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/cytomics.mdx
@@ -0,0 +1,152 @@
+---
+title: Cytomics
+category: "documentation"
+---
+
+## Cytomics
+
+#### Introduction
+
+Human bodies are built of >30 trillion cells specialized to fulfill diverse roles within our tissues, organs, and organ systems. All these cells originate from a single cell, a zygote formed at conception. From zygote to fetus, and throughout childhood, adolescence, and adulthood, cells divide and commit to different fates in order for the organism to develop, sustain and regenerate. The series of steps that lead from an undifferentiated progenitor cell, such as a stem cell, to one of a number of possible specialized descendants constitutes a cell lineage path. Recent technological advances have allowed researchers to collect high-throughput omics data from single cells of multicellular organisms and use it to track and manipulate cell fates [(Burgess 2018; Saelens et al. 2019)](). This opens the door to the possibility of deciphering cell lineage paths at single-cell resolution, a critical requirement for the advancement of regenerative medicine and cancer medicine.
+
+Researchers who conduct experiments that involve cell lineage tracing through the study of genomic, epigenomic, transcriptomic, and proteomic features of single cells currently expend significant effort extracting information from published sources to create short-lived, limited-scope local annotations of biological markers that are needed to interpret and analyze their high-throughput experimental data. This creates the urgent need for an open source, comprehensive, continuously maintained reference repository of cell lineage paths framed on the pre-existing knowledge of tissue morphogenesis, as well as a cell type-specific pathway resource, which can serve as a framework for interpretation and integration of the latest experimental findings.
+
+Cytomics Reactome aims to become an integrative systems biology resource of cell lineage paths and a toolset for the analysis of single-cell omics data. The Reactome web application and pathway visualization tools [(Croft et al. 2011)]() have been extended to enable Cytomics Reactome visual display and search functions (Milacic et al. 2024) on the live website.
+
+#### Database schema features to support Cytomics
+
+The Reactome data model provides a robust framework for the annotation of biological entities and events [(Joshi-Tope et al. 2005)](). The basic unit of the Reactome data model is a Reaction, an event that converts input to output physical entities. The participants in reactions are PhysicalEntities, which can be simple or complex molecules or molecular complexes. These PhysicalEntities can also act as reaction catalysts and regulators. Reactions are grouped into causal chains to form Pathways [(Joshi-Tope et al. 2005)]().
+
+The Reactome data schema was expanded as described below to accommodate new classes needed for Cytomics Reactome annotations.
+
+ * Cell: A subclass of PhysicalEntity, represents a type of cell in a particular state of development/differentiation (Figure 1A).
+ * A Cell instance has CellType, TissueLayer, Tissue, and Organ as single-valued attributes which are populated using terms from public open source databases, the Cell Ontology [(Sarntivijai et al. 2014; Osumi-Sutherland 2017)]() and UBERON [(Haendel et al. 2014)](), and cross-reference these databases.
+ * Cell instances also have multi-valued ProteinMarker and RNAMarker attributes, which are specific to the cell type and/or the cell state or are differentially upregulated in the Cell. (Figure 1B). The markers (protein or RNA) are manually curated; each marker instance is associated with one or more literature references for evidence, including, if applicable, citations of CellMarker (Hu et al. 2022) and PanglaoDB databases (Franzén et al. 2019) (Figure 1B)
+
+
+
+**Figure 1**. Cell Physical Entity class A and new attributes B
+
+ * Cell Development Step and Cell Lineage Path are two Event subclasses (Figure 2A), that describe developmental/differentiation relationships among Cells:
+ * A Cell Development Step is a ReactionLikeEvent that contains Cell instances as its inputs and outputs. The input attribute represents the cell of origin, and the output attribute represents the destination cell type. Additional Cell Development Step attributes include regulators (molecules promoting or inhibiting the step) and required input components (input cell proteins required for the action of regulators) (Figure 2B).
+ * A Cell Lineage Path, organizationally similar to the preexisting Pathway subclass, is composed of other Cell Lineage Path instances or Cell Development Steps as subevents that are connected through shared input/output Cells (Figure 2C).
+ * Both Cell Development Step and Cell Lineage Path are associated with a relevant Gene Ontology (GO) biological process term [(The Gene Ontology Consortium 2019)]() and with a “tissue” term from UBERON [(Haendel et al. 2014)]().
+
+
+
+**Figure 2.** Cell Development Step and Cell Lineage Path schema (A) and attributes (B,C).
+
+The Cytomics content is searchable on the Reactome website. Running a search from the homepage search bar may list hits within various categories including pathway, reaction, complex, and cell. Clicking on an item in the search results list will redirect to a Details page, which displays more information about the specific record.
+
+ * For example, a details page for Cell (Figure 3A) includes information on location in the pathway hierarchy, cell type, and histological description of the given cell with ontology terms, a list of protein and RNA markers, and corresponding literature references for each marker. A scroll bar appears when there are more than 5 markers in the same category. The page also lists event(s) (i.e., Cell Development Step), in which the Cell participates as an input/output.
+ * Clicking on marker or Cell Development Step, will redirect to the details page of the marker or event respectively.
+ * Clicking on the Cell in the expanded hierarchical view of the Developmental Biology pathway (Figure 3B) in the details page, will redirect to the Pathway Browser view highlighting the Cell Lineage Path in the hierarchy panel and showing the selected Cell in the pathway diagram.
+
+
+
+**Figure 3**. Details page for Cell (A)
+
+
+
+**Figure 3**. Expanded view of the Cell location in the Pathway Browser (B)
+
+ * See the details pages for marker instance (Figure 4), Cell Development Step (Figure 5), Cell Lineage Path (Figure 6) below.
+ * The content of individual events can be exported from the event’s details pages (Figure 5, 6) as SBML, PDF, SVG, PNG, PPTX, SBGN files.
+
+
+
+**Figure 4**. Details page for Marker instance
+
+
+
+**Figure 5**. Details page for Cell Development Step
+
+
+
+**Figure 6**. Details page for Cell Lineage Path
+
+#### Navigation of a Cell Lineage Path in the Pathway Browse
+
+Cell Lineage Paths are available under the “Developmental Biology” pathway in the Reactome Pathway Browser, where they are grouped in the “Developmental Cell Lineages” subpathway (Figure 7).
+
+
+Although the pathway diagram depicts a selection of cell types that are planned for annotation, only one type has been annotated so far and this is indicated by a selectable blue box label in the diagram.
+
+
+
+**Figure 7.** Pathway browser view of “Developmental Cell Lineages” subpathway
+
+Selecting an individual Cell Lineage Path in the pathway hierarchy will display all Cell Development Steps of the selected path in the pathway diagram (Figure 8). An individual Cell is depicted with a double-layered outer compartment representing the plasma membrane and the cytosol, along with an inner blue rectangle symbolizing the nucleus.
+
+The Description tab in the Details panel of a selected Cell Lineage Path provides additional information about the Cell Lineage Path including name, species, assigned GO Biological Process term (if applicable), and literature reference(s) (Figure 8).
+
+
+
+**Figure 8.** Selecting a Cell Lineage Path from the pathway hierarchy
+
+Clicking on an individual Cell Development Step either in the pathway hierarchy or in the pathway diagram will display relevant information for the selected event in the details panel below the diagram (Figure 9). The description tab of the details panel shows input and output, which are defined by Cell instances, regulators of the event (when applicable), GO biological process, and UBERON tissue terms. It may also display preceding event(s) connecting individual Cell Development Steps within the same Cell Lineage Path (Figure 9). Details of participating Cell instances including histological terms, specific protein and RNA markers, and corresponding marker references are accessible from the description tab of Cell Development Step upon clicking on the plus sign to the right of the input/output attributes.
+
+
+
+**Figure 9.** Selecting a Cell Development Step
+
+Alternatively, the Cell details can be displayed in the Details panel by selecting a “Cell” icon in the diagram of the Cell Lineage Path (Figure 10). Right-clicking or clicking the blue info icon when hovering over a “Cell” icon in the pathway diagram will open a popup list of protein and RNA markers for the selected Cell (Figure 11).
+
+
+
+**Figure 10.** Selecting a Cell
+
+
+
+**Figure 11.** Displaying the list of Cell markers
+
+The Molecules tab in the Details panel (Figure 12A) displays a downloadable list of all markers for a Cell and both the markers and regulators for a Cell Development Step and a Cell Lineage Path.
+
+The Structure tab (Figure 12B) in the details panel displays the Protein Data Bank structures of molecules listed in the Molecular tab and the Expression tab shows gene expression data from Expression Atlas.
+
+
+
+**Figure 12.** Displaying Molecules Tab (A)
+
+
+
+**Figure 12.** Structures tab (B)
+
+The Analysis tab (Figure 13, 14) in the details panel displays data after running the dataset analysis. Refer to (Rothfels et al. 2023) for detailed instructions on how to use the Reactome Analysis Tool.
+
+The results of the analysis are also visualized in the pathway diagram panel, where the “Cell” icons are colored (Figure 13, 14). The colored area in the nucleus of the individual Cell corresponds to the coverage of the cell markers in the submitted data set.
+
+In Pathway Enrichment Analysis, olive green is applied to color the nucleus of the cells in the Cell Lineage Path that contain markers from the query list. The extent of coloration is proportional to the number of markers hit (Figure 13).
+
+The results of gene expression analysis, including Reactome Gene Set Analysis (Reactome GSA), appear as a single vertical bar of color when zoomed out representing the average expression of the markers from the query list (Figure 14a). Zooming in on the “Cell” icon reveals bars for individual cell markers with their expression values, as shown in Figure 14b.Expression values of cell markers from the query data set can be visualized in a popup information panel of protein and RNA markers for the selected Cell (Figure 14b).
+
+Expression values for the markers of a selected cell are displayed as blue horizontal lines within the expression scale (Figure 14C). The median cell marker expression among the cell's matching markers is shown in black. On the left, when a specific marker bar is hovered over, the currently hovered marker’s expression value is displayed in red, and the expression values of other markers in the same cell are shown in yellow.
+
+
+
+**Figure 13.** Overrepresentation analysis displayed within the nuclei of the cells in the Cell Lineage Path
+
+
+
+**Figure 14.** Gene expression analysis results are displayed within the nuclei of the cells. The level with coloration reflects the average expression value of the cell markers (A)
+
+
+
+**Figure 14.** A zoomed-in view of the expression bars of individual markers (B)
+
+
+
+**Figure 14. T** he expression scale (C)
+
+ 1. Burgess, Darren J. 2018. “Tracing Cell-Lineage Histories.” _Nature Reviews. Genetics_.
+ 2. Croft, David, Gavin O’Kelly, Guanming Wu, Robin Haw, Marc Gillespie, Lisa Matthews, Michael Caudy, et al. 2011. “Reactome: A Database of Reactions, Pathways and Biological Processes.” _Nucleic Acids Research_ 39 (Database issue): D691–97.
+ 3. Franzén O, Gan LM, Björkegren JLM, "PanglaoDB: a web server for exploration of mouse and human single cell RNA sequencing data", Database (Oxford), 2019, 2019.
+ 4. Haendel, Melissa A., James P. Balhoff, Frederic B. Bastian, David C. Blackburn, Judith A. Blake, Yvonne Bradford, Aurelie Comte, et al. 2014. “Unification of Multi-Species Vertebrate Anatomy Ontologies for Comparative Biology in Uberon.” _Journal of Biomedical Semantics_ 5 (May): 21.
+ 5. Hu C, Li T, Xu Y, Zhang X, Li F, Bai J, Chen J, Jiang W, Yang K, Ou Q, Li X, Wang P, Zhang Y, "CellMarker 2.0: an updated database of manually curated cell markers in human/mouse and web tools based on scRNA seq data", Nucleic Acids Res, 2022.
+ 6. Joshi-Tope, G., M. Gillespie, I. Vastrik, P. D’Eustachio, E. Schmidt, B. de Bono, B. Jassal, et al. 2005. “Reactome: A Knowledgebase of Biological Pathways.” _Nucleic Acids Research_ 33 (Database issue): D428–32.
+ 7. Milacic M, Beavers D, Conley P, et al. The Reactome Pathway Knowledgebase 2024. _Nucleic Acids Res_. 2024;52(D1):D672-D678.
+ 8. Osumi-Sutherland, David. 2017. “Cell Ontology in an Age of Data-Driven Cell Classification.” _BMC Bioinformatics_ 18 (Suppl 17): 558.
+ 9. Saelens, Wouter, Robrecht Cannoodt, Helena Todorov, and Yvan Saeys. 2019. “A Comparison of Single-Cell Trajectory Inference Methods.” _Nature Biotechnology_ 37 (5): 547–54.
+ 10. Rothfels Karen, Milacic Marija, Matthews Lisa et al. 2023. “Using the Reactome Database.” _Current protocols_ vol. 3,4: e722.
+ 11. Sarntivijai, Sirarat, Yu Lin, Zuoshuang Xiang, Terrence F. Meehan, Alexander D. Diehl, Uma D. Vempati, Stephan C. Schürer, et al. 2014. “CLO: The Cell Line Ontology.” _Journal of Biomedical Semantics_ 5 (August): 37.
+ 12. The Gene Ontology Consortium. 2019. “The Gene Ontology Resource: 20 Years and Still GOing Strong.” _Nucleic Acids Research_ 47 (D1): D330–38.
diff --git a/projects/website-angular/content/documentation/userguide/details-panel.mdx b/projects/website-angular/content/documentation/userguide/details-panel.mdx
new file mode 100644
index 0000000..475f79f
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/details-panel.mdx
@@ -0,0 +1,168 @@
+---
+title: Details Panel
+category: "documentation"
+---
+
+## Details Panel
+
+The Details Panel is the bottom right panel of the Pathway Browser. It displays the details of molecular objects or events when selected in the Pathway Diagram or Hierarchy. The details shown depend on the type of molecule or event selected. The Details Panel can be revealed/hidden using the middle Layout button.
+
+The Details Panel has several tabs:
+
+### **The Description tab**
+
+This tab contains the defining details of the selected diagram object. These details vary, depending on what is selected in the diagram. For example, reaction details always describe the input and output molecules, and where relevant, the enzyme catalyst. Other items in the Description tab may include:
+
+Summation – a summary of the molecular event, often with background information and additional literature citations
+
+Preceding/Following Events - Links to reactions that precede/follow this reaction.
+
+Cellular Compartment - Identifies the cellular compartment for the reaction, with the associated Gene Ontology (GO) term.
+
+Inferred from another species - Reactome represents human biology. Wherever possible key references contain experimental data obtained using human reagents. If the only data available is from model organisms, a human event may be inferred from this data, if the Author, Curator and Reviewer agree that this inference is valid. The inferred event is labelled with an icon ‘Inferred from another species’ in the Hierarchy and in the Details panel, where there is a link to the molecular details of the event as it occurs in the model organism and the supporting literature references. A description of the inference process can be found here.
+
+Computationally Inferred To – This drop-down list represents the species available for computationally predicted pathways, inferred from Reactome’s human pathway. References – all events in Reactome include links to original research publications containing experimental data, the evidence that the event has been shown to exist.
+
+Authored - an expert biologist who contributed materials allowing construction of the reaction/pathway. All Reactome pathways are authored by an expert biologist.
+
+Reviewed – all Reactome pathways are peer reviewed by an expert biologist who verifies the content provided by the Author.
+
+Details are organised into panels. Panels that have a plus icon on the right hand side can be expanded to reveal further levels of detail by clicking on the icon or the detail name.
+
+
+
+Clicking the plus icon reveals further details:
+
+
+
+Panels that represent molecules have an icon on the left hand side that represents the molecule subtype, e.g. the protein icon is a green circle. Mouse-over the icon to see an explanation of the subtype. The icon is a selection button; clicking the icon replaces the current details with a new panel of details specific to the selected molecule, which is highlighted on the pathway diagram.
+
+### **The Molecules Tab**
+
+This tab details all the molecules involved in the pathway displayed in the panel above. When a pathway name is highlighted in the hierarchy, the total number of molecules in that pathway is shown in a bubble on the molecules tab.
+
+Molecules are grouped into the subtypes Protein, Chemical Compound, Sequences (for DNA/RNA) and Other.
+
+If an item is selected in the diagram, the corresponding molecules are highlighted in the details panel, and the bubble next to the ‘Molecules’ tab enumerates the fraction of the total pathway molecules that are represented in that item.
+
+
+
+The Download button to the right of the pathway name and species buttons allows details of the molecules to be downloaded in several formats. Click on the buttons to select the fields you want included in the output file, and the format of the file. Click Download to save the file.
+
+### **The Structures Tab**
+
+The content displayed in this tab depends on the type of object or event selected in the pathway diagram. For Reactions it will display equivalent reaction diagrams from the [Rhea]() database if available:
+
+
+
+The panel has a link top-left to details in the Rhea database.
+
+The molecular structures are linked to the [ChEBI]() database:
+
+
+
+For Proteins or pathway objects that represent sets or complexes, corresponding structure information from [PDBe]() is represented if available:
+
+
+
+For simple molecules, the structures tab represents information from the ChEBI database:
+
+
+
+### **The Expression Tab**
+
+This tab represents expression information obtained from the [Expression Atlas](). Note that at the moment, only information from the first 50 genes in the pathway is displayed or available for download from the Expression Atlas widget.
+
+
+
+### **The Analysis Tab**
+
+This tab displays the results of analyses – see the section Reactome Tools for further details
+
+### **The Downloads Tab**
+
+The Downloads Tab contains buttons to start a download of the currently displayed pathway in several human-readable or computationally reusable formats.
+
+
+
+### Get Started:
+
+### **Exercises:**
+
+ 1. Find the reaction ‘Activated type I receptor phosphorylates SMAD2/3 directly’. What pathway does it belong to?
+ 2. In which cellular compartment does this reaction take place?
+ 3. What is the GO molecular function associated with the catalyst?
+ 4. What reference has experimental verification of this reaction?
+ 5. Is this reaction predicted to occur in Canis familiaris? In C. elegans?
+ 6. Is this event likely to occur in liver?
+ 7. Are 3D structures available for TGF-beta1?
+
+### More info:
+
+**Pathway Description tab:**
+
+The Description tab for a pathway includes:
+
+ * pathway name
+ * species
+ * stable identifier
+ * pathway summation
+ * a drop-down menu to select species for which the pathway is computationally predicted to be conserved, where appropriate. A description of the inference process can be found here.
+ * the GO biological process term for the pathway, where appropriate. Expanding the panel provides a link-out to GO.
+ * literature references
+
+**Reaction Description tab:**
+
+The Description tab for a reaction includes:
+
+ * reaction name
+ * a stable identifier
+ * pathway summation
+ * links to external identifiers, where appropriate. For instance, a reaction that describes a binding event may link out to IntAct records that corroborate the interaction.
+ * Input/Output: Identifies the input/output molecules, sets or complexes for this reaction. Icons to the right of these named items link to further information in Reactome.
+ * Catalyst (when relevant): The protein or complex that catalyzes the reaction. The Gene Ontology molecular function term that represents the activity of a catalyst or transporter within the reaction will be listed. If the catalyst is a complex, the component that enables the reaction to occur will be identified as the ‘active unit’.
+ * Preceding event(s): A list of events that occur immediately before the event being viewed.
+ * Following event(s): A list of events that occur immediately after the event being viewed.
+ * Cellular compartment:
+ * Inferred from another species (when relevant): This indicates that event has not been experimentally demonstrated in humans, but has been inferred on the basis of data acquired for another species.
+ * Clicking on the ‘+’ button to the right of the reaction name in this panel reveals the summation and literature references for the supporting reaction in the other species;
+ * clicking on the reaction icon to the left of the reaction name in this panel switches the user to the corresponding pathway of the species from which the human reaction has been inferred. Note that the manually curated event in the non-human species is not currently displayed in this pathway; instead, a computationally-inferred reaction is shown. This will be addressed in future updates. The human representation of the pathway can be restored either by clicking on the back button of the browser or by re-selecting ‘Homo sapiens’ in the species selector in the top panel of the Reactome website.
+ * Computationally inferred to: Links to descriptions of the events in other species that are either confirmed to occur in a very similar way in both species, or have been electronically inferred.
+ * Positively (or Negatively) regulated by: The protein or complex that regulates the reaction.
+ * References that contain experimental data verifying the reaction, with a link-out to PubMed, when applicable.
+ * Authors: The expert biologists that contributed materials that allowed this reaction to be created in Reactome.
+
+ * Reviewers: The expert biologists that verified the content for this pathway.
+
+**Physical Entity Description tab:**
+
+The details of any protein, small molecule, complex or set represented in a pathway diagram can be displayed in the Details panel by selecting the object within the diagram. Clicking a ‘+’ or ‘-‘ icon associated with the different displayed fields within the Detail panel tabs will show or hide more information about the protein, small molecule, complex and set.
+
+The Description tab may display include the following information depending upon what physical entity is selected:
+
+ * Link to corresponding entries in other databases: Cross-reference to identifiers used for this molecule in external reference databases, with hyperlinks to the record.
+ * Cellular compartment: The cellular compartment that contains this molecule, with hyperlink to the corresponding GO term.
+ * Computationally inferred orthologues: Lists equivalent molecules in other species if these have been inferred to exist. A description of the inference process can be found here.
+ * Components: If a complex is selected in the diagram, its components are listed here.
+ * Produced by: If a complex is selected in the diagram, this tab in the details panel will list all the reactions in Reactome that produce it as an output. Clicking on the ‘+’ reveals summation and literature references for these reactions.
+ * Consumed by events: If a complex is selected in the diagram, this tab in the details panel will list all reactions in Reactome that use it as an input. Clicking on the ‘+’ reveals summation and literature references for these reactions.
+
+**Download tab:**
+
+The Download tab provides options for the user to download different files types for the selected pathways. These files include:
+
+ * SBML: An exchange format used by systems biologists for their models.
+ * SBGN: An exchange format used to represent pathway and network diagrams.
+ * BioPAX2: An exchange format used by systems biologists for their models.
+ * BioPAX3: An exchange format used by systems biologists for their models.
+ * PDF: Text dump of the pathway, organized to look like a research report.
+ * Word: A document format compatible with Microsoft and other word processor software.
+ * Protege: A format used for ontology exchange.
+
+[SBML](),[ SBGN]() and[ BioPAX]() are exchange formats of interest to bioinformaticians.[ Word]() and[ PDF]() are familiar document formats, providing you with a convenient "document" of the pathway.[ Protégé]() is an extensible, platform-independent environment for creating and editing ontologies and knowledge bases, this download is likely to be of interest to those wishing to extend Reactome functionality.
+
+The complete Reactome textbook of biological processes in PDF or RTF format, the complete set of human reactions in Reactome (in SBML or BioPAX level 2 or 3 format), and a list of human protein-protein interaction pairs are available to download from the Download page, linked to the Menu Bar on the Reactome homepage.
+
+To find out about how Reactome generates SBML, see[ SBML At Reactome]().
+
+For general information about SBML,[ click here](). For more information about BioPAX[ click here]().
diff --git a/projects/website-angular/content/documentation/userguide/diseases.mdx b/projects/website-angular/content/documentation/userguide/diseases.mdx
new file mode 100644
index 0000000..0bdf099
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/diseases.mdx
@@ -0,0 +1,206 @@
+---
+title: Diseases
+category: "documentation"
+---
+
+## Diseases
+
+Reactome can annotate and display pathways associated with disease. Reactome disease annotations include cancer, metabolic and immune disease as well as infectious diseases, among others. Where possible, Reactome disease pathways also include the interaction of relevant therapeutic drugs. (Disease pathways are contained in a separate top-level Chapter in the hierarchy, and are designated with a red “+” symbol to the left of the pathway name.)
+
+Pathway Diagrams:
+
+Disease events or processes are shown in the context of the normal pathway diagram to provide context to the disease event.
+
+Disease events have red connecting lines; abnormal molecules involved in these processes are outlined in red.
+
+For infection processes, events involved in the life cycle of the infecting agent and host interactions with the infecting agent have red connecting lines. Molecules derived from the infecting agent are outlined in red, while those derived from the host are outlined as normal in black.
+
+Disease events may be loss-of-function or gain-of-function compared to the normal process.
+
+For loss-of-function events (where a protein has lost all or most of its functional activity) the disease reaction is automatically overlaid on top of the corresponding normal reaction. The loss-of-function disease entity is outlined with a dashed red line, and products that are no longer made are greyed out with a superimposed red cross. These loss-of-function events represent “stop points” in the pathway.
+
+
+
+For gain-of-function events (where a protein has acquired a novel function not performed by the wild-type protein), disease events are represented alongside the normal events.
+
+
+
+When a gain-of-function mutant performs a normal function (either a higher rate or efficiency, or after being activated in an novel manner, for instance), these disease events are overlaid on the corresponding normal event in the pathway diagram.
+
+
+
+Infectious disease events (which don’t occur in the absence of the infectious agent and therefore have no normal counterpart) have their own diagrams.
+
+
+
+Drug annotations:
+
+Where applicable, the effect of drugs on disease pathways is annotated.
+
+
+
+Details panel:
+
+The details panel for a disease pathway/event has a number of extra panels in addition to those displayed in a normal pathway/event.
+
+Disease tag:
+
+Disease events and entities are tagged with a disease term, taken from the [Disease Ontology]() and displayed in the “Disease” panel.
+
+Individual proteins are labeled with all the relevant disease terms, while sets of disease entities are tagged with the most specific term that represents all of the set members.
+
+
+
+Where possible, disease proteins are linked to [OMIM]() (Online Inheritance in Man),
+
+
+
+and cancer variants are linked to [COSMIC]() (Catalogue of Somatic Mutations in Cancer).
+
+
+
+Functional status:
+
+As described above, disease events are tagged as either gain- or loss-of-function, and this is displayed in the “Functional Status” panel.
+
+
+
+Normal Pathway/Reaction:
+
+Where applicable, the corresponding normal pathway or reaction is identified in the details panel of the disease event.
+
+
+
+Disease entities:
+
+Disease entities are annotated in molecular detail. Changes to protein sequence that arise as a result of variation in DNA sequence in disease are displayed in the details panel when a disease entity is highlighted in the diagram. Missense, nonsense, frameshift, deletion and fusion mutations are all annotated. See “More info”, below, for details of these annotations.
+
+Primary references describing the identification of the mutants proteins are annotated, but this information is not currently displayed on the website. Note also that on the website, these genetic modifications are displayed in the same panel as post-translational modifications such as phosphorylations, and are therefore (mis)labeled as ‘Post-translational modifications’.
+
+Missense mutation:
+
+
+
+Deletion mutation:
+
+
+
+Drugs:
+
+Drugs are cross-referenced to [ChEBI]() and to [IUPHAR]() where possible and applicable, and this information is displayed in the details panel when a drug is highlighted in the diagram.
+
+Get started:
+
+**Example 1 - drug-target interaction in disease** :
+
+The cystic fibrosis transmembrane conductance regulator (CFTR) is a low conductance chloride-selective channel that mediates the transport of chloride ions in human airway epithelial cells. Chloride ions plays a key role in maintaining homeostasis of epithelial secretions in the lungs. Defects in CFTR can cause cystic fibrosis (CF), resulting in an ionic imbalance that impairs clearance of secretions, not only in the lung, but also in the pancreas, gastrointestinal tract and liver. More than 1500 mutations in the CFTR gene have been identified.
+
+Gain-of-function events involving CFTR F508del:
+
+Deletion of phenylalanine 508 in CFTR is the most prevalent mutation causing cystic fibrosis. F508 deletion causes destabilization and subsequent targeting for co-translational degradation by the ER-associated degradation machinery (ERAD). F508del is ubiquitinated by ERAD-associated E3 ligases including RNF5 and RNF185, targeting it for VCP-mediated retrotranslocation and 26S proteasomal degradation. These events are novel for the mutant protein and are represented in the disease diagram “ABC transporter disorders”.
+
+
+
+Loss-of-function events for CFTR mutants:
+
+Some loss-of-function CFTR mutants are properly transported to the plasma membrane but are unable to transport chloride ions to the extracellular space. These are represented as a loss-of-function event in an overlay of the corresponding WT reaction, and displayed in the “ABC transporter disorders” disease diagram.
+
+
+
+Drug interactions in Cystic Fibrosis:
+
+CF patients with a particular mutation, G551D, have shown lung function improvements when given the drug Ivacaftor. Reactions showing ivacaftor binding the mutant protein (top right, above) and the following reaction (centre) showing channel functionality restored are displayed in the disease diagram.
+
+**Example 2 - diseases caused by accumulation of substrate over time** :
+
+Special case in Reactome where the reaction describing a particular substrate's metabolism occurs in normal physiology but over time, accumulation of the substrate leads to toxic consequences which can lead to disease. Examples are neurodegenerative diseases such as Alzheimer's and Parkinson's diseases, chronic obstructive pulmonary disease and retinal macular degeneration.
+
+The bisretinoid A2E (di-retinoid-pyridinium-ethanolamine) is a major component of lipofuschin, a yellow-brown pigment grain composed mainly of lipids but also sugars and certain metals whose accumulation is associated with degenerative diseases. In the eye, A2E is the end-product of the condensation of 2 molecules of all-trans-retinal and phosphatidylethanolamine in photoreceptor outer disc membranes. Once formed, A2E is phagocytosed, together with outer segments, to retinal pigment epithelial (RPE) cells where it accumulates. There is no evidence as yet to indicate that A2E can be catabolised.
+
+The relationship between lipofuscin accumulation and retinal degeneration is illustrated by Stargardt disease type 1. Because the reactions can be considered "normal" (they occur as part of the normal metabolism of retinoids) as well as disease-causing, they appear in the normal pathway diagram (Visual phototransduction) and coloured red.
+
+
+
+To display in the disease view of the diagram, these reactions are added as components of the disease pathway "Diseases associated with visual transduction". These reactions will now display in both normal and disease diagrams.
+
+
+
+### **Getting Started**
+
+### **Exercises:**
+
+ 1. Search for the pathway FGFR1 mutant receptor activation and open it. What disease(s) is/are associated with this pathway? (_Hint: look at the Description tab in the Details section_).
+ 2. Search for the reaction Defective MMADHC does not bind MMACHC:B12r. What type of defect has caused this loss of function? (_Hint: look at the Functional Status category for the reaction_).
+ 3. How many mutations of MMADHC are represented?
+
+More info:
+
+**Annotating genetic alteration of protein sequence for disease pathways:**
+
+The following image shows the portion of the Reactome data model describing the relationship between possible modifications to protein sequence. The subclass “GeneticallyModifiedResidue” is used for the annotation of disease entities, while “TranslationalModification” is used to annotate the consequence of processes such as phosphorylation, acylation, cross-linking and other similar non-genetic events. TranslationalModifications will not be described further here.
+
+
+
+The ReplacedResidue class is used for amino-acid substitutions. This class is also used for “simple” nonsense mutations that change a coding amino acid for a stop codon.
+
+The FragmentModification class describes more extensive changes to the coding sequence through insertions and deletions in the gene, and includes three subclasses:
+
+ * FragmentDeletionModification is used for in-frame deletions of amino-acids
+ * FragmentInsertionModification is used for in-frame insertions of amino-acids, including genomic events that result in fusion proteins
+ * FragmentReplacedModification is used for frameshifts.
+
+Examples of each annotation type are further described below.
+
+**Simple missense mutation: HHAT G287V**
+
+Hedgehog (Hh) is a secreted morphogen that regulates a number of developmental processes in vertebrates, including limb development and neural tube patterning, among others. Maturation of Hh ligand includes a number of proteolytic processing and lipid modification steps. These modifications are required for normal transit of the ligand to the surface of the secreting cell and mutations that affect these processes are associated with decreased Hh ligand secretion, abrogated Hh signaling and disease.
+
+HHAT is an O-acyltransferase that palmitoylates the N-terminal fragment of Hh. A G287V loss-of-function mutation in HHAT was identified in a rare case of Syndromic 46 XY Disorder of Sex Development, which results in testis dysgenesis. This mutant is not able to palmitoylate the Hh ligand. Details of the amino-acid substitution are displayed in the details panel when the entity is highlighted in the diagram:
+
+
+
+**Simple nonsense mutation:**
+
+The disorder “Ehlers-Danlos syndrome, musculocontractural type 1” (EDSMC1) is caused by loss-of-function mutations in the carbohydrate sulfotransferase 14 (CHST14) gene. A nonsense mutation causing this disorder is a 205A-T transversion in the CHST14 gene, resulting in a lys69-to-ter (K69*) substitution. This is represented in the details panel as “L-lysine 69 replaced with unknown”.
+
+
+
+**FragmentDeletionModification:**
+
+This class is used for in-frame deletions of amino-acids leading to internally truncated proteins. These variants are named “core protein name” [first amino acid of deletion_last amino acid of deletion]del, as shown below for PIK3R1 Y463_L466del, which has a deletion of residues Y463 to L466, inclusive:
+
+
+
+**FragmentInsertionModification**
+
+This class is used for in-frame insertions of amino-acids and for fusion proteins.
+
+**In-frame insertions:**
+
+These variants are named “core protein name” [aa prior to insertion_aa following insertion]ins[inserted aa’s]. The amino-acid string of the inserted residues is added manually to the mutant protein name, as shown below for EGFR V738_K739insKIPVAI. This represents a variant where the amino acids 739-744 (KIPVAI) from EGFR are inserted at aa 739 of EGFR, and is thus a duplication of these residues:
+
+
+
+**Fusion proteins:**
+
+The FragmentInsertionModification class is also used to annotate proteins that arise as the result of genomic changes that bring two genes together to result in a fusion protein, as in the ZMYM2-FGFR1 fusion described below. This fusion puts the ZMYM2 dimerization region (1-914) together with the kinase domain of the FGFR1 receptor (residues 429-822) and results in constitutive activation of the kinase domain by virtue of ligand-independent dimerization (for reference, the full length aa sequence of these two proteins are 1-1377 and 1-822 for ZMYM2 and FGFR1, respectively).
+
+By convention, the N-terminal most partner of the fusion is set as the reference protein for the variant protein while the C-terminal fusion partner sequence is captured in the FragmentInsertionModification record, as in the example below for the ZMYM2-FGFR1 fusion:
+
+
+
+Post-translational modifications to either partner in the fusion protein are numbered according to each respective WT reference gene product and do not reflect the aa position in the fusion. For instance, if in the context of the fusion protein, the FGFR1 partner is phosphorylated at (WT FGFR1 position) Y766, the fusion EWAS would be ZMYM2-pY766-FGFR1, despite the fact that in linear sequence the phosphorylation occurs at residue 1250 of the fusion (913+(766-429)).
+
+**FragmentReplacedModification**
+
+This class is used to annotate frameshift mutations that alter the amino-acid sequence of the protein as in the case of EXT1, described below.
+
+The disorder “Hereditary multiple exostoses 1” (EXT1) is caused by loss-of-function mutations in Exostosin 1 (EXT1). One such mutation is a 1-bp deletion at nucleotide 1469 in the EXT1 gene, resulting in a frameshift mutation with a premature stop codon nine amino acids downstream. Variants of this type are named “core protein name” [aa prior to insertion_fs*(number of aa to the first stop codon)]. The novel amino acids that occur as a result of the frameshift are identified in the mutant protein record, as shown below for EXT1 L490Rfs*9:
+
+
+
+For cancer-related processes, it is useful to view the altered disease events alongside their normal counterparts in the same diagram. The user can then see where normal processes diverge into ones that are implicated in cancer.
+
+For metabolic processes, proteins that have lost all or most of their functional activity towards a substrate causes the majority of defects. The diagram displays these disease reactions as 'stop points' in the pathway. Defective enzyme catalysts are outlined by a red dashed line; products that are no longer made are shown greyed-out with a superimposed red cross.
+
+
diff --git a/projects/website-angular/content/documentation/userguide/pathway-browser.mdx b/projects/website-angular/content/documentation/userguide/pathway-browser.mdx
new file mode 100644
index 0000000..59e7f9a
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/pathway-browser.mdx
@@ -0,0 +1,203 @@
+---
+title: The Pathway Browser
+category: "documentation"
+---
+
+## The Pathway Browser
+
+The Pathway Browser is the primary means of viewing and interacting with pathways in Reactome.
+
+See our Youtube Video explaining [navigating the Pathway Browser]() for more information.
+
+The Pathway Browser includes tools for analysing datasets and exploring pathways. These tools allow several types of analysis:
+
+ * Pathway over-representation analysis and pathway topology-based analysis
+ * Comparison of a pathway with its equivalent in another species
+ * The overlay of user-supplied expression data onto a pathway
+ * The overlay of protein-protein or protein-compound interaction data from external databases or user-supplied data onto a pathway
+
+See the section [Reactome Analysis Tools ]()for details of the analysis tools.
+
+The Pathway Browser is launched by clicking the Browse Pathways button on the Homepage:
+
+
+
+When the Pathway Browser opens, it displays an overview of all Reactome pathways. Pathways are organized hierarchically and often have sub-pathways. At the highest hierarchical level, all pathways are represented in an Overview. At intermediate levels, pathways are often represented as interactive illustrations, with selectable regions that act as links to the lower levels, which are represented as detailed Pathway Diagrams. Reactome’s Overview uses a unique graphical visualization of the hierarchy, representing pathways at the uppermost hierarchical level as central nodes (circles) with subpathways arranged concentrically (in rings) around them. Many have further concentric rings of ‘child’ subpathways. Pathway nodes are connected to their subpathways by edges (lines). This overview displays all pathways and allows rapid zooming to details. As you zoom in, labels appear for the subpathway nodes. This view is also used to display data analysis results.
+
+Nodes in the overview link to detailed pathway diagrams that use a style based on Systems Biology Graphical Notation. They have several features intended to ensure an appropriate level of information detail, including subpathway shading and fade-in of labels, commonly occurring small molecules and structures.
+
+To see the details of a specific pathway, select and click (or double-click) the node representing the pathway. Alternatively select and click on the name in the Pathway Hierarchy panel on the left.
+
+Features of the Pathway Browser are labelled in the diagram below.
+
+
+
+**Home** – the Reactome logo is a button linked to the homepage.
+
+**Species** – Reactome is a database of curated human biological pathways. These human pathways are used to computationally infer equivalent pathways in model organisms (described in detail [here]()). Use the Species drop-down to select a species and view the predicted pathway. Note that infectious disease pathways that show the interaction of pathogens with human proteins are included in Homo sapiens.
+
+**Layout** – The pathway browser is divided into 3 main sections. These are: the Hierarchy Panel, on the left, the Details Panel, bottom right, the Pathway Panel, top right. The layout buttons allow you to show or hide these panels.
+
+**Analyse Data** – Opens a panel where analysis can be performed.
+
+**Key** – this opens a key explaining the objects present in the pathway panel.
+
+**Zoom/Move Toolbar** \- Click on the arrows to move the diagram. Click on the plus and minus buttons to zoom in and out. Alternatively, click and drag the diagram, and use your mouse scroll wheel to zoom.
+
+**In-Diagram Search** – click the button to open the search panel. When the Overview or a Pathway Diagram is displayed, objects represented in the displayed diagram or all diagrams are searchable.
+
+**Fit to Page** \- resizes the Pathway Overview or Diagram to fit the available space.
+
+**Open Diagram** – when a pathway node is selected in the Pathway Overview, this button opens the corresponding Pathway Diagram
+
+**Illustrations** – some Reactome pathways have a corresponding high-quality graphic. Select this button to view.
+
+**Export** – use this button to download the visible region of the Pathway Overview or Diagram, including any analysis overlay, as a PNG file or upload it to GenomeSpace.
+
+**Thumbnail** – Provides an overview of the entire pathway when zoomed in so that only a region of the pathway is displayed.
+
+**Pathway Panel** \- This is where Pathway Diagrams are displayed when they are selected in the Pathway Hierarchy or in search results. If no pathway is selected, this panel displays a brief Reactome help guide.
+
+**Details Panel** \- Contains details of objects when they are selected in the Pathway Diagram. The content depends on the type of object, i.e. pathway, reaction, complex, set, protein or small molecule (see the Details section).
+
+**Event Hierarchy Panel** – Shows the hierarchical organisation of Reactome pathways and pathway events or steps, known as reactions. When the Pathway Browser is opened, only the top level of the hierarchy is shown, as an alphabetical list of Reactome’s main topics (superpathways). Most topics are split into sub-pathways, which may be further divided into sub-sub-pathways. Most topics in Reactome are organised in this hierarchical manner. Sub-pathways can be revealed by clicking on the + icon to the left of the pathway name, and hidden by clicking on the - icon.
+
+**Settings Sidebar** – Used for context-sensitive help, to configure colour schemes and the default source of interactors.
+
+## Event Hierarchy
+
+The order of reactions from top to bottom in the Event Hierarchy Panel usually follows their order as steps in the pathway so that the preceding reaction is above and the subsequent reaction below, but this is not always the case. Pathways may not be linear; they can be circular or branched. Consequently, reactions often have multiple preceding or subsequent reactions. To view a complete list of the events that precede or follow a reaction, refer to the Preceding and Following Events section in the Details Panel, as described in the Details Panel chapter below.
+
+The event hierarchy panel provides a nested structure containing large topics that are organised as pathways and subpathways and can be expanded to show individual events and steps. The icons next to the event describe the event.
+
+You can hover over the icon in the Event Hierarchy panel to show a tool-tip that describes the event, as illustrated in the figure below.
+
+
+
+This table shows a legend of the icons and their respective descriptions in Reactome:
+
+
+
+## **Pathway Diagrams**
+
+Pathway Diagrams represent pathways as a series of connected molecular events, known in Reactome as ‘reactions’, which can be considered as steps in the pathway.
+
+Cellular compartments are represented as pink/orange boxes with a double boundary. A typical diagram has a box to represent the cytosol, bounded by a double-line that represents the plasma membrane. The white area outside this box represents the extracellular space. Other organelles are represented as additional labelled boxes within the cytosol.
+
+Molecules, represented in diagrams as blue/green boxes, are placed in the physiologically-correct cellular compartment, or lie on the boundary of a compartment to indicate that they are in the corresponding membrane, e.g. a plasma membrane protein will be placed on the boundary of the cytosol.
+
+Reactions typically include:
+
+ * Input and output molecules, and when relevant a catalyst (see diagram A below)
+ * Inputs, outputs and catalyst are represented as boxes or ovals
+ * Green boxes with rounded corners are proteins
+ * Green boxes with square corners are proteins that have no UniProt accession (or did not at the time the reaction was created).
+ * Green ovals are small molecules or sets of small molecules
+ * Blue boxes with a double boundary are sets, i.e. proteins or small molecules that are functionally equivalent.
+ * Blue boxes with cut corners are complexes, i.e. proteins and/or other molecules that are bound in a multimolecular entity.
+ * Green boxes with a white inner box are sub-pathways.
+ * Reaction input and output molecules are joined by lines to a central ‘reaction node’ (surrounded by a green box in Figure A below). Clicking this node selects the reaction.
+ * The outputs of a reaction have an arrowhead on the line connecting them to the reaction node.
+ * Reaction inputs/output molecules are often connected by arrows to preceding or subsequent reactions (i.e. the preceding/subsequent steps in the pathway).
+ * Catalysts are connected to the reaction node by a line ending in a circle.
+ * Numbered boxes on the line between an input/output and the reaction node indicates the number of molecules of this type in the reaction (when n >1).
+ * Molecules that regulate a reaction are connected to the reaction node by a line ending in an open triangle for positive regulation or a ‘T’-shaped head for negative regulation (see B below).
+ * A white box labelled P on the boundary of an object indicates a phosphorylation event.
+ * Proteins or small molecules that are also part of a displayed set may be connected to the set object by a line of short dashes.
+ * Sets with overlapping content may be connected by a line of long dashes.
+ * Reactions that represent a disease process use red connecting lines. Objects associated with a disease are bordered in red.
+
+
+
+## **Enhanced high-level diagrams and Subpathway icons**
+
+Many pathways at the higher levels of the hierarchy are too large to represent as a single, detailed pathway diagram. Instead they are often represented as an illustration, known as an enhanced high-level diagram, which graphically represents the subpathways as selectable regions of an illustration. When the mouse pointer is moved over a region, it becomes surrounded by a blue highlighting 'halo'. When you click on a region it becomes selected and outlined in dark blue, the corresponding subpathway is selected in the Hierarchy panel and the Details panel updates to show details of the subpathway. Double-clicking on a region opens the corresponding detailed Pathway Diagram.
+
+In the Hierarchy panel, pathways that have an enhanced high-level diagram have a blue icon to the left of their name.
+
+
+
+A few higher level pathways use an older visualization of large topics, where subpathways are represented by a box with a green boundary, the subpathway icon (see example below). Selecting a subpathway icon has the same result as selecting a region of an enhanced-level pathway diagram as described above.
+
+
+
+In some pathways, Reactome links to other related pathways through green boxes. In order to differentiate between related subpathways that are children of the same parent pathway (subpathways) and related subpathways that are children of a different parent pathway (interacting pathways), the Pathway Browser uses slightly different glyphs as illustrated in the figure below.
+
+
+
+### **Subpathway Shading**
+
+When a Pathway Diagram contains subpathways, the area containing each subpathway is overlaid with a coloured box, labelled with the subpathway name, to help locate it in the larger diagram. Below is the diagram for ERBB4 signaling. The four subpathways are represented as 4 boxed and shaded areas.
+
+
+
+## **Pathway Diagram zoom detail level**
+
+The pathway diagram represents different levels of detail, depending on the zoom level. When fully zoomed-out, regions corresponding to subpathways are shaded. The boxes representing molecules or groups of molecules have no text labels and trivial molecules, such as water and ATP, are not represented. At the next level of zoom, subpathway shading disappears, while trivial molecules and the circular red icon indicating that a protein has known interactors fade into view. At closer levels of zoom the reaction nodes appear, as do boxes containing a number on the lines connecting molecules to the reaction node if stoichiometry is greater than one. At this level of zoom, if interactors are displayed their names appear. At the highest level of zoom, if available, proteins, small molecules and interactors display structural diagrams from PDBe or ChEBI with associated details.
+
+
+
+
+
+
+
+
+
+## **Navigating Pathway Diagrams**
+
+The Pathway Diagram, Hierarchy and Details Panels are interactively connected. Selecting an entity (molecular object) or event in the diagram will reveal relevant information in the Details Panel. Clicking a pathway name in the Hierarchy will open the corresponding Pathway Diagram in the Pathway Diagram Panel, and reveal details of the pathway in the Details Panel. If a subpathway is contained within the diagram that is displayed, clicking its name in the hierarchy will cause all the reactions in the pathway to be highlighted in blue. In the figure below, the diagram for Platelet homeostasis is displayed in the Pathway Panel, and the sub-pathway Prostacyclin signalling through prostacyclin receptor has been selected in the Hierarchy, causing all the reactions in this sub-pathway to be highlighted (blue) on the Pathway Diagram.
+
+
+
+Similarly, selecting a reaction in the Hierarchy causes that reaction to be highlighted in the pathway diagram and updates the Details Panel to show details of the reaction. If the reaction is not in the visible region of the diagram, the view will re-centre and zoom to show it. If instead of clicking an event, you hover the mouse pointer over its name, it will be highlighted in yellow. In the figure above, the sub-pathway Platelet calcium homeostasis is highlighted in yellow.
+
+### **Getting Started**
+
+### Hierarchy Panel **Exercises**
+
+This exercise is to check that you understand the organisation of the Hierarchy Panel. You don’t need to look at the Diagram Panel.
+
+From the Home page, search for PDGF signaling.
+
+In the results page, open the expandable hierarchy in the section Locations in the Pathway Browser by clicking on the + button. **DON’T** click on the location before expanding the hierarchy! This will open the pathway diagram in the Pathway Browser.
+
+Look at the Hierarchy Panel on the left.
+
+ 1. How many sub-pathways does this pathway have?
+ 2. How many reactions are in the first sub-pathway?
+ 3. What reaction(s) follow ‘Translocation of PDGF from ER to Golgi’? _Hint:Look in the Details Panel, bottom-right. If it is hidden, use the layout buttons in the top right corner, above the diagram, to reveal it._
+
+### Subpathway **Exercises**
+
+This exercise is to check that you understand Subpathway icons and the relationship between the Pathway Hierarchy and Pathway Diagram.
+
+Open the pathway Hemostasis in the Pathway Browser.
+
+Select the subpathway ‘Platelet Hemostasis’.
+
+ 1. What happens if you hover your mouse pointer in the Hierarchy on the sub-pathway ‘Platelet calcium hemostasis’?
+ 2. What happens if you click in the Hierarchy on the sub-pathway ‘Platelet calcium hemostasis’?
+ 3. What happens if you click in the hierarchy on the reaction ‘Binding of ATP to P2X receptors’?
+
+**More Information**
+
+There are 5 subtypes of reaction node, indicating reaction subclasses.
+
+
+
+ * Open squares represent ‘transition’, i.e. a change of state that is not one of the defined subclasses.
+ * Solid circles represent ‘association’, i.e. binding
+ * Double-bordered circles represent ‘dissociation’
+ * A square with two slashes represents an ‘omitted process’. This is used when the full details of a reaction have been deliberately omitted. This is most commonly used for events that include representative members of a large family to illustrate the general behaviour of the group. It can be used for reactions that occur with no fixed order or stoichiometry, or for degradation events where the output is a random set of fragments.
+ * Squares containing a question mark represent an ‘uncertain process’, where some details of the reaction are known, but the process is thought to be more complex than represented. Explanatory details are typically included in the Description.
+
+### Reaction Node **Exercises**
+
+This exercise is to check that you understand reaction node subtypes.
+
+From the Home page, search for the pathway ‘Effects of PIP2 hydrolysis’ and open it in the Pathway Browser (in any of the several locations)
+
+ 1. What symbol represents the reaction for ‘Binding of IP3 to the IP3 receptor’?
+ 2. What symbol represents the reaction ‘IP3R tetramer:I(1,4,5)P3:4xCa2+ transports Ca2+ from platelet dense tubular system to cytosol? What subtype of reaction is this?
+ 3. Open the subpathway ‘Arachidonate production from DAG’. What is the name of the catalyst for ‘2-AG hydrolysis to arachidonate by MAGL’? Can you name the three outputs of this reaction?
+ 4. Can you find the UniProt ID for the catalyst? Hint: There are several ways to find this, two require you to select something!
diff --git a/projects/website-angular/content/documentation/userguide/reactome-fiviz.mdx b/projects/website-angular/content/documentation/userguide/reactome-fiviz.mdx
new file mode 100644
index 0000000..770b6a2
--- /dev/null
+++ b/projects/website-angular/content/documentation/userguide/reactome-fiviz.mdx
@@ -0,0 +1,842 @@
+---
+title: ReactomeFIVIz
+category: "documentation"
+---
+
+## ReactomeFIVIz
+
+## Contents
+
+ * [1. Overview](<#Overview>)
+ * [2. Download and Launch ReactomeFIViz](<#Download_and_Launch_ReactomeFIViz>)
+ * [3. Use Reactome Pathways](<#Use_Reactome_Pathways>)
+ * [3.1. Explore Reactome Pathways](<#Explore_Reactome_Pathways>)
+ * [3.2. Display Reactome Pathways in the FI Network View](<#Display_Reactome_Pathways_in_the_FI_Network_View>)
+ * [3.3. Pathway Enrichment Analysis](<#Pathway_Enrichment_Analysis>)
+ * [3.4. Probabilistic Graphical Model Based Pathway Analysis](<#Probabilistic_Graphical_Model_based_Pathway_Analysis>)
+ * [3.5. Boolean Network Based Pathway Analysis](<#Boolean_Network_based_Pathway_Analysis>)
+ * [3.6. Visualization of Structural Variants in the Context of Reactome Pathways](<#Structural_Variants_Visualization>)
+ * [4. Use the Reactome Functional Interaction (FI) Network](<#Use_the_Reactome_Functional_Interaction_.28FI.29_Network>)
+ * [4.1. Gene Set/Mutation Analysis](<#Gene_Set.2FMutation_Analysis>)
+ * [4.2. PGM Impact Analysis](<#PGM_Impact_Analysis>)
+ * [4.3. Microarray Data Analysis](<#Microarray_Data_Analysis>)
+ * [5. Visualize Drugs in the Contexts of Reactome Pathways and FI Network](<#Visualize_Cancer_Drugs_in_Contexts_of_Reactome_Pathways_and_FI_Network>)
+ * [5.1. Visualize Cancer Drugs in Reactome Pathways](<#Visualize_Cancer_Drugs_in_Reactome_Pathways>)
+ * [5.2. Visualize Cancer Drugs in the FI Network](<#Visualize_Cancer_Drugs_in_the_FI_Network>)
+ * [5.3. Visualize Drug Central Drugs](<#Visualize_DrugCentral_Drugs>)
+ * [5.4. Simulate Impact Drugs on Pathway Activities](<#Simulate_Impact_of_Cancer_Drugs_on_Pathway_Activities>)
+ * [6. Perform scRNA-seq Data Analysis and Visualization](<#scRNA_seq>)
+ * [6.1. Standard Analysis via scanpy](<#scanpy_analysis>)
+ * [6.2. RNA Velocity Analysis via scVelo](<#scvelo_analysis>)
+ * [7. Other Features Related to the FI Network](<#Other_Features_Related_to_the_FI_Network>)
+ * [7.1. Query FI Source](<#Query_FI_Source>)
+ * [7.2. Fetch FIs for Node](<#Fetch_FIs_for_Node>)
+ * [7.3. Show Pathway Diagram](<#Show_Pathway_Diagram>)
+ * [7.4. Load Cancer Gene Index Annotations](<#Load_Cancer_Gene_Index_Annotations>)
+ * [7.5. Survival Analysis](<#Survival_Analysis>)
+
+### Overview
+
+The [ReactomeFIViz]() app is designed to find pathways and network patterns related to cancer and other types of diseases. This app accesses the [Reactome]() pathways stored in the database, help you to do pathway enrichment analysis for a set of genes, visualize hit pathways using manually laid-out pathway diagrams directly in Cytoscape, and investigate functional relationships among genes in hit pathways. The app can also access the Reactome Functional Interaction (FI) network, a highly reliable, manually curated pathway-based protein functional interaction network covering over 60% of human proteins, and allows you to construct a FI sub-network based on a set of genes, query the FI data source for the underlying evidence for the interaction, build and analyze network modules of highly-interacting groups of genes, perform functional enrichment analysis to annotate the modules, expand the network by finding genes related to the experimental data set, display pathway diagrams, and overlay with a variety of information sources such as cancer gene index annotations. Recently we have also added features to help users visualize FDA-approved cancer drugs in the contexts of the FI network and Reactome pathways, and use Boolean network models directly built from Reactome pathways to investigate potential functional impacts of displayed cancer drugs.
+
+For an example how we use Reactome FIs for cancer data analysis, please see our publication: [A human functional protein interaction network and its application to cancer data analysis]().
+
+### Download and Launch ReactomeFIViz
+
+ReactomeFIViz app 6 needs Cytoscape 3.7.0 or above. If you have not installed Cytoscape 3.7.0 or above, please download it from Cytoscape's web site: [http://www.cytoscape.org](). After launching Cytoscape, use menu "Apps/App Manager" to open the "App Manager" dialog, and search for "ReactomeFI". You should see the ReactomeFIViz app listed in the middle panel (See the Figure below. You may see a different version number. **Note: The listed name of this app is "ReactomeFIPlugIn"** , which is the original name of the app.). Choose the app, and then click the "Install" button at the bottom of the dialog. Follow the procedures to finish the installation.
+
+
+
+Install ReactomeFIViz app From App Store
+
+### Use Reactome Pathways
+
+Using the pathway visualization and analysis features, you can load pathways in the Reactome database into Cytoscape, visualize Reactome pathways in either the native pathway diagram view or the FI network view, do pathway enrichment analysis for a set of genes, and check genes from your list in hit pathways.
+
+#### Explore Reactome Pathways
+
+ 1. Load Reactome pathways: Use menu "Apps/Reactome FI/Reactome Pathways" to load pathways into Cytoscape. The loaded pathways are organized in a hierarchical way as in the Reactome web application ([https://reactome.org/PathwayBrowser/]()), and listed in the left side "Control Panel" in the tab called "Reactome".
+
+
+
+Reactome Pathways
+
+ 2. View pathways in Reactome: After selecting a pathway in the pathway hierarchy, you can choose "View Reactome Source" from the popup menu (right click in Windows or Control-click in Macs to get the popup menu)
+
+
+
+Pathway Popup Menu
+
+to view its detailed annotation in Reactome. Or you can choose "View in Reactome" to view the detailed information in the Reactome web application.
+**Note** : The ancestor pathways (container pathways) for a selected pathway are displayed in the middle panel, "Selected Event Branch", in the Reactome tab. You can click an ancestor pathway in this middle panel to view the clicked pathway's location in the original pathway hierarchical tree. However, the ancestor pathway will not be selected in the original tree. This is a designed behavior to keep the selection in the original tree.
+ 3. Search pathways: Choose "Search" in the popup menu to bring up the search dialog. The found pathway(s) will be highlighted in blue in the pathway tree.
+**Note** : Search will be against all loaded pathways, not limited to the selected pathway and its contained sub-pathways.
+
+
+
+Search Pathways
+
+ 4. Open Reactome Reacfoam: The Reacfoam view provides a holistic view of all (exclude disease) human pathways in the Reactome database. Choose "Open Reactome Reacfoam" in the popup menu to open the Reactome Reacfoam in the default browser.
+
+
+
+Open Reacfoam
+
+**Note** : Pressing your mouse and then holding it to a pathway box will select the pathway in the tree of ReactomeFIV automatically.
+
+
+
+Reactome Reacfoam
+
+ 5. Open pathway diagram: Pathways in Reactome are organized in a hierarchical way. Not all pathways have their own pathway diagrams. A smaller pathway (called sub-pathway) may be drawn in a bigger pathway, which has its own pathway diagram. Most of top-level pathways (called modules or super pathways) are used to organize related pathways (e.g. Disease, Signaling Transduction), and therefore contain only rectangle boxes representing canonical pathways.
+ 1. **Show Diagram** : If a selected pathway has its own pathway diagram, you can choose "Show Diagram" in the popup menu to open its pathway diagram into the central Cytoscape desktop.
+ 2. **View in Diagram** : If a selected pathway is laid-out as a sub-pathway in a bigger one, you can choose "View in Diagram" in the popup menu to view its drawing in its container pathway. Reactions contained by the selected pathway will be highlighted in blue after the diagram is opened. For example, see pathway "G1/S DNA Damage Checkpoints" opened in pathway "Cell Cycle Checkpoints" below:
+
+
+
+Pathway Diagram
+
+ 6. Search diagram: Objects displayed in a pathway diagram can be searched using "Search Diagram" from the popup menu (Right click in Windows or Control click in Macs without selecting any object in the pathway diagram to get the popup menu). The found objects will be selected and highlighted in blue.
+**Note** : Reactions will not be searched in the diagram. Use the search feature in the pathway tree to search for reactions.
+ 7. Export diagram: Displayed diagram can be exported as a PDF, JPG or PNG file. Use "Export Diagram" from the popup menu to export the displayed diagram.
+ 8. View Reactome Source or View in Reactome: Select an object, and then right-click (or control click) to get the popup menu. Choose "View Reactome Source" to view the detailed annotation for the selected object in a table (See Figure below for an example). Or choose "View in Reactome" to view the selected object in the Reactome web application.
+
+
+
+Reactome Instance View
+
+ 9. List Genes: Genes contained by a complex or protein set, or a gene related by a displayed protein can be viewed by using a menu item "List Genes" after selecting an object. For example, the following dialog shows genes contained by complex hBUBR1:hBUB3:MAD2*:CDC20. Clicking a gene symbol will bring you to the web page for that gene in the GeneCard web site.
+
+
+
+List Genes
+
+#### Display Reactome Pathways in the FI Network View
+
+ 1. Display pathway in the FI network view: A Reactome pathway can be converted into a functional interaction network using the method we have established (see [A human functional protein interaction network and its application to cancer dat analysis]()). Use "Convert to FI Network" in the popup menu brought up by right-clicking (Windows) or control-clicking (Macs) an empty area without any selection in the pathway diagram panel. The original pathway diagram will be moved to the bottom-left corner, and a new FI network will be generated based on the original pathway diagram, which will be displayed in a new network panel.
+**Note** : sub-pathways contained by the displayed pathway will be extracted into the FI network too.
+
+
+
+Pathway in the FI Network View
+
+ 2. Explore objects in the pathway and network views: Object selection in three views has been synchronized. Objects that can be selected include: events in the pathway tree view, objects in the pathway view at the bottom-left corner, and genes and FIs in the network view. You can select an object in one of three views, and corresponding objects in other two views should be selected too. Also you should use features implemented in popup menus in each individual view to explore objects as in a single view.
+
+
+**Note** : Using Cytoscape's built-in "Saving Session" feature can save the converted FI networks from pathways. However, displayed pathways cannot be saved into a session file for the time being. We will implement this function in a future release.
+
+#### Pathway Enrichment Analysis
+
+ 1. **Pathway enrichment analysis** : A list of genes can be used to check if any of Reactome pathways have been enriched. To do this, use the popup menu item, "Analyze Pathway Enrichment" (below left figure), to get the dialog for choosing a gene set file (below right figure). You can use a gene set file in one of three file formats: one gene per line, all genes in the same line and delimited by commas, or all genes in the same line and delimited by tabs. You can also manually input genes by clicking the "Click to Enter" button (one line for one gene).
+**Note** : Dependent on the size of your gene list, it may take over 1 minute for running the pathway enrichment analysis. Pathways used in this feature are different from Reactome pathways for annotating a FI network or network modules. Here all over 2,000 pathways are used. For annotation, only a subset of Reactome pathways, which have been pre-selected for a certain size, are used.
+
+**Note** : To get a holistic view of the pathway enrichment analysis results, open the Reactome Reacfoam after the analysis using the popup menu "Open Reactome Reacfoam" for the pathway tree. You may also download the Reacfoam view by clicking the download button at the top-right corner. For windows 10 users, to open the Reacfoam view, you need to allow "public" access to Cytoscape by checking "public" in the settings for "Allow an app through Windows Firewall" in the "System and Security" control settings.
+
+ 2. **View enrichment analysis results** : Pathway enrichment results are displayed as a table labeled as "Reactome Pathway Enrichment" in the "Table Panel" at the bottom of the main Cytoscape window. You can use "Views in Diagram" to view hit pathways in the pathway diagram view, and use "Export Annotations" to save the results in the table. Pathways in the Reactome pathway tree are highlighted in different colors based on their FDR values. Objects containing genes from your gene list are highlighted in a purple background with a white font in the pathway diagram view. Hit genes are displayed in a thick purple border in the FI network view for a hit pathway.
+**Note** : Hit genes are displayed with same colors in the "Gene List" dialog from the "List Genes" feature.
+
+
+
+Pathway Enrichment Results
+
+ 3. **Perform GSEA analysis** : Gene Set Enrichment Analysis ([GSEA]()) is a rank-based pathway enrichment analysis approach, widely used in pathway-based data analysis. ReactomeFIViz provides support to perform GSEA analysis for Reactome pathways using a gene score file. Gene score may be t-score from differential gene expression analysis or other type of scores that can be ranked. To perform the GSEA pathway enrichment analysis, you need to provide a tab-delimited text file containing two columns: the first for gene symbols (human only) and the second for gene scores. The first row is reserved for the column headers, and will not be imported for analysis. To perform GSEA analysis, use popup menu "Perform GSEA Analysis" in the pathway tree to bring up the GSEA configuration dialog, where you can enter the gene score file and choose the minimum and maximum size of pathways along with the permutation number.
+
+
+
+Perform GSEA Analysis
+
+
+
+Configure GSEA Analysis
+
+The GSEA analysis results are displayed in the table labeled as "Reactome GSEA Analysis" in Cytoscape Table Panel. Pathways subject to GSEA analysis in the pathway tree are highlighted based on FDR values as in the gene set-based pathway enrichment analysis (See above). For details about the meanings of columns shown in the results table, please consult the original GSEA document: [GSEA Document]().
+
+
+
+GSEA Analysis Results
+
+ 4. **Overlay Gene Scores onto Pathways** : For significant pathways produced from the GSEA analysis, you can overlay gene scores to investigate locations of products of genes having significant high or low scores, therefore to understand potential pathway activity impact caused by these extreme scores. To do this, use popup menu "Overlay Gene Scores" in pathway diagram view to choose the gene score file in the configuration dialog. After the file loading, entities in pathway diagrams will be highlighted based on scores. You may choose one or more genes in the right gene scores Table View to visualize related entities in the pathway diagram.
+
+
+
+Overlay Gene Scores
+
+
+
+Gene Score Overlay Results
+
+**Note** : If an entitiy (e.g. a complex or an EntitySet) is composed of more than one gene, the score for the entity is the mean of all genes annotated for that entity. To remove overlaid gene scores in the pathway diagram, use popup menu "Remove Gene Scores". To view the distribution of scores for genes annotated in the displayed pathway diagram, choose "Plot View" in the "Gene Scores" tab in Cytoscape Results Panel (see below).
+
+
+
+Gene Score Distribution
+
+#### Probabilistic Graphical Model based Pathway Analysis
+
+We adapted the PARADIGM approach for Reactome pathways by converting reactions drawn in pathway diagrams into factors in factor graphs, a type of probabilistic graphical models (PGMs). For details about the PARADIGM approach, see: [Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM](). For introduction to factor graphs, see this wikipedia entry: [Factor Graph](). For test purposes, you can download two sample data files for 100 TCGA ovarian cancer patients: [CNVs]( "ov.CNV.100.txt.zip") and [mRNA gene expression]( "ov.mRNA.100.txt.zip"). The original TCGA OV files were downloaded from [the Broad GDAC]()[ Firehose]() web site.
+
+ 1. **Run graphical model analysis in batch:** This feature is used to perform a batch graphical model analysis for all Reactome pathways having manual layout diagrams.
+ 1. **Start the analysis:** Choose the popup menu, "Run Graphical Model Analysis", in the pathway hierarchical tree. After choosing this menu, you will be asked to choose data files and provide parameters for inference algorithms in the following two tabs in the "Run Graphical Model Analysis" dialog:
+
+
+
+LoadData
+
+
+
+SetUpAlgorithms
+
+
+**Notes** :
+1). If you choose "Use empirical distribution" in the data loading dialog, your loaded data will be used directly to construct factor functions without discretizing. At present, we recommend to use "Choose threshold values for discretizing".
+2). It is recommended to use the default parameters for inference algorithms for a batch analysis for quick performance. You can try different parameters for some specific pathways after you find interesting pathways from the batch analysis. _**If you want to perform two-case study (e.g. case-control, drug sensitive/insensitive, etc), check the checkbox, "Used for pathway analysis for samples with two cases", and provide a sample information file as required. For two-case analysis, a random data set will not be generated. Results will be presented by comparing two types of samples in your uploaded data files.**_
+ 2. **Run the analysis:** Click the "OK" button to start the batch analysis. Depending on your sample size, it may take hours to finish the whole analysis.
+ 3. **Finish the analysis:** After the batch analysis done, you may see the following list if some of pathways cannot be analyzed because the inference algorithm cannot converge. Please make sure the following list is small (probably less than 10 pathways) so that you can get enough results.
+
+
+
+FailedPathwaysList
+
+ 4. **View the results:** The results from the batch analysis are displayed at the bottom table panel of the Cytoscape desktop as the following:
+
+
+
+BatchResults
+
+**Note** : There are 7 columns in this table: ReactomePathway for pathway names analyzed by the App; AverageUpIPA shows how much a pathway is up-perturbated by comparing to a random background (IPA: integrated pathway activity. See the above PARADIGM paper for details); AverageDownIPA shows how much a pathway is down-perturbated; CombinedPValue is a p-value indicating how significant this pathway is perturbed based on pathway outputs and the Fisher's method; MinimumPValue is the minimum p-value for pathway outputs; the last two columns are FDRs for two p-values based on the Benjamini–Hochberg method. AverageUpIPA or AverageDownIPA may be NaN, which indicates there is no detected up or down perturbation based on this analysis. The FDR filtering works based on the FDR values displayed in the last two columns with "OR" operation.
+ 5. **Save the results:** To keep the results, use the popup menu in the table, "Export Annotations", to save the results into an external text file. The saved results can be loaded later on by using popup menu, "Load Graphical Model Results", in the pathway tree.
+ 2. **Run graphical model analysis for a single pathway:** This feature is used to perform a graphical model analysis for a pathway displayed in the Cytoscape desktop.
+ 1. **Open a pathway:** As before, you can choose a pathway in the pathway tree, and open its diagram in the Cytoscape desktop. Or you can choose an interesting pathway from the batch analysis results table by choosing popup menu, "View in Diagram".
+ 2. **Start the analysis:** Choose popup menu, "Run Graphical Model Analysis", from the popup menu list in the pathway diagram window. You will be asked to provide data files and set up inference algorithms as in the batch analysis. After clicking the "OK" button, you will be asked to provide a list of escape names for entities in the pathway that will not be considered in the graphical model (e.g. ATP, ADP, etc) in the following dialog:
+
+
+
+EscapeNameDialog
+
+
+**Note** : If you have loaded data files, you may choose to use the loaded data files without displaying the data loading dialog.
+ 3. **View the results:** After the analysis is done, three tabs are displayed in the table pane of Cytoscape: IPA Pathway Analysis, IPA Sample Analysis and IPA Node Values. IPA Pathway analysis displays inference results for entities in the pathway by comparing samples in your data files and in a random data set generated dynamically by the App based on your data files. IPA Sample Analysis shows results for each individual samples as up or down perturbation. You may choose to show/hide p-values and FDR values for samples in the table. IPA Node Values show inference results for selected entities in the pathway diagram for each sample. Entities in the pathway diagram are highlighted based on values in the MeanDiff column in the IPA Pathway Analysis tab.
+
+
+
+CellCycleCheckPointsResults
+
+
+**Note** : You may change the color spectrum mapping for pathway diagram highlighting by double-clicking the color spectrum bar at the bottom of pathway diagram window to get the dialog for setting min/max values. You can save the analysis results for a pathway by using popup menu, "Save Analysis Results", and load the results back later on by "Open Analysis Results".
+ 4. **Analyze gene level results:** The up or down perturbation results are inferred based on genomic data files for individual genes. The App provides features to analyze observation results and inference results for individual genes. You can view gene-level observation and inference results for the whole pathway by using popup menus, "Show Gene Level Analysis Results" and "Show Observations". You can also view these results for genes contained by an entity displayed in the pathway diagram after selecting that entity and then using these two popup menus. The following two dialogs show gene level observations and inference results for genes whose products are contained by complex "hBUBR1:hBUB3:MAD2*:CDC20 complex [cytosol]" in the cell cycle checkpoints pathway:
+
+
+
+ObservationsForEntity
+
+
+
+GeneLevelResultsForEntity
+
+ 5. **Analyze and visualize results for individual samples:** The inference results and loaded observation data are displayed in the right "Results Panel" (see below). By checking "Highlight pathway for sample", entities in the displayed pathway diagram will be highlighted based on inferred IPA values for the selected sample displayed in the "Choose sample" box. You can also enable animation by clicking the play button. There are two tabs in the "Results Panel": "Inference" for showing inferred IPA values, and "Observation" tab for loaded observed data related to entities in the pathway (Note: if you choose "discretizing", the displayed observation values are discretized: 0 for lower than normal, 1 for normal, and 2 for higher normal). Objects in three views (pathway diagram, inference table, and observation table) are synchronized for selection.
+
+
+
+PGM Sample View: Inference
+
+
+
+PGM Sample View: Observation
+
+ 6. **Compare analysis results for two samples:** You can compare observation data and inference results for two samples. To do this, choose two samples in the "IPA Sample Analysis" tab in the bottom results pane, and use popup menu "Compare Samples" to bring out another tab called "Sample Comparison". You can view comparing results for inference and observation data.
+
+
+
+Two Sample Comparison
+
+#### Boolean Network Based Pathway Analysis
+
+We have developed an approach (Manuscript in preparation) to convert biochemical reactions-based Reactome pathways into Boolean networks and then perform pathway simulation based on the constrained fuzzy logic method according to [Training Signaling Pathway Maps to Biochemical Data with Constrained Fuzzy Logic: Quantitative Analysis of Liver Cell Responses to Inflammatory Stimuli]() and [Querying quantitative logic models (Q2LM) to study intracellular signaling networks and cell-cytokine interactions](). Based on this approach, the user can perform pathway simulation inside Cytoscape using rich Reactome pathways based on fuzzy logic built upon Boolean networks.
+
+ 1. **Set up and run logic model simulation** : Choose popup menu "Run Logic Model Analysis" in the Pathway Diagram View to get the New Simulation dialog. Enter a name for the simulation and the default value, which usually should be 1.0 to enable that the simulation can proceed, and then choose either PROD or MIN for the AND gate mode (the default choice PROD usually should be fine).
+**Note** : You may also choose an Transfer Function and adjust parameters for Hill function. However, for simplicity, it is suggested to use "Identity Function" first. For how to apply drugs for logic model simulation, see below.
+
+
+
+Run Boolean Network Analysis
+
+
+
+New BN Simulation
+
+
+After clicking the OK button in the New Simulation dialog, the default initial configuration will be displayed in the Results Panel. You may change the variable Type and Modification in the set up table by clicking the cell for the selected variable. To run the simulation, click the Simulate button in the Results Panel.
+
+
+
+Set up Boolean Network Analysis
+
+
+
+Choose BN Variable Type
+
+
+
+Choose BN Modification Type
+
+ 2. **Visualize the simulation results** : After the simulation is done, entities in the pathway diagram will be highlighted based on simulated values, which should be between 0 and 1. You may choose one or more entities to visualize their temporal behaviors inside the Table Panel at the bottom of Cytoscape. Attractors computed from the simulation are also listed in the right columns in the original set up table inside the Results Panel. **Note** : After simulation, you will not be able to modify any initial configuration.
+
+
+
+Boolean Network Simulation Results
+
+
+**Note** : To avoid clutter, if too many time steps have been generated for a logic model simulation, only the last 20 time steps are displayed in the table. However, the plot shows all time steps. You may choose different columns to display in the table by use popup menu "Configure Columns" after selecting any variables in the table.
+ 3. **Perform pathway simulation via modification** : Simulation with Boolean network can help users uncover the impact of modification of entity activities (e.g. inhibition or activation caused by somatic mutation) on the pathway behaviors. For example, in PIP3 activates AKT signaling, Complex AKT:PIP3 forms a complex with EntitySet THEM4/TRIB3 to form another complex, which inhibits the activation of AKT (For details see [Reactome PIP3 activates AKT signaling]()). To perform simulation with modification, choose modification type in the simulation set up table and assign the strength to the modification.
+
+
+
+Choose Inhibition for Boolean Network Simulation
+
+
+Clicking the Simulate button invokes the constrained fuzzy logic model simulation with this configured inhibition. You can compare the simulation results between the two configurations by selecting an entity and then toggling the simulation result tables at the bottom of Cytoscape.
+
+
+
+Active AKT in Inihibition
+
+
+
+
+
+Active AKT in Default
+
+
+You can also use the "Compare" button in the Results Panel to get the comparison dialog and then choose two simulations for comparison. The comparison results are displayed in a new table listed at the bottom Table Panel.
+
+
+
+Comparison Dialog
+
+
+
+
+
+Active AKT in Comparison
+
+
+**Note:** The RelativeDifference in the comparison result table is calculated based on relative change for each fuzzy logic variable, calculated as (valueInSim2 - valueInSim1) / (valueInSim2 + valueInSim1). The time course may be interpolated based on the attractor until this relative difference converges.
+ 4. To remove all displayed constrained fuzzy logic simulation results, use popup menu, "Remove Analysis Results" under "Run Logic Model Analysis". The pathway diagram should be reset to the original colors, and all tables related to logic model simulations will be deleted.
+
+#### Visualization of Structural Variants in the Context of Reactome Pathways
+
+By collaborating with Drs. Francesco Raimondi and Rob Russell at the University of Heidelberg to utilize protein-protein interaction 3D structures provided by [Mechismo](), a platform developed by [Dr. Russell's group]() to study the contributions of individual amino acid residues to protein structure and function, we have systematically analyzed mutations in the TCGA dataset and collected a set of reactions and functional interactions that involve proteins significantly enriched with mutations in their interaction interfaces (manuscript in preparation). We have added features to ReactomeFIViz to visualize these 3D structures and mutated residues in the contexts of Reactome pathways, reactions, and interactions.
+
+ 1. **Visualize analysis results in the context of a pathway and its reactions** : In the opened pathway diagram view (See [Explore Reactome Pathways](<#Explore_Reactome_Pathways>) for how to open a pathway diagram), use the popup menu "Load Mechismo Results" to load the analysis results into the opened pathway diagram. After the results are loaded, reactions are colored based on FDR values as listed in the bottom table labeled as "Mechismo Reaction".
+
+
+
+Mechismo Reaction View
+
+**Note** : You may choose results from a different cancer type or pancancer by clicking the list at the top of the bottom tab labeled as "Choose a cancer type to highlight reactions based on FDRs in the table". Some of reactions are not highlighted by any color since there are no structural variants found for proteins annotated for these reactions. To remove the loaded results from the pathway diagram, use another popup menu "Remove Mechismo Results".
+ 2. **Visualize analysis results in the context of a pathway FI network** : After the Mechismo results are loaded into the pathway diagram, you can convert the diagram into a FI network by using popup menu "Convert to FI Network" as usual. The edges displayed in the FI network view are highlighted based on FDR values listed in the bottom table labeled as "Mechismo Interaction".
+
+
+
+Mechismo Interaction View
+
+**Note** : To make the FI network view simplier, check "Show FIs Only for Selected" at the left-bottom corner for the pathway diagram view. You may choose a reaction or complex and then view extracted FIs for the selected object in the pathway diagram view. Some of reactions and complexes may not contain any FI (e.g. a complex composed of a protein and a chemical or a reaction between a protein and a chemical).
+ 3. **Visualize structural variants in the protein-protein 3D structures** : In the FI network view, choose the popup menu called "Fetch Mechismo Results" to bring the view for protein-protein interaction 3D models. Structural variants collected from the TCGA data set are mapped to the original protein-protein interaction 3D structures based on the [Mechismo]() platform.
+
+
+
+Mechismo Structure View
+
+**Note** : In the structure view, amino acid residues whose coordinates are mapped to structural variants are displayed in balls. The residues that are mapped to the selected rows in the bottom table are highlighted in yellow. ReactomeFIViz uses [Jmol]() for protein 3D structure visualization. For more information about how to use Jmol, see its documentation: .
+
+### Use the Reactome Functional Interaction (FI) Network
+
+After the ReactomeFIViz app installed, you should see a menu item called "Reactome FI" under the Apps menu. Clicking this menu, you will see 6 sub-menus: [Gene Set/Mutation Analysis](<#Gene_Set.2FMutation_Analysis>), [PGM Impact Analysis](<#PGM_Impact_Analysis>), [Microarray Data Analysis](<#Microarray_Data_Analysis>), [Reactome Pathways](<#Use_Reactome_Pathways>) and [User Guide.]() Gene set/mutation analysis is for doing FI network-based data analysis for a set of genes or a mutation data file, PGM Impact analysis for performing functional impact analysis based on a probabilistic graphical model for the Reactome FI network using multiple omics data types, HotNet mutation analysis for the HotNet algorithm to search for network modules (see ), microarray data analysis for doing MCL (Markov Graph Clustering, ) based FI network clustering analysis by converting a non-weighted FI network to weighted network using correlations among genes in the network, Reactome pathways for loading pathways from the Reactome database, visualizing Reactome pathways directly in Cytoscape in a their native way, and doing pathway enrichment analysis, and user guide brings you to this user guide.
+
+
+
+#### Gene Set/Mutation Analysis
+
+ 1. You can enter a list of genes directly into ReactomeFIViz by clicking the "Enter" button, or load it from a local file. Currently ReactomeFIViz supports three file formats for gene set/mutation analysis:
+ 1. **Simple gene set** : one line per gene. For example, [GWASFuzzyGenes.txt](), a list of T2D GWAS genes.
+ 2. **Gene/sample number pair**. For example, [GeneSampleNumber.txt](), which contains two required columns, gene and number of samples having gene mutated, and an optional third column listing sample names (delimited by ";").
+ 3. **NCI MAF (mutation annotation file)**. For example, [GlioblastomaMutationTable.txt](), the mutation file from the TCGA GBM project.
+ 2. Choose a FI network version from listed three versions.
+**Note** : you may get different results using different FI network versions because a later version may contain more proteins/genes and more FIs. But based on our experience, a significant FI network module is usually stable across multiple versions.
+ 3. Enter genes directly by clicking the "Enter" button or choose a file containing genes you want to use to construct a functional interaction network. To choose a file, select an appropriate file format and parameters to load genes and construct FI network in the dialog. Click the "OK" button to start the FI network building process.
+
+
+
+ 4. The constructed FI network will be displayed in the network view panel. A FI specific visual style will be created automatically for the FI network.
+
+
+
+Reactome FI Sub-Network
+
+ 5. The main features of Reactome FI plug-in should be invoked from a popup menu, which can be displayed by right clicking an empty space in the network view panel.
+
+
+
+ * **Fetch FI annotations** : query detailed information on selected FIs. Three FI related edge attribues will be created: FI Annotation, FI Direction, and FI Score. Edges will be displayed based on FI direction attribute values. In the following screenshot, "->" for activating/catalyzing, "-|" for inhibition, "-" for FIs extracted from complexes or inputs, and "---" for predicted FIs. See the "VizMapper" tab, Edge Source Arrow Shape and Edge Target Arrow Shape values for details.
+
+
+
+FI Annotations
+
+
+**Note** : Here is a short explanation about displayed columns: GeneSet for pathways collected in the Reactome FI network hit by the query gene list; RatioOfProteinInGeneSet for ratios of numbers of genes contained in pathways to total genes in the Reactome FI network; NumberOfProteinInGeneSet for numbers of genes in pathways; ProteinFromNetwork for numbers of hit genes from the query gene list; P-value for pvalues calculated based on binomial test; FDR for FDRs calculated based on p-values using Benjamini-Hocherberg method; Nodes for hit genes in pathways.
+ * **Analyze network functions** : pathway or GO term ennrichment analysis for the displayed network. You can choose to filter enrichment results by a FDR cutoff value. Also you can choose to display nodes in the network panel for a selected row or rows by checking "Hide nodes in not selected rows". The letter in parentheses after each pathway gene set name corresponds to the source of the pathway annotations: C - CellMap, R – Reactome, K – KEGG, N – NCI PID, P - Panther, and B – BioCarta. The following screenshot shows results from a pathway enrichment analysis.
+
+
+
+Pathways in FI Sub-Network
+
+
+_Tip: To analyze pathway or GO term enrichment on a set of genes that are not linked together, select the "Show genes not linked to others" option in the "Set Parameters for FI Network" dialog._
+ * **Cluster FI network** : run a network clustering algorithm (spectral partition based network clustering by [Newman 2006]()) on the displayed FI network. Nodes in different network modules will be shown in different colors (different colors used only for first 15 modules based on sizes).
+
+
+
+Network Modules
+
+ * **Analyze module functions** : pathway or GO term enrichment analysis for each individual network modules. You can select a size cutoff to filter out network modules that are too small, choose a FDR cutoff to view enriched pathways or GO terms under a certain FDR value, and view nodes in a selected row or rows only in the network diagram.
+ * **Analyze functions for a set of selected genes** : select a set of nodes displayed in the network view and then choose the popup menu, Analyze Nodes Functions, to perform pathway or GO term enrichment analysis. The results will be displayed in a dialog.
+ * **Load Cancer Gene Index** : load cancer gene index annotations. For details, see section [Load Cancer Gene Index](<#Load_Cancer_Gene_Index_Annotations>).
+
+#### PGM Impact Analysis
+
+ 1. We have developed a probabilistic graphical model (PGM)-based functional impact analysis using the Reactome FI network by integrating multiple omics data types together. The current version of ReactomeFIViz supports four omics data types: CNV, mRNA expression, DNA methylation, and somatic mutation. PGMs used for this analysis are based on [Markov random field (MRF)](). Currently we support two types of MRFs: [Pairwise MRF]() and [Nearest neighbor Gibbs MRF](). You can choose one of these two models. We recommend pair-wise MRF for its simplicity.
+ 2. To perform PGM-based functional impact analysis, choose menu, Apps/Rectome FI/PGM Impact Analysis/Analyze. You can enter your omics data by using the PGM configuration dialog.
+
+
+
+FI-PGM Configuration Dialog
+
+
+**Notes** :
+1). ReactomeFIViz supports continuous observation variables without discretizing by choosing "Use empirical distribution" in the configuration. For somatic mutation, currently it supports NCI MAF file format only and requires a specific column named "MA_FI.score" for mutation function impact score collected from [Mutation Assessor]() or from some other sources.
+2). The default MRF model used is PairwiseMRF. You can choose the default setting for the first test.
+3). Several parameters are needed for MRF models. We have tuned these parameters based on a small toy model. In the current version of ReactomeFIViz, these parameters cannot be changed.
+ 3. It may take several hours to finish the whole analysis. The actual running time will be dependent on the size of your data. The progress of the job running is displayed in the following progress pane. You can cancel the running at any time.
+
+
+
+Progress of FI PGM Running
+
+ 4. After the analysis is finished, you should see the result dialog similar to the following screenshot. You can use filtering features to filter to a list of genes that you want to use to construct a FI subnetwork for further analysis.
+
+
+
+FI-PGM Result Dialog
+
+
+**Note** : If you select one or more genes in the result table, only these selected genes will be used to construct a FI sub-network. You can save the full analysis results by clicking the "Save" button (Results for all genes, not just displayed ones, will be saved). We strongly recommend to save your results first before clicking the "OK" button so that you can visit your results back.
+ 5. After choosing the "OK" button, a FI subnetwork will be constructed and displayed in a network view. The sizes of displayed nodes are proportional to impact scores inferred from the FI-PGM model. To view impact scores and loaded observation data, you can choose a sample in the Sample List tab in the Results Panel. You can also enable sample-based network visualization of the network by checking "Highlight network for sample" in the Sample List tab. If you want to review the original results used to construct the FI subnetwork, click "Show All Results" in the "Impact Gene Values" tab in "Table Panel".
+
+
+
+FI-PGM Result Network
+
+#### Microarray Data Analysis
+
+The ReactomeFIViz app can load gene expression data file, calculate correlations among genes involved in the same FIs, use the calculated correlations as weights for edges (i.e. FIs) in the whole FI network, apply MCL graph clustering algorithm to the weighted FI network, and generate a sub-network for a list of selected network modules based on module size and average correlation. The generated FI sub-network will be displayed in the network panel, and can be used for analysis as in Gene Set/Mutation Analysis. For details about this method, please see our publication: [A network module-based method for identifying cancer prognostic signatures]().
+
+An array data file should be a tab-delimited text file with table headers. The first column should be gene names. All other columns should be expression values in different samples. **The data set in the file should be pre-normalized.** For example, see this gene expression file for breast cancer: [NejmLogRatioNormGlobalZScore_070111.txt.zip](). This data set was download from [van de Vijver et al in 2002](), and has been normalized.
+
+ 1. **Select a microarray data file and run MCL network clustering** : After selecting sub-menu "Microarray Data Analysis" from menu Plugins/Reactome FIs, you should see the following dialog. Choose a microarray data file, check if you want to use absolute values as weights for edges, and input an inflation parameter (-I) for the MCL clustering algorithm. The smaller the inflation parameter is, the bigger the average size of generated network modules. Based on our own experience, we use 5.0 for the inflation parameter, the highest recommended value, and choose the absolute value for edge weights. For more details on how to choose the inflation parameter, please see . After you have set these parameters, click the OK button to load the data file, calculate correlations, and apply the MCL clustering algorithm.
+
+
+
+Set Parameters for Microarray Data Analysis
+
+ 2. **Select network modules and build a FI sub-network** : The generated network modules are listed in the MCL clustering results dialog (see below). Only modules having more than 2 genes can be listed, and used in the FI sub-network building. You can choose a module size or an average correlation value (absolute value if absolute has been checked before) to filter out modules that may not be significant (Note: after set these cutoff values, please press the "Enter" key to commit your changes.). In our analysis, we choose modules having 7 or more genes with average correlation values no less than 0.25. These values have been used as default in the dialog. In the dialog, you can see how many modules and genes will be chosen for building FI sub-network under your selected filter values. Click the OK button to start the sub-network building. The built sub-network will be displayed, and can be analyzed as with sub-networks generated from the gene set/mutation analysis.
+
+
+
+Choose MCL Modules
+
+### Visualize Drugs in the Contexts of Reactome Pathways and FI Network
+
+ReactomeFIViz provide a suite of features to assist users to visualize drugs in the contexts of Reactome pathways and networks. The drug data sources include two: Cancer Targetome ([Blucher et al 2017]()), which collected all FDA-approved cancer drugs (prior to 2018) and their target interactions from four sources, including DrugBank, Therapeutic Targets Database, IUPHAR, and BindingDB; [DrugCentral](), a comprehensive drug database supported by [NIH IDG program]().
+
+#### Visualize Drugs in Reactome Pathways
+
+ 1. **List all FDA approved cancer drugs** : Use popup menu "View Cancer Drugs" in the Reactome pathway tree to get the list of all FDA approved cancer drugs collected in Cancer Targetome in a table dialog.
+
+
+
+View Cancer Drugs
+
+
+A screenshot of this drug list is shown below:
+
+
+
+List of Cancer Drugs
+
+ 2. **View drug/target interactions** : Select a drug in the drug table (See Figure above) to perform Google search, or view its targets by clicking one of buttons in the dialog. Two views are provided for drug/target interactions. Drug targets table view (below) shows targets for the selected drug with collected binding affinities, which are divided into different categories (e.g.): KD, IC50, Ki, and EC50. The displayed targets are pre-filtered using a set of default filters. You may change the default filters in the Drug/Target Interaction Filter dialog by clicking the "Filter" button.
+
+
+
+Drug Targets Table View
+
+
+
+Drug Target Interaction Filter
+
+In the Drug targets table view, you can choose an interaction and then click the "View Details" button to view the detailed information for the selected interaction. In the details view, you can browse the original pubmed references for experimental evidence, Gene Card entry for target, or Google drug by clicking the hyper links.
+
+
+
+Cancer Drug Interaction Details View
+
+You can also select multiple rows in the table to perform pathway enrichment analysis by clicking "Overlay Targets to Pathways". See [Pathway Enrichment Analysis](<#Pathway_Enrichment_Analysis>) for details about pathway enrichment analysis results.
+
+Drug targets plot view (below) provides a stagged plot of bar charts for evidence-supported interactions between the selected drug and all its targets, suggesting primary target(s) and secondary targets in a single graphical view.
+
+
+
+Drug Targets Plot View
+
+ 3. **View cancer drugs in pathway diagrams** : Cancer drugs can be overlaid directly onto displayed pathway diagrams using the popup menu, "Fetch Cancer Drugs", in the [Pathway Diagram View](). The fetched cancer drugs are displayed in the pathway diagram, linked to entities containing targets for the displayed drugs.
+
+
+
+Fetch Cancer Drugs
+
+
+
+Drugs in Pathway Diagram
+
+**Note:** Use Popup menu "Filter Drugs" to adjust filters to select drugs and interactions in the pathway diagram view; Use "Remove Overlaid Interactions" to remove displayed drugs; You may choose a displayed entity in the pathway diagram and then use popup menu "Fetch Cancer Drugs" to display drugs targeted to the selected entity only.
+
+
+
+To view the detailed information about the interaction between the displayed drug and its targeted entity, choose the link and use pupup menu "Show Details" to bring up the drug targets view.
+
+
+
+View Details
+
+ 4. **Perform systems pathway impact analysis** : In the cancer drug list table (see above), you may perform pathway impact analysis for all Reactome pathways having entity level view drawn by choosing the "Run Pathway Impact Analysis" button. After the analysis is done, the impact results are displayed in the table labled with the selected drug in the Cytoscape Table Panel. See below for an example of the results for Imatinib:
+
+
+
+Pathway Impact Analysis Results
+
+**Note** : You may open the pathway diagram using popup menu "View in Diagram" and export the table into a text file using menu "Export Table" in the results table.
+
+#### Visualize Cancer Drugs in the FI Network
+
+The Reactome FI network provides a network-based view among proteins/genes, where each gene/protein is displayed only once. Visualizing cancer drugs in a FI network context shows a simplified relationship between cancer drugs and their targets. In the [FI Network View](<#Gene_Set.2FMutation_Analysis>), use popup menu, "Reactome FI/Overlay Cancer Drugs/Fetch Cancer Drugs", to load cancer drugs for proteins/genes displayed in the FI network. The loaded drugs and interactions between drugs and proteins/genes are rendered in green diamonds and blue edges, respectively.
+
+
+
+Fetch Cancer Drugs in FI Network
+
+
+
+Show Drug Target In FI Network
+
+
+**Note** : To remove overlaid drugs in the FI network view, use popup menu, "Reactome FI/Overlay Cancer Drugs/Remove Drugs"; As in the Pathway diagram view, you can also apply filters by using "Filte Drugs" popup menu; To view the details about an interaction between a drug and a target, select the edge, and then use popup menu "Reactome FI/Show Drug/Target Interaction Details". (You may need to zoom into the selected edge.)
+
+#### Visualize DrugCentral Drugs
+
+Visualize drugs collected in DrugCentral is simlar with cancer drugs collected in Cancer Targetome. You can use popup menu "View DrugCentral Drugs" in the pathway tree to list all drugs collected in the DrugCentral database, "Fetch DrugCentral Drugs" popup menu in the pathway diagram view to overlay drugs onto pathways, or "ReactomeFI/Overlay Drugs/Fetch DrugCentral Drugs" to overlay drugs onto the FI network in the network view.
+
+#### Simulate Impact of Drugs on Pathway Activities
+
+Overlaying cancer drugs onto the contexts of Reactome pathways and its FI network helps users to understand the potential impact of applying drugs on the pathway activities and network behavior. However, the actual perturbation of drugs on pathways may be much more complicated. Performing pathway simulation may help users to understand the actual impact. ReactomeFIViz implements features to assist users to perform Boolean network-based drug simulation. Before you do drug simulation, please read section [Boolean Network Based Pathway Analysis](<#Boolean_Network_based_Pathway_Analysis>) first. In this section, we will use pathway [HDR through Homologous Recombination (HR) or Single Strand Annealing (SSA)]() and drug Imatinib as an example (Note: To get the following screenshot, open diagram for this pathway, and then fetch cancer drugs and filter drugs to Imatinib using the name filter).
+
+
+
+Imatinib in Pathway
+
+ 1. **Set up new simulation for drug** : In the pathway diagram view, use popup menu, "Run Logic Model Analysis". In the New Simulation dialog, enter a name (e.g. Imtatinib) and default value for the simulation, and then choose the mode for the AND gate as in the regular logic model simulation. To perform drug simulation, choose the Drug Application tab and a drug data source, and then click the "..." button to bring up the Drug Selection dialog. **
+Note** : You will need to adjust the drug filters to show all targets for Imatinib as displayed in the following screenshot. Based on the collected annotations for interactions between cancer drugs and their targets, default modification types will be selected (as expected, most of them are inhibition). Strengths of modifications are pre-configured based on affinities collected in our aggregated drug/target database.
+
+
+
+Imatinib BN Simulation I
+
+
+
+Imatinib BN Simulation II
+
+
+
+
+
+Select Drugs for BN Simulation
+
+
+**Note:** Since no affinities can be found for interactions between Imatinib and CHEK1 or CDK2, CHEK1 and CDK2 are not listed in the New Simulation dialog. Checking "Filter members in sets to drug targets" will select members in EntitySet instances that are targeted by drugs to force showing of potential pathway impact caused by drugs.
+ 2. **Perform simulation with drug** : The configuration for drug in the Drug Selection dialog will be copied into the Boolean Network configuration table displayed in the Results Panel. Click the "Simulate" button to perform simulation. After the simulation is done, Entities in the pathway diagrams are highlighted in different colors based on values in the attractor with detailed temporal values displayed in the table at the bottom Table Panel.
+
+
+
+Imatinib BN Setup
+
+
+
+
+
+Imatinib BN Results
+
+ 3. **Investigate the drug impact on pathway activities** : To see the impact of a drug on the pathway activities, perform another Boolean network simulation without applying cancer drugs (here as Default) (For details, see [Boolean Network Based Pathway Analysis](<#Boolean_Network_based_Pathway_Analysis>)). The screenshot for the logic model simulation results with the default initial configuration for pathway "HDR through Homologous Recombination (HR) or Single Strand Annealing (SSA)" is displayed below:
+
+
+
+HDR Through HR or SSA Default Results
+
+
+To see the drug impact to the activity of an entity displayed in the pathway, choose that entity and check its temporal behavior in both BN:Default and BN:Imatinib tables. For example, below a complex (see above screenshot) related to ABL1 is selected (Up for imatinib applied and down for default without drug):
+
+
+
+Imatinib One Variable
+
+
+
+
+
+Default One Variable
+
+
+You may also use the "Compare" button to check the detailed difference in the computed attractors from two simulations.
+
+
+
+p-S456-ABL1 Values
+
+
+**Note** : From the above comparison, we can see that application of imatinib will significantly impact the formation of the complex in HDR, an effect of imanitib on DNA repair pathway has been reported by others (e.g. [Imatinib (STI571) induces DNA damage in BCR/ABL-expressing leukemic cells but not in normal lymphocytes]()). You may see a little bit different simulation results because of update in pathway annotations in new versions of ReactomeFIViz.
+
+### Perform scRNA-seq Data Analysis and Visualization
+
+ReactomeFIViz implements a suite of features for users to conduct scRNA-seq data analysis and visualization. To do this, we have packaged several popuplar Python packages developed for scRNA-seq data analysis and visualization together into a Python standalone application. These packages include [scanpy]() for routine scRNA-seq data analysis and visualization and [scVelo]() for RNA velocity based data analysis and visualization.
+**Note** : For scRNA-seq data analysis and visualization, you need to have Python 3.7 installed at your computer. If you have not installed Python at your computer, you can do so by downloading an installer from [https://www.python.org/downloads]() for your computer. We have tested Python 3.7 only and thefore suggest that you use 3.7 for these features. However, you don't need to install our standalone Python application indepedently from ReactomeFIViz. When needed, ReactomeFIViz will automatically download and update the application for you as long as you point to the correct Python application path (i.e. directory and application file).
+
+#### Standard Analysis via Scanpy
+
+ 1. **Set up the analysis:** The Python package, [scanpy](), provides a set of powerful analysis and visualization features for scRNA-seq data. ReactomeFIViz wraps these features for users to take advantage of pathway and network analysis and visualization facilities provided by Cytoscape in general and ReactomeFIViz in particular. To conduct a scRNA-seq analysis using scanpy, choose menu Apps/Reactome FI/Single Cell Analysis/Analyze to get the configuration window as shown below:
+
+
+
+scRNA-seq Analysis Configuration
+
+ReactomeFIViz supports scRNA-seq data generated from mouse and human. You should choose the species for your data and the format. If your data is in the 10x-Genomics-mtx format, you should choose the directory containing the files in that format. You may check an imputation method. Currently, ReactomeFIViz supports the [MAGIC]() approach only. You may also check total_counts and/or pct_counts_mt for [regress out]() to control unwanted variations.
+**Note** : All analysis steps and their paramters are logged into CytoscapeConfiguration/ReactomeFIViz/ReactomeFIViz.{date}.log in your user folder for your review.
+ 2. **Configure Python for ReactomeFIViz:** If you have not done so, you will be asked to set up Python for ReactomeFIViz using the following configuration dialog when ReactomeFIViz downloads the Reactome Python app for scRNA-seq data analysis and visualization.
+
+
+
+Set up Python
+
+**Note** : Currently only Python 3.7 is supported.
+ReactomeFIViz uses the functions provided by scanpy for pre-processing, normalization, UMAP analysis, cell clustering and all other scRNA-seq analysis except imputation, which is handled by [MAGIC](), if checked. See details in the scanpy document: . For paramters used for these functions, open the ReactomeFIViz.{data}.log file (see above).
+ 3. **Visuzlize Cell Networks** : Dependent on the sample size and the computing power, it may take several minutes to finish the analysis. After that, two networks, one for cell clusters and another for single cells, are displayed in Cytoscape and listed under "SingleCellClusterNetwork" and "SingleCellNetwork" in the left-side, Network tab, respectively.
+
+
+
+ScRNA-seq Cluster Network
+
+
+
+ScRNA-seq Cell Network
+
+**Note** : You may use the built-in Cytoscape Style features and other configuration properties to adjust the rendering of these two networks. See Cytoscape's user manual by clicking menu Help/User Manual. To show or hide edges in the networks, use popup menu Reactome FI/Show Edges (see below for a screenshot). Cluster in the cell cluster network are named based on the rank of cell clusters sorted by cell numbers in the clusters. For example, cluster0 has the largest number of cells.
+ 4. **Analyze scRNA-seq Data** : To explore the loaded scRNA-seq data and perform further analysis, you can use the popup menu provided in the cell cluster or single cell network view as shown below:
+
+
+
+ScRNA-seq Standard Analysis Popup Menus
+
+ * **Load Gene Expression** : Overlay expression value for a gene onto the network. You may enter the gene name from the input dialog after clicking this menu.
+**Note** : Cell clusters use the median values of cells in the clusters for coloring for gene expression and cell features (below).
+ * **Load Cell Features** : Overlay cell features by choosing a sub-menu, e.g., n_genes (total genes), n_genes_by_counts (total genes having counts), total_counts, total_count_mt (for mitonchorian genes), pct_counts_mt (percent of mitochondria genes), and leiden (network clustering results based on the [Leiden]() algorithm).
+**Note** : Both the cell cluster network and the single cell network are colored based on the leiden clustering results when they are rendered after the analysis. To get back to the orignal colors, choose Load Cell Feature/leiden.
+ * **CytoTrace Analysis** : Perform CytoTrace analysis to predict the differential state of cells based on the number of detected expressed genes per cell. ReactomeFIViz provides a Python implementation of CytoTrace based on the original R code published in . For details about CytoTrace, see the original paper: [Single-cell transcriptional diversity is a hallmark of developmental potential](). You can find more information in the CytoTrace's web site: